+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-25798 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | AtTPC1 D454N with 1 mM Ca2+ | |||||||||
 Map data Map data | Non-uniform refinement, post-processed with Deepemhancer. Inputs are the two provided half maps, high resolution protocol. | |||||||||
 Sample Sample |
| |||||||||
| Function / homology |  Function and homology information Function and homology informationregulation of jasmonic acid biosynthetic process /  seed germination / regulation of stomatal movement / plant-type vacuole / seed germination / regulation of stomatal movement / plant-type vacuole /  vacuole / vacuolar membrane / monoatomic ion channel complex / vacuole / vacuolar membrane / monoatomic ion channel complex /  voltage-gated calcium channel activity / calcium-mediated signaling / calcium ion transport ...regulation of jasmonic acid biosynthetic process / voltage-gated calcium channel activity / calcium-mediated signaling / calcium ion transport ...regulation of jasmonic acid biosynthetic process /  seed germination / regulation of stomatal movement / plant-type vacuole / seed germination / regulation of stomatal movement / plant-type vacuole /  vacuole / vacuolar membrane / monoatomic ion channel complex / vacuole / vacuolar membrane / monoatomic ion channel complex /  voltage-gated calcium channel activity / calcium-mediated signaling / calcium ion transport / voltage-gated calcium channel activity / calcium-mediated signaling / calcium ion transport /  calcium ion binding / calcium ion binding /  Golgi apparatus / identical protein binding / Golgi apparatus / identical protein binding /  plasma membrane / plasma membrane /  cytosol cytosolSimilarity search - Function | |||||||||
| Biological species |   Arabidopsis thaliana (thale cress) Arabidopsis thaliana (thale cress) | |||||||||
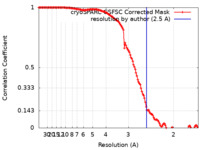
| Method |  single particle reconstruction / single particle reconstruction /  cryo EM / Resolution: 2.5 Å cryo EM / Resolution: 2.5 Å | |||||||||
 Authors Authors | Dickinson MS / Stroud RM | |||||||||
| Funding support |  United States, 1 items United States, 1 items
| |||||||||
 Citation Citation |  Journal: Proc Natl Acad Sci U S A / Year: 2022 Journal: Proc Natl Acad Sci U S A / Year: 2022Title: Molecular basis of multistep voltage activation in plant two-pore channel 1. Authors: Miles Sasha Dickinson / Jinping Lu / Meghna Gupta / Irene Marten / Rainer Hedrich / Robert M Stroud /   Abstract: Voltage-gated ion channels confer excitability to biological membranes, initiating and propagating electrical signals across large distances on short timescales. Membrane excitation requires channels ...Voltage-gated ion channels confer excitability to biological membranes, initiating and propagating electrical signals across large distances on short timescales. Membrane excitation requires channels that respond to changes in electric field and couple the transmembrane voltage to gating of a central pore. To address the mechanism of this process in a voltage-gated ion channel, we determined structures of the plant two-pore channel 1 at different stages along its activation coordinate. These high-resolution structures of activation intermediates, when compared with the resting-state structure, portray a mechanism in which the voltage-sensing domain undergoes dilation and in-membrane plane rotation about the gating charge-bearing helix, followed by charge translocation across the charge transfer seal. These structures, in concert with patch-clamp electrophysiology, show that residues in the pore mouth sense inhibitory Ca and are allosterically coupled to the voltage sensor. These conformational changes provide insight into the mechanism of voltage-sensor domain activation in which activation occurs vectorially over a series of elementary steps. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_25798.map.gz emd_25798.map.gz | 191.1 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-25798-v30.xml emd-25798-v30.xml emd-25798.xml emd-25798.xml | 17.9 KB 17.9 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_25798_fsc.xml emd_25798_fsc.xml | 12.7 KB | Display |  FSC data file FSC data file |
| Images |  emd_25798.png emd_25798.png | 65.3 KB | ||
| Others |  emd_25798_half_map_1.map.gz emd_25798_half_map_1.map.gz emd_25798_half_map_2.map.gz emd_25798_half_map_2.map.gz | 203.4 MB 203.4 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-25798 http://ftp.pdbj.org/pub/emdb/structures/EMD-25798 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-25798 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-25798 | HTTPS FTP |
-Related structure data
| Related structure data |  7tbgMC  7tddC  7tdeC  7tdfC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_25798.map.gz / Format: CCP4 / Size: 219.4 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_25798.map.gz / Format: CCP4 / Size: 219.4 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Non-uniform refinement, post-processed with Deepemhancer. Inputs are the two provided half maps, high resolution protocol. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.835 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
