[English] 日本語
 Yorodumi
Yorodumi- EMDB-31005: CryoEM structure of the human Kv4.2-KChIP1 complex, transmembrane... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-31005 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | CryoEM structure of the human Kv4.2-KChIP1 complex, transmembrane region | |||||||||
 Map data Map data | map B | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | ion channel / MEMBRANE PROTEIN | |||||||||
| Function / homology |  Function and homology information Function and homology informationKv4.2-KChIP2 channel complex / A-type (transient outward) potassium channel activity / Phase 1 - inactivation of fast Na+ channels / voltage-gated monoatomic ion channel activity involved in regulation of postsynaptic membrane potential / membrane repolarization / Voltage gated Potassium channels / postsynaptic specialization membrane / anchoring junction / regulation of heart contraction / neuronal cell body membrane ...Kv4.2-KChIP2 channel complex / A-type (transient outward) potassium channel activity / Phase 1 - inactivation of fast Na+ channels / voltage-gated monoatomic ion channel activity involved in regulation of postsynaptic membrane potential / membrane repolarization / Voltage gated Potassium channels / postsynaptic specialization membrane / anchoring junction / regulation of heart contraction / neuronal cell body membrane / action potential / voltage-gated potassium channel activity / plasma membrane raft / locomotor rhythm / neuronal action potential / GABA-ergic synapse / voltage-gated potassium channel complex / potassium ion transmembrane transport / sensory perception of pain / muscle contraction / protein homooligomerization / cellular response to hypoxia / chemical synaptic transmission / perikaryon / postsynaptic membrane / dendritic spine / neuronal cell body / glutamatergic synapse / metal ion binding / plasma membrane Similarity search - Function | |||||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | |||||||||
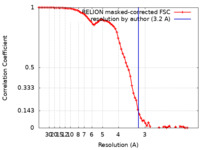
| Method | single particle reconstruction / cryo EM / Resolution: 3.2 Å | |||||||||
 Authors Authors | Kise Y / Nureki O | |||||||||
| Funding support |  Japan, 1 items Japan, 1 items
| |||||||||
 Citation Citation |  Journal: Nature / Year: 2021 Journal: Nature / Year: 2021Title: Structural basis of gating modulation of Kv4 channel complexes. Authors: Yoshiaki Kise / Go Kasuya / Hiroyuki H Okamoto / Daichi Yamanouchi / Kan Kobayashi / Tsukasa Kusakizako / Tomohiro Nishizawa / Koichi Nakajo / Osamu Nureki /  Abstract: Modulation of voltage-gated potassium (Kv) channels by auxiliary subunits is central to the physiological function of channels in the brain and heart. Native Kv4 tetrameric channels form ...Modulation of voltage-gated potassium (Kv) channels by auxiliary subunits is central to the physiological function of channels in the brain and heart. Native Kv4 tetrameric channels form macromolecular ternary complexes with two auxiliary β-subunits-intracellular Kv channel-interacting proteins (KChIPs) and transmembrane dipeptidyl peptidase-related proteins (DPPs)-to evoke rapidly activating and inactivating A-type currents, which prevent the backpropagation of action potentials. However, the modulatory mechanisms of Kv4 channel complexes remain largely unknown. Here we report cryo-electron microscopy structures of the Kv4.2-DPP6S-KChIP1 dodecamer complex, the Kv4.2-KChIP1 and Kv4.2-DPP6S octamer complexes, and Kv4.2 alone. The structure of the Kv4.2-KChIP1 complex reveals that the intracellular N terminus of Kv4.2 interacts with its C terminus that extends from the S6 gating helix of the neighbouring Kv4.2 subunit. KChIP1 captures both the N and the C terminus of Kv4.2. In consequence, KChIP1 would prevent N-type inactivation and stabilize the S6 conformation to modulate gating of the S6 helices within the tetramer. By contrast, unlike the reported auxiliary subunits of voltage-gated channel complexes, DPP6S interacts with the S1 and S2 helices of the Kv4.2 voltage-sensing domain, which suggests that DPP6S stabilizes the conformation of the S1-S2 helices. DPP6S may therefore accelerate the voltage-dependent movement of the S4 helices. KChIP1 and DPP6S do not directly interact with each other in the Kv4.2-KChIP1-DPP6S ternary complex. Thus, our data suggest that two distinct modes of modulation contribute in an additive manner to evoke A-type currents from the native Kv4 macromolecular complex. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_31005.map.gz emd_31005.map.gz | 5.9 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-31005-v30.xml emd-31005-v30.xml emd-31005.xml emd-31005.xml | 13.5 KB 13.5 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_31005_fsc.xml emd_31005_fsc.xml | 7.6 KB | Display |  FSC data file FSC data file |
