[English] 日本語
 Yorodumi
Yorodumi- EMDB-30014: V1-ATPase built from cryo-EM map of V/A-ATPase from Thermus therm... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-30014 | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Title | V1-ATPase built from cryo-EM map of V/A-ATPase from Thermus thermophilus. | ||||||||||||
 Map data Map data | Cryo-EM maps of V/A-ATPase from Thermus thermophilus. Soluble domain including V1, d, two EG stalks, and N-terminal domain of a-subunit. | ||||||||||||
 Sample Sample |
| ||||||||||||
 Keywords Keywords | rotary ATPase / V/A-ATPase / molecular motor / MOTOR PROTEIN | ||||||||||||
| Function / homology |  Function and homology information Function and homology informationproton motive force-driven plasma membrane ATP synthesis / proton-transporting ATPase activity, rotational mechanism / H+-transporting two-sector ATPase / proton-transporting ATP synthase complex / proton-transporting ATP synthase activity, rotational mechanism / ATP binding Similarity search - Function | ||||||||||||
| Biological species |   Thermus thermophilus HB8 (bacteria) Thermus thermophilus HB8 (bacteria) | ||||||||||||
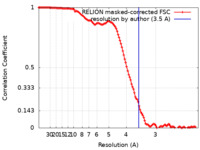
| Method | single particle reconstruction / cryo EM / Resolution: 3.5 Å | ||||||||||||
 Authors Authors | Kishikawa J / Nakanishi A / Furuta A / Kato T / Namba K / Tamakoshi M / Mitsuoka K / Yokoyama K | ||||||||||||
| Funding support |  Japan, 3 items Japan, 3 items
| ||||||||||||
 Citation Citation |  Journal: Elife / Year: 2020 Journal: Elife / Year: 2020Title: Mechanical inhibition of isolated V from V/A-ATPase for proton conductance. Authors: Jun-Ichi Kishikawa / Atsuko Nakanishi / Aya Furuta / Takayuki Kato / Keiichi Namba / Masatada Tamakoshi / Kaoru Mitsuoka / Ken Yokoyama /  Abstract: V-ATPase is an energy converting enzyme, coupling ATP hydrolysis/synthesis in the hydrophilic V domain, with proton flow through the V membrane domain, via rotation of the central rotor complex ...V-ATPase is an energy converting enzyme, coupling ATP hydrolysis/synthesis in the hydrophilic V domain, with proton flow through the V membrane domain, via rotation of the central rotor complex relative to the surrounding stator apparatus. Upon dissociation from the V domain, the V domain of the eukaryotic V-ATPase can adopt a physiologically relevant auto-inhibited form in which proton conductance through the V domain is prevented, however the molecular mechanism of this inhibition is not fully understood. Using cryo-electron microscopy, we determined the structure of both the V/A-ATPase and isolated V at near-atomic resolution, respectively. These structures clarify how the isolated V domain adopts the auto-inhibited form and how the complex prevents formation of the inhibited V form. | ||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_30014.map.gz emd_30014.map.gz | 10.7 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-30014-v30.xml emd-30014-v30.xml emd-30014.xml emd-30014.xml | 23 KB 23 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_30014_fsc.xml emd_30014_fsc.xml | 12.1 KB | Display |  FSC data file FSC data file |
| Images |  emd_30014.png emd_30014.png | 70.8 KB | ||
| Filedesc metadata |  emd-30014.cif.gz emd-30014.cif.gz | 7.7 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-30014 http://ftp.pdbj.org/pub/emdb/structures/EMD-30014 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-30014 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-30014 | HTTPS FTP |
-Related structure data
| Related structure data |  6ly8MC  6ly9C M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_30014.map.gz / Format: CCP4 / Size: 149.9 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_30014.map.gz / Format: CCP4 / Size: 149.9 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Cryo-EM maps of V/A-ATPase from Thermus thermophilus. Soluble domain including V1, d, two EG stalks, and N-terminal domain of a-subunit. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.1 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : Soluble domain of V/A-ATPase including V1, d, two EG stalks, and ...
| Entire | Name: Soluble domain of V/A-ATPase including V1, d, two EG stalks, and N-terminal of a-subunit |
|---|---|
| Components |
|
-Supramolecule #1: Soluble domain of V/A-ATPase including V1, d, two EG stalks, and ...
