[English] 日本語
 Yorodumi
Yorodumi- EMDB-31142: Cryo-EM structure of human ABCB8 transporter in nucleotide bindin... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-31142 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
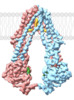
| Title | Cryo-EM structure of human ABCB8 transporter in nucleotide binding state | |||||||||
 Map data Map data | Electron density map | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | ABC transporter / MEMBRANE PROTEIN | |||||||||
| Function / homology |  Function and homology information Function and homology informationmitochondrial ATP-gated potassium channel complex / mitochondrial potassium ion transmembrane transport / ABC-type peptide transporter activity / Mitochondrial ABC transporters / cell volume homeostasis / potassium ion transmembrane transport / ATP-binding cassette (ABC) transporter complex / mitochondrial membrane / transmembrane transport / mitochondrial inner membrane ...mitochondrial ATP-gated potassium channel complex / mitochondrial potassium ion transmembrane transport / ABC-type peptide transporter activity / Mitochondrial ABC transporters / cell volume homeostasis / potassium ion transmembrane transport / ATP-binding cassette (ABC) transporter complex / mitochondrial membrane / transmembrane transport / mitochondrial inner membrane / nucleolus / ATP hydrolysis activity / mitochondrion / nucleoplasm / ATP binding / membrane Similarity search - Function | |||||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | |||||||||
| Method | single particle reconstruction / cryo EM | |||||||||
 Authors Authors | Li SJ / Yang X | |||||||||
 Citation Citation |  Journal: Biochem Biophys Res Commun / Year: 2021 Journal: Biochem Biophys Res Commun / Year: 2021Title: Cryo-EM structure of human ABCB8 transporter in nucleotide binding state. Authors: Shunjin Li / Yue Ren / Xuhang Lu / Yuequan Shen / Xue Yang /  Abstract: Human ATP-binding cassette transporter 8 of subfamily B (hABCB8) is an ABC transporter that located in the inner membrane of mitochondria. The ABCB8 is involved in the maturation of Fe-S and protects ...Human ATP-binding cassette transporter 8 of subfamily B (hABCB8) is an ABC transporter that located in the inner membrane of mitochondria. The ABCB8 is involved in the maturation of Fe-S and protects the heart from oxidative stress. Here, we present the cryo-EM structure of human ABCB8 binding with AMPPNP in inward-facing conformation with resolution of 4.1 Å. hABCB8 shows an open-inward conformation when ATP is bound. Unexpectedly, cholesterol molecules were identified in the transmembrane domain of hABCB8. Our results provide structural basis for the transport mechanism of the ABC transporter in mitochondria. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_31142.map.gz emd_31142.map.gz | 49.7 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-31142-v30.xml emd-31142-v30.xml emd-31142.xml emd-31142.xml | 10 KB 10 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_31142.png emd_31142.png | 149.7 KB | ||
| Filedesc metadata |  emd-31142.cif.gz emd-31142.cif.gz | 5.3 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-31142 http://ftp.pdbj.org/pub/emdb/structures/EMD-31142 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-31142 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-31142 | HTTPS FTP |
-Validation report
| Summary document |  emd_31142_validation.pdf.gz emd_31142_validation.pdf.gz | 501 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_31142_full_validation.pdf.gz emd_31142_full_validation.pdf.gz | 500.6 KB | Display | |
| Data in XML |  emd_31142_validation.xml.gz emd_31142_validation.xml.gz | 6.1 KB | Display | |