| Images |  emd_31005.png emd_31005.png | 44.8 KB | ||
| Masks |  emd_31005_msk_1.map emd_31005_msk_1.map | 36.3 MB |  Mask map Mask map | |
| Filedesc metadata |  emd-31005.cif.gz emd-31005.cif.gz | 5 KB | ||
| Others |  emd_31005_half_map_1.map.gz emd_31005_half_map_1.map.gz emd_31005_half_map_2.map.gz emd_31005_half_map_2.map.gz | 24.7 MB 24.8 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-31005 http://ftp.pdbj.org/pub/emdb/structures/EMD-31005 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-31005 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-31005 | HTTPS FTP |
-Validation report
| Summary document |  emd_31005_validation.pdf.gz emd_31005_validation.pdf.gz | 679.3 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_31005_full_validation.pdf.gz emd_31005_full_validation.pdf.gz | 678.9 KB | Display | |
| Data in XML |  emd_31005_validation.xml.gz emd_31005_validation.xml.gz | 13 KB | Display | |
| Data in CIF |  emd_31005_validation.cif.gz emd_31005_validation.cif.gz | 18 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-31005 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-31005 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-31005 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-31005 | HTTPS FTP |
-Related structure data
| Related structure data |  7e7zMC  7e83C  7e84C  7e87C  7e89C  7e8bC  7e8eC  7e8gC  7e8hC  7f0jC  7f3fC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_31005.map.gz / Format: CCP4 / Size: 36.3 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_31005.map.gz / Format: CCP4 / Size: 36.3 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | map B | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.00226 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
-Mask #1
| File |  emd_31005_msk_1.map emd_31005_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
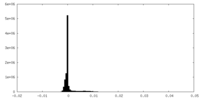
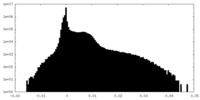
| Density Histograms |
-Half map: #1
| File | emd_31005_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #2
| File | emd_31005_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : CryoEM structure of the human Kv4.2-KChIP1 complex, transmembrane...
| Entire | Name: CryoEM structure of the human Kv4.2-KChIP1 complex, transmembrane region |
|---|---|
| Components |
|
-Supramolecule #1: CryoEM structure of the human Kv4.2-KChIP1 complex, transmembrane...
| Supramolecule | Name: CryoEM structure of the human Kv4.2-KChIP1 complex, transmembrane region type: organelle_or_cellular_component / ID: 1 / Parent: 0 / Macromolecule list: all |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
-Macromolecule #1: Potassium voltage-gated channel subfamily D member 2
| Macromolecule | Name: Potassium voltage-gated channel subfamily D member 2 / type: protein_or_peptide / ID: 1 / Number of copies: 4 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 28.327346 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: PTMTARQRVW RAFENPHTST MALVFYYVTG FFIAVSVIAN VVETVPCGSS PGHIKELPCG ERYAVAFFCL DTACVMIFTV EYLLRLAAA PSRYRFVRSV MSIIDVVAIL PYYIGLVMTD NEDVSGAFVT LRVFRVFRIF KFSRHSQGLR ILGYTLKSCA S ELGFLLFS ...String: PTMTARQRVW RAFENPHTST MALVFYYVTG FFIAVSVIAN VVETVPCGSS PGHIKELPCG ERYAVAFFCL DTACVMIFTV EYLLRLAAA PSRYRFVRSV MSIIDVVAIL PYYIGLVMTD NEDVSGAFVT LRVFRVFRIF KFSRHSQGLR ILGYTLKSCA S ELGFLLFS LTMAIIIFAT VMFYAEKGSS ASKFTSIPAA FWYTIVTMTT LGYGDMVPKT IAGKIFGSIC SLSGVLVIAL PV PVIVSNF SRIYHQNQ UniProtKB: Potassium voltage-gated channel subfamily D member 2 |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 8 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 BIOQUANTUM (6k x 4k) / Average electron dose: 48.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
+ Image processing
Image processing
-Atomic model buiding 1
| Refinement | Space: REAL |
|---|---|
| Output model |  PDB-7e7z: |
 Movie
Movie Controller
Controller


























 Z
Z Y
Y X
X