-Half map: Half map 1 from non-uniform refinement
| File | emd_25798_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Half map 1 from non-uniform refinement | ||||||||||||
| Projections & Slices |
| ||||||||||||
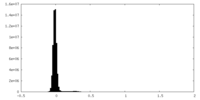
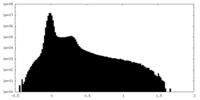
| Density Histograms |
-Half map: Half map 2 from non-uniform refinement
| File | emd_25798_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Half map 2 from non-uniform refinement | ||||||||||||
| Projections & Slices |
| ||||||||||||
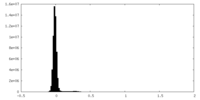
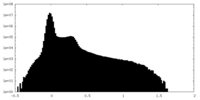
| Density Histograms |
- Sample components
Sample components
-Entire : Two-pore channel 1 dimer
| Entire | Name: Two-pore channel 1 dimer |
|---|---|
| Components |
|
-Supramolecule #1: Two-pore channel 1 dimer
| Supramolecule | Name: Two-pore channel 1 dimer / type: complex / Chimera: Yes / ID: 1 / Parent: 0 / Macromolecule list: #1 |
|---|---|
| Source (natural) | Organism:   Arabidopsis thaliana (thale cress) Arabidopsis thaliana (thale cress) |
| Recombinant expression | Organism:   Saccharomyces cerevisiae (brewer's yeast) Saccharomyces cerevisiae (brewer's yeast) |
| Molecular weight | Theoretical: 168 KDa |
-Macromolecule #1: Two pore calcium channel protein 1
| Macromolecule | Name: Two pore calcium channel protein 1 / type: protein_or_peptide / ID: 1 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:   Arabidopsis thaliana (thale cress) Arabidopsis thaliana (thale cress) |
| Molecular weight | Theoretical: 84.956836 KDa |
| Recombinant expression | Organism:   Saccharomyces cerevisiae (brewer's yeast) Saccharomyces cerevisiae (brewer's yeast) |
| Sequence | String: MEDPLIGRDS LGGGGTDRVR RSEAITHGTP FQKAAALVDL AEDGIGLPVE ILDQSSFGES ARYYFIFTRL DLIWSLNYFA LLFLNFFEQ PLWCEKNPKP SCKDRDYYYL GELPYLTNAE SIIYEVITLA ILLVHTFFPI SYEGSRIFWT SRLNLVKVAC V VILFVDVL ...String: MEDPLIGRDS LGGGGTDRVR RSEAITHGTP FQKAAALVDL AEDGIGLPVE ILDQSSFGES ARYYFIFTRL DLIWSLNYFA LLFLNFFEQ PLWCEKNPKP SCKDRDYYYL GELPYLTNAE SIIYEVITLA ILLVHTFFPI SYEGSRIFWT SRLNLVKVAC V VILFVDVL VDFLYLSPLA FDFLPFRIAP YVRVIIFILS IRELRDTLVL LSGMLGTYLN ILALWMLFLL FASWIAFVMF ED TQQGLTV FTSYGATLYQ MFILFTTSNN PDVWIPAYKS SRWSSVFFVL YVLIGVYFVT NLILAVVYDS FKEQLAKQVS GMD QMKRRM LEKAFGLIDS DKNGEIDKNQ CIKLFEQLTN YRTLPKISKE EFGLIFDELD DTRDFKINKD EFADLCQAIA LRFQ KEEVP SLFEHFPQIY HSALSQQLRA FVRSPNFGYA ISFILIINFI AVVVETTLNI EESSAQKNWQ VAEFVFGWIY VLEMA LKIY TYGFENYWRE GANRFDFLVT WVIVIGETAT FITPDENTFF SNGEWIRYLL LARMLRLIRL LMNVQRYRAF IATFIT LIP SLMPYLGTIF CVLCIYCSIG VQVFGGLVNA GNKKLFETEL AEDDYLLFNF NDYPNGMVTL FNLLVMGNWQ VWMESYK DL TGTWWSITYF VSFYVITILL LLNLVVAFVL EAFFTELDLE EEEKCQGQDS QEKRNRRRSA GSKSRSQRVD TLLHHMLG D ELSKPECSTS DT |
-Macromolecule #2: CALCIUM ION
| Macromolecule | Name: CALCIUM ION / type: ligand / ID: 2 / Number of copies: 4 / Formula: CA |
|---|---|
| Molecular weight | Theoretical: 40.078 Da |
-Macromolecule #3: SODIUM ION
| Macromolecule | Name: SODIUM ION / type: ligand / ID: 3 / Number of copies: 1 |
|---|---|
| Molecular weight | Theoretical: 22.99 Da |
-Macromolecule #4: water
| Macromolecule | Name: water / type: ligand / ID: 4 / Number of copies: 20 / Formula: HOH |
|---|---|
| Molecular weight | Theoretical: 18.015 Da |
| Chemical component information |  ChemComp-HOH: |
-Experimental details
-Structure determination
| Method |  cryo EM cryo EM |
|---|---|
 Processing Processing |  single particle reconstruction single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 5 mg/mL |
|---|---|
| Buffer | pH: 7.5 |
| Grid | Model: Quantifoil R1.2/1.3 / Material: GOLD / Mesh: 300 / Support film - Material: CARBON / Support film - topology: HOLEY / Pretreatment - Type: GLOW DISCHARGE / Pretreatment - Atmosphere: AIR |
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 280 K / Instrument: FEI VITROBOT MARK IV |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | C2 aperture diameter: 50.0 µm / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD Bright-field microscopy / Cs: 2.7 mm / Nominal defocus max: 1.5 µm / Nominal defocus min: 0.8 µm / Nominal magnification: 105000 Bright-field microscopy / Cs: 2.7 mm / Nominal defocus max: 1.5 µm / Nominal defocus min: 0.8 µm / Nominal magnification: 105000 |
| Specialist optics | Energy filter - Name: GIF Bioquantum / Energy filter - Slit width: 20 eV |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Image recording | Film or detector model: GATAN K3 BIOQUANTUM (6k x 4k) / Average electron dose: 66.0 e/Å2 |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
- Image processing
Image processing
-Atomic model buiding 1
| Refinement | Space: REAL / Protocol: FLEXIBLE FIT |
|---|---|
| Output model |  PDB-7tbg: |
 Movie
Movie Controller
Controller













 Z
Z Y
Y X
X