| Supramolecule | Name: Soluble domain of V/A-ATPase including V1, d, two EG stalks, and N-terminal of a-subunit type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1-#4 / Details: V/A-type ATPase from Thermus thermophilus |
|---|---|
| Source (natural) | Organism:   Thermus thermophilus HB8 (bacteria) / Strain: HB8 Thermus thermophilus HB8 (bacteria) / Strain: HB8 |
| Molecular weight | Theoretical: 500 KDa |
-Macromolecule #1: V-type ATP synthase alpha chain
| Macromolecule | Name: V-type ATP synthase alpha chain / type: protein_or_peptide / ID: 1 / Number of copies: 3 / Enantiomer: LEVO / EC number: H+-transporting two-sector ATPase |
|---|---|
| Source (natural) | Organism:   Thermus thermophilus HB8 (bacteria) / Strain: HB8 Thermus thermophilus HB8 (bacteria) / Strain: HB8 |
| Molecular weight | Theoretical: 63.69998 KDa |
| Sequence | String: MIQGVIQKIA GPAVIAKGML GARMYDICKV GEEGLVGEII RLDGDTAFVQ VYEDTSGLKV GEPVVSTGLP LAVELGPGML NGIYDGIQR PLERIREKTG IYITRGVVVH ALDREKKWAW TPMVKPGDEV RGGMVLGTVP EFGFTHKILV PPDVRGRVKE V KPAGEYTV ...String: MIQGVIQKIA GPAVIAKGML GARMYDICKV GEEGLVGEII RLDGDTAFVQ VYEDTSGLKV GEPVVSTGLP LAVELGPGML NGIYDGIQR PLERIREKTG IYITRGVVVH ALDREKKWAW TPMVKPGDEV RGGMVLGTVP EFGFTHKILV PPDVRGRVKE V KPAGEYTV EEPVVVLEDG TELKMYHTWP VRRARPVQRK LDPNTPFLTG MRILDVLFPV AMGGTAAIPG PFGSGKTVTQ QS LAKWSNA DVVVYVGCGE RGNEMTDVLV EFPELTDPKT GGPLMHRTVL IANTSNMPVA AREASIYVGV TIAEYFRDQG FSV ALMADS TSRWAEALRE ISSRLEEMPA EEGYPPYLAA RLAAFYERAG KVITLGGEEG AVTIVGAVSP PGGDMSEPVT QSTL RIVGA FWRLDASLAF RRHFPAINWN GSYSLFTSAL DPWYRENVAE DYPELRDAIS ELLQREAGLQ EIVQLVGPDA LQDAE RLVI EVGRIIREDF LQQNAYHEVD AYCSMKKAYG IMKMILAFYK EAEAAIKRGV SIDEILQLPV LERIGRARYV SEEEFP AYF EEAMKEIQGA FKALA UniProtKB: V-type ATP synthase alpha chain |
-Macromolecule #2: V-type ATP synthase beta chain
| Macromolecule | Name: V-type ATP synthase beta chain / type: protein_or_peptide / ID: 2 / Number of copies: 3 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:   Thermus thermophilus HB8 (bacteria) / Strain: HB8 Thermus thermophilus HB8 (bacteria) / Strain: HB8 |
| Molecular weight | Theoretical: 53.2195 KDa |
| Sequence | String: MDLLKKEYTG ITYISGPLLF VENAKDLAYG AIVDIKDGTG RVRGGQVIEV SEEYAVIQVF EETTGLDLAT TSVSLVEDVA RLGVSKEML GRRFNGIGKP IDGLPPITPE KRLPITGLPL NPVARRKPEQ FIQTGISTID VMNTLVRGQK LPIFSGSGLP A NEIAAQIA ...String: MDLLKKEYTG ITYISGPLLF VENAKDLAYG AIVDIKDGTG RVRGGQVIEV SEEYAVIQVF EETTGLDLAT TSVSLVEDVA RLGVSKEML GRRFNGIGKP IDGLPPITPE KRLPITGLPL NPVARRKPEQ FIQTGISTID VMNTLVRGQK LPIFSGSGLP A NEIAAQIA RQATVRPDLS GEGEKEEPFA VVFAAMGITQ RELSYFIQEF ERTGALSRSV LFLNKADDPT IERILTPRMA LT VAEYLAF EHDYHVLVIL TDMTNYCEAL REIGAAREEI PGRRGYPGYM YTDLATIYER AGVVEGKKGS VTQIPILSMP DDD RTHPIP DLTGYITEGQ IQLSRELHRK GIYPPIDPLP SLSRLMNNGV GKGKTREDHK QVSDQLYSAY ANGVDIRKLV AIIG EDALT ENDRRYLQFA DAFERFFINQ GQQNRSIEES LQIAWALLSM LPQGELKRIS KDHIGKYYGQ KLEEIWGAPQ ALD UniProtKB: V-type ATP synthase beta chain |
-Macromolecule #3: V-type ATP synthase subunit D