| Data in CIF |  emd_31142_validation.cif.gz emd_31142_validation.cif.gz | 6.9 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-31142 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-31142 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-31142 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-31142 | HTTPS FTP |
-Related structure data
| Related structure data |  7ehlMC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_31142.map.gz / Format: CCP4 / Size: 52.7 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_31142.map.gz / Format: CCP4 / Size: 52.7 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Electron density map | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.014 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : ABCB8
| Entire | Name: ABCB8 |
|---|---|
| Components |
|
-Supramolecule #1: ABCB8
| Supramolecule | Name: ABCB8 / type: cell / ID: 1 / Parent: 0 / Macromolecule list: #1 |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
-Macromolecule #1: Mitochondrial potassium channel ATP-binding subunit
| Macromolecule | Name: Mitochondrial potassium channel ATP-binding subunit / type: protein_or_peptide / ID: 1 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 73.628375 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: SLLRAVAHLR SQLWAHLPRA PLAPRWSPSA WCWVGGALLG PMVLSKHPHL CLVALCEAEE APPASSTPHV VGSRFNWKLF WQFLHPHLL VLGVAVVLAL GAALVNVQIP LLLGQLVEVV AKYTRDHVGS FMTESQNLST HLLILYGVQG LLTFGYLVLL S HVGERMAV ...String: SLLRAVAHLR SQLWAHLPRA PLAPRWSPSA WCWVGGALLG PMVLSKHPHL CLVALCEAEE APPASSTPHV VGSRFNWKLF WQFLHPHLL VLGVAVVLAL GAALVNVQIP LLLGQLVEVV AKYTRDHVGS FMTESQNLST HLLILYGVQG LLTFGYLVLL S HVGERMAV DMRRALFSSL LRQDITFFDA NKTGQLVSRL TTDVQEFKSS FKLVISQGLR SCTQVAGCLV SLSMLSTRLT LL LMVATPA LMGVGTLMGS GLRKLSRQCQ EQIARAMGVA DEALGNVRTV RAFAMEQREE ERYGAELEAC RCRAEELGRG IAL FQGLSN IAFNCMVLGT LFIGGSLVAG QQLTGGDLMS FLVASQTVQR SMANLSVLFG QVVRGLSAGA RVFEYMALNP CIPL SGGCC VPKEQLRGSV TFQNVCFSYP CRPGFEVLKD FTLTLPPGKI VALVGQSGGG KTTVASLLER FYDPTAGVVM LDGRD LRTL DPSWLRGQVV GFISQEPVLF GTTIMENIRF GKLEASDEEV YTAAREANAH EFITSFPEGY NTVVGERGTT LSGGQK QRL AIARALIKQP TVLILDEATS ALDAESERVV QEALDRASAG RTVLVIAHRL STVRGAHCIV VMADGRVWEA GTHEELL KK GGLYAELIRR QALDAPRTAA PPPKKPEGPR SHQHKS UniProtKB: Mitochondrial potassium channel ATP-binding subunit |
-Macromolecule #2: CHOLESTEROL
| Macromolecule | Name: CHOLESTEROL / type: ligand / ID: 2 / Number of copies: 4 / Formula: CLR |
|---|---|
| Molecular weight | Theoretical: 386.654 Da |
| Chemical component information |  ChemComp-CLR: |
-Macromolecule #3: PHOSPHOAMINOPHOSPHONIC ACID-ADENYLATE ESTER
| Macromolecule | Name: PHOSPHOAMINOPHOSPHONIC ACID-ADENYLATE ESTER / type: ligand / ID: 3 / Number of copies: 2 / Formula: ANP |
|---|---|
| Molecular weight | Theoretical: 506.196 Da |
| Chemical component information |  ChemComp-ANP: |
-Macromolecule #4: MAGNESIUM ION
| Macromolecule | Name: MAGNESIUM ION / type: ligand / ID: 4 / Number of copies: 2 / Formula: MG |
|---|---|
| Molecular weight | Theoretical: 24.305 Da |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 6.5 |
|---|---|
| Vitrification | Cryogen name: ETHANE / Instrument: FEI VITROBOT MARK IV |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Detector mode: COUNTING / Average electron dose: 56.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
- Image processing
Image processing
| Startup model | Type of model: OTHER / Details: Ab-initio in CryoSPARC-2 |
|---|---|
| Final reconstruction | Number images used: 43131 |
| Initial angle assignment | Type: ANGULAR RECONSTITUTION |
| Final angle assignment | Type: ANGULAR RECONSTITUTION |
 Movie
Movie Controller
Controller












 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)





















