+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Human pyruvate carboxylase in BCCP-CTS state without BC | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | PC / Mitochondrial / CYTOSOLIC PROTEIN | |||||||||
| Function / homology |  Function and homology information Function and homology informationpyruvate carboxylase / pyruvate carboxylase activity / Defective HLCS causes multiple carboxylase deficiency / Biotin transport and metabolism / viral RNA genome packaging / NADP+ metabolic process / host-mediated activation of viral process / Pyruvate metabolism / Gluconeogenesis / NAD+ metabolic process ...pyruvate carboxylase / pyruvate carboxylase activity / Defective HLCS causes multiple carboxylase deficiency / Biotin transport and metabolism / viral RNA genome packaging / NADP+ metabolic process / host-mediated activation of viral process / Pyruvate metabolism / Gluconeogenesis / NAD+ metabolic process / biotin binding / viral release from host cell / gluconeogenesis / lipid metabolic process / mitochondrial matrix / negative regulation of gene expression / mitochondrion / ATP binding / metal ion binding / identical protein binding / cytoplasm / cytosol Similarity search - Function | |||||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 3.83 Å | |||||||||
 Authors Authors | Liu DS / Su JY | |||||||||
| Funding support |  China, 1 items China, 1 items
| |||||||||
 Citation Citation |  Journal: Nat Struct Mol Biol / Year: 2025 Journal: Nat Struct Mol Biol / Year: 2025Title: Structural insight into synergistic activation of human 3-methylcrotonyl-CoA carboxylase. Authors: Jiayue Su / Xuyang Tian / Hang Cheng / Desheng Liu / Ziyi Wang / Shan Sun / Hong-Wei Wang / Sen-Fang Sui /   Abstract: The enzymes 3-methylcrotonyl-coenzyme A (CoA) carboxylase (MCC), pyruvate carboxylase and propionyl-CoA carboxylase belong to the biotin-dependent carboxylase family located in mitochondria. They ...The enzymes 3-methylcrotonyl-coenzyme A (CoA) carboxylase (MCC), pyruvate carboxylase and propionyl-CoA carboxylase belong to the biotin-dependent carboxylase family located in mitochondria. They participate in various metabolic pathways in human such as amino acid metabolism and tricarboxylic acid cycle. Many human diseases are caused by mutations in those enzymes but their structures have not been fully resolved so far. Here we report an optimized purification strategy to obtain high-resolution structures of intact human endogenous MCC, propionyl-CoA carboxylase and pyruvate carboxylase in different conformational states. We also determine the structures of MCC bound to different substrates. Analysis of MCC structures in different states reveals the mechanism of the substrate-induced, multi-element synergistic activation of MCC. These results provide important insights into the catalytic mechanism of the biotin-dependent carboxylase family and are of great value for the development of new drugs for the treatment of related diseases. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_36044.map.gz emd_36044.map.gz | 62.2 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-36044-v30.xml emd-36044-v30.xml emd-36044.xml emd-36044.xml | 14.3 KB 14.3 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_36044.png emd_36044.png | 116.1 KB | ||
| Filedesc metadata |  emd-36044.cif.gz emd-36044.cif.gz | 5.9 KB | ||
| Others |  emd_36044_half_map_1.map.gz emd_36044_half_map_1.map.gz emd_36044_half_map_2.map.gz emd_36044_half_map_2.map.gz | 115.8 MB 115.9 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-36044 http://ftp.pdbj.org/pub/emdb/structures/EMD-36044 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-36044 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-36044 | HTTPS FTP |
-Related structure data
