+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Methylcrotonoyl-CoA carboxylase core-Dimer | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | MCC / Mitochondria / CYTOSOLIC PROTEIN | |||||||||
| Function / homology |  Function and homology information Function and homology information3-Methylcrotonyl-CoA carboxylase deficiency / methylcrotonoyl-CoA carboxylase / methylcrotonoyl-CoA carboxylase complex / methylcrotonoyl-CoA carboxylase activity / L-leucine catabolic process / Defective HLCS causes multiple carboxylase deficiency / Biotin transport and metabolism / Branched-chain amino acid catabolism / branched-chain amino acid catabolic process / coenzyme A metabolic process ...3-Methylcrotonyl-CoA carboxylase deficiency / methylcrotonoyl-CoA carboxylase / methylcrotonoyl-CoA carboxylase complex / methylcrotonoyl-CoA carboxylase activity / L-leucine catabolic process / Defective HLCS causes multiple carboxylase deficiency / Biotin transport and metabolism / Branched-chain amino acid catabolism / branched-chain amino acid catabolic process / coenzyme A metabolic process / mitochondrial matrix / mitochondrion / ATP binding / cytosol Similarity search - Function | |||||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 4.16 Å | |||||||||
 Authors Authors | Liu DS / Su JY | |||||||||
| Funding support |  China, 1 items China, 1 items
| |||||||||
 Citation Citation |  Journal: Nat Struct Mol Biol / Year: 2025 Journal: Nat Struct Mol Biol / Year: 2025Title: Structural insight into synergistic activation of human 3-methylcrotonyl-CoA carboxylase. Authors: Jiayue Su / Xuyang Tian / Hang Cheng / Desheng Liu / Ziyi Wang / Shan Sun / Hong-Wei Wang / Sen-Fang Sui /   Abstract: The enzymes 3-methylcrotonyl-coenzyme A (CoA) carboxylase (MCC), pyruvate carboxylase and propionyl-CoA carboxylase belong to the biotin-dependent carboxylase family located in mitochondria. They ...The enzymes 3-methylcrotonyl-coenzyme A (CoA) carboxylase (MCC), pyruvate carboxylase and propionyl-CoA carboxylase belong to the biotin-dependent carboxylase family located in mitochondria. They participate in various metabolic pathways in human such as amino acid metabolism and tricarboxylic acid cycle. Many human diseases are caused by mutations in those enzymes but their structures have not been fully resolved so far. Here we report an optimized purification strategy to obtain high-resolution structures of intact human endogenous MCC, propionyl-CoA carboxylase and pyruvate carboxylase in different conformational states. We also determine the structures of MCC bound to different substrates. Analysis of MCC structures in different states reveals the mechanism of the substrate-induced, multi-element synergistic activation of MCC. These results provide important insights into the catalytic mechanism of the biotin-dependent carboxylase family and are of great value for the development of new drugs for the treatment of related diseases. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_36024.map.gz emd_36024.map.gz | 62.5 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-36024-v30.xml emd-36024-v30.xml emd-36024.xml emd-36024.xml | 15.3 KB 15.3 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_36024.png emd_36024.png | 109.1 KB | ||
| Filedesc metadata |  emd-36024.cif.gz emd-36024.cif.gz | 5.6 KB | ||
| Others |  emd_36024_half_map_1.map.gz emd_36024_half_map_1.map.gz emd_36024_half_map_2.map.gz emd_36024_half_map_2.map.gz | 116 MB 115.9 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-36024 http://ftp.pdbj.org/pub/emdb/structures/EMD-36024 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-36024 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-36024 | HTTPS FTP |
-Related structure data
| Related structure data |  8j73MC  7ybuC  8hwlC  8j4zC  8j78C  8j7dC  8j7oC  8jakC  8jawC  8jxlC  8jxmC  8jxnC  8k2vC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_36024.map.gz / Format: CCP4 / Size: 125 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_36024.map.gz / Format: CCP4 / Size: 125 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.0979 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Half map: #2
| File | emd_36024_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
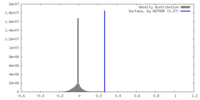
| Density Histograms |
-Half map: #1
| File | emd_36024_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Methylcrotonoyl-CoA carboxylase core-Dimer
| Entire | Name: Methylcrotonoyl-CoA carboxylase core-Dimer |
|---|---|
| Components |
|
-Supramolecule #1: Methylcrotonoyl-CoA carboxylase core-Dimer
| Supramolecule | Name: Methylcrotonoyl-CoA carboxylase core-Dimer / type: complex / ID: 1 / Parent: 0 / Macromolecule list: all |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
-Macromolecule #1: Methylcrotonoyl-CoA carboxylase beta chain, mitochondrial
| Macromolecule | Name: Methylcrotonoyl-CoA carboxylase beta chain, mitochondrial type: protein_or_peptide / ID: 1 / Number of copies: 12 / Enantiomer: LEVO / EC number: methylcrotonoyl-CoA carboxylase |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 61.406027 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: MWAVLRLALR PCARASPAGP RAYHGDSVAS LGTQPDLGSA LYQENYKQMK ALVNQLHERV EHIKLGGGEK ARALHISRGK LLPRERIDN LIDPGSPFLE LSQFAGYQLY DNEEVPGGGI ITGIGRVSGV ECMIIANDAT VKGGAYYPVT VKKQLRAQEI A MQNRLPCI ...String: MWAVLRLALR PCARASPAGP RAYHGDSVAS LGTQPDLGSA LYQENYKQMK ALVNQLHERV EHIKLGGGEK ARALHISRGK LLPRERIDN LIDPGSPFLE LSQFAGYQLY DNEEVPGGGI ITGIGRVSGV ECMIIANDAT VKGGAYYPVT VKKQLRAQEI A MQNRLPCI YLVDSGGAYL PRQADVFPDR DHFGRTFYNQ AIMSSKNIAQ IAVVMGSCTA GGAYVPAMAD ENIIVRKQGT IF LAGPPLV KAATGEEVSA EDLGGADLHC RKSGVSDHWA LDDHHALHLT RKVVRNLNYQ KKLDVTIEPS EEPLFPADEL YGI VGANLK RSFDVREVIA RIVDGSRFTE FKAFYGDTLV TGFARIFGYP VGIVGNNGVL FSESAKKGTH FVQLCCQRNI PLLF LQNIT GFMVGREYEA EGIAKDGAKM VAAVACAQVP KITLIIGGSY GAGNYGMCGR AYSPRFLYIW PNARISVMGG EQAAN VLAT ITKDQRAREG KQFSSADEAA LKEPIIKKFE EEGNPYYSSA RVWDDGIIDP ADTRLVLGLS FSAALNAPIE KTDFGI FRM UniProtKB: Methylcrotonoyl-CoA carboxylase beta chain, mitochondrial |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 8 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 (6k x 4k) / Average electron dose: 50.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 1.8 µm / Nominal defocus min: 1.3 µm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller
















 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)