| Macromolecule | Name: V-type ATP synthase subunit D / type: protein_or_peptide / ID: 3 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:   Thermus thermophilus HB8 (bacteria) / Strain: HB8 Thermus thermophilus HB8 (bacteria) / Strain: HB8 |
| Molecular weight | Theoretical: 24.715566 KDa |
| Sequence | String: MSQVSPTRMN LLQRRGQLRL AQKGVDLLKK KRDALVAEFF GLVREAMEAR KALDQAAKEA YAALLLAQAF DGPEVVAGAA LGVPPLEGV EAEVENVWGS KVPRLKATFP DGALLSPVGT PAYTLEASRA FRRYAEALIR VANTETRLKK IGEEIKKTTR R VNALEQVV ...String: MSQVSPTRMN LLQRRGQLRL AQKGVDLLKK KRDALVAEFF GLVREAMEAR KALDQAAKEA YAALLLAQAF DGPEVVAGAA LGVPPLEGV EAEVENVWGS KVPRLKATFP DGALLSPVGT PAYTLEASRA FRRYAEALIR VANTETRLKK IGEEIKKTTR R VNALEQVV IPGIRAQIRF IQQVLEQRER EDTFRLKRIK GKIEAREAEE EGGRPNPQVE IGAGL UniProtKB: V-type ATP synthase subunit D |
-Macromolecule #4: V-type ATP synthase subunit F
| Macromolecule | Name: V-type ATP synthase subunit F / type: protein_or_peptide / ID: 4 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:   Thermus thermophilus HB8 (bacteria) / Strain: HB8 Thermus thermophilus HB8 (bacteria) / Strain: HB8 |
| Molecular weight | Theoretical: 11.294904 KDa |
| Sequence | String: MAVIADPETA QGFRLAGLEG YGASSAEEAQ SLLETLVERG GYALVAVDEA LLPDPERAVE RLMRGRDLPV LLPIAGLKEA FQGHDVEGY MRELVRKTIG FDIKL UniProtKB: V-type ATP synthase subunit F |
-Macromolecule #5: ADENOSINE-5'-DIPHOSPHATE
| Macromolecule | Name: ADENOSINE-5'-DIPHOSPHATE / type: ligand / ID: 5 / Number of copies: 1 / Formula: ADP |
|---|---|
| Molecular weight | Theoretical: 427.201 Da |
| Chemical component information |  ChemComp-ADP: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 3.0 mg/mL | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Buffer | pH: 8 Component:
| |||||||||
| Grid | Model: Quantifoil R1.2/1.3 / Material: MOLYBDENUM / Mesh: 300 / Pretreatment - Type: GLOW DISCHARGE / Pretreatment - Time: 60 sec. / Pretreatment - Atmosphere: AIR | |||||||||
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 277 K / Instrument: FEI VITROBOT MARK IV | |||||||||
| Details | The sample was purified from cell membrane of Thermus thermophilus and incorporated into nanodisc. |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: FEI FALCON II (4k x 4k) / Detector mode: INTEGRATING / Digitization - Dimensions - Width: 4096 pixel / Digitization - Dimensions - Height: 4096 pixel / Digitization - Frames/image: 1-34 / Number real images: 3694 / Average exposure time: 2.0 sec. / Average electron dose: 45.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Calibrated defocus max: 3.5 µm / Calibrated defocus min: 2.0 µm / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal magnification: 75000 |
| Sample stage | Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
+ Image processing
Image processing
-Atomic model buiding 1
| Initial model | PDB ID: Chain - Source name: PDB / Chain - Initial model type: experimental model |
|---|---|
| Refinement | Space: REAL / Protocol: FLEXIBLE FIT / Target criteria: Correlation coefficient |
| Output model |  PDB-6ly8: |
 Movie
Movie Controller
Controller













 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)