| Related structure data |  8j7oMC  7ybuC  8hwlC  8j4zC  8j73C  8j78C  8j7dC  8jakC  8jawC  8jxlC  8jxmC  8jxnC  8k2vC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_36044.map.gz / Format: CCP4 / Size: 125 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_36044.map.gz / Format: CCP4 / Size: 125 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.0979 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Half map: #2
| File | emd_36044_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
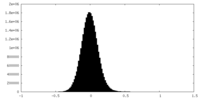
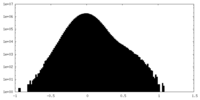
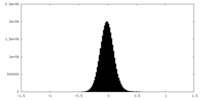
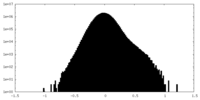
| Density Histograms |
-Half map: #1
| File | emd_36044_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Tetramer complex of human pyruvate carboxylase
| Entire | Name: Tetramer complex of human pyruvate carboxylase |
|---|---|
| Components |
|
-Supramolecule #1: Tetramer complex of human pyruvate carboxylase
| Supramolecule | Name: Tetramer complex of human pyruvate carboxylase / type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1 |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
-Macromolecule #1: Pyruvate carboxylase, mitochondrial
| Macromolecule | Name: Pyruvate carboxylase, mitochondrial / type: protein_or_peptide / ID: 1 / Number of copies: 4 / Enantiomer: LEVO / EC number: pyruvate carboxylase |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 129.799359 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: MLKFRTVHGG LRLLGIRRTS TAPAASPNVR RLEYKPIKKV MVANRGEIAI RVFRACTELG IRTVAIYSEQ DTGQMHRQKA DEAYLIGRG LAPVQAYLHI PDIIKVAKEN NVDAVHPGYG FLSERADFAQ ACQDAGVRFI GPSPEVVRKM GDKVEARAIA I AAGVPVVP ...String: MLKFRTVHGG LRLLGIRRTS TAPAASPNVR RLEYKPIKKV MVANRGEIAI RVFRACTELG IRTVAIYSEQ DTGQMHRQKA DEAYLIGRG LAPVQAYLHI PDIIKVAKEN NVDAVHPGYG FLSERADFAQ ACQDAGVRFI GPSPEVVRKM GDKVEARAIA I AAGVPVVP GTDAPITSLH EAHEFSNTYG FPIIFKAAYG GGGRGMRVVH SYEELEENYT RAYSEALAAF GNGALFVEKF IE KPRHIEV QILGDQYGNI LHLYERDCSI QRRHQKVVEI APAAHLDPQL RTRLTSDSVK LAKQVGYENA GTVEFLVDRH GKH YFIEVN SRLQVEHTVT EEITDVDLVH AQIHVAEGRS LPDLGLRQEN IRINGCAIQC RVTTEDPARS FQPDTGRIEV FRSG EGMGI RLDNASAFQG AVISPHYDSL LVKVIAHGKD HPTAATKMSR ALAEFRVRGV KTNIAFLQNV LNNQQFLAGT VDTQF IDEN PELFQLRPAQ NRAQKLLHYL GHVMVNGPTT PIPVKASPSP TDPVVPAVPI GPPPAGFRDI LLREGPEGFA RAVRNH PGL LLMDTTFRDA HQSLLATRVR THDLKKIAPY VAHNFSKLFS MENWGGATFD VAMRFLYECP WRRLQELREL IPNIPFQ ML LRGANAVGYT NYPDNVVFKF CEVAKENGMD VFRVFDSLNY LPNMLLGMEA AGSAGGVVEA AISYTGDVAD PSRTKYSL Q YYMGLAEELV RAGTHILCIK DMAGLLKPTA CTMLVSSLRD RFPDLPLHIH THDTSGAGVA AMLACAQAGA DVVDVAADS MSGMTSQPSM GALVACTRGT PLDTEVPMER VFDYSEYWEG ARGLYAAFDC TATMKSGNSD VYENEIPGGQ YTNLHFQAHS MGLGSKFKE VKKAYVEANQ MLGDLIKVTP SSKIVGDLAQ FMVQNGLSRA EAEAQAEELS FPRSVVEFLQ GYIGVPHGGF P EPFRSKVL KDLPRVEGRP GASLPPLDLQ ALEKELVDRH GEEVTPEDVL SAAMYPDVFA HFKDFTATFG PLDSLNTRLF LQ GPKIAEE FEVELERGKT LHIKALAVSD LNRAGQRQVF FELNGQLRSI LVKDTQAMKE MHFHPKALKD VKGQIGAPMP GKV IDIKVV AGAKVAKGQP LCVLSAMKME TVVTSPMEGT VRKVHVTKDM TLEGDDLILE IE UniProtKB: Pyruvate carboxylase, mitochondrial |
-Macromolecule #2: 5-(HEXAHYDRO-2-OXO-1H-THIENO[3,4-D]IMIDAZOL-6-YL)PENTANAL
| Macromolecule | Name: 5-(HEXAHYDRO-2-OXO-1H-THIENO[3,4-D]IMIDAZOL-6-YL)PENTANAL type: ligand / ID: 2 / Number of copies: 4 / Formula: BTI |
|---|---|
| Molecular weight | Theoretical: 228.311 Da |
| Chemical component information |  ChemComp-BTI: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 8 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 (6k x 4k) / Average electron dose: 50.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 1.8 µm / Nominal defocus min: 1.3 µm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
- Image processing
Image processing
| Startup model | Type of model: INSILICO MODEL |
|---|---|
| Final reconstruction | Resolution.type: BY AUTHOR / Resolution: 3.83 Å / Resolution method: FSC 0.143 CUT-OFF / Number images used: 21326 |
| Initial angle assignment | Type: RANDOM ASSIGNMENT |
| Final angle assignment | Type: ANGULAR RECONSTITUTION |
 Movie
Movie Controller
Controller























 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)