+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-22014 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | hEAAT3-OFS-Asp | |||||||||
 Map data Map data | outward facing hEAAT3 protomer with Asp and sodium bound | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | hEAAT3 outward-facing / full-bound / TRANSPORT PROTEIN | |||||||||
| Function / homology |  Function and homology information Function and homology informationD-aspartate transmembrane transport / regulation of protein targeting to membrane / D-aspartate transmembrane transporter activity / Defective SLC1A1 is implicated in schizophrenia 18 (SCZD18) and dicarboxylic aminoaciduria (DCBXA) / response to decreased oxygen levels / distal dendrite / response to anesthetic / cysteine transmembrane transporter activity / cysteine transport / neurotransmitter receptor transport to plasma membrane ...D-aspartate transmembrane transport / regulation of protein targeting to membrane / D-aspartate transmembrane transporter activity / Defective SLC1A1 is implicated in schizophrenia 18 (SCZD18) and dicarboxylic aminoaciduria (DCBXA) / response to decreased oxygen levels / distal dendrite / response to anesthetic / cysteine transmembrane transporter activity / cysteine transport / neurotransmitter receptor transport to plasma membrane / high-affinity L-glutamate transmembrane transporter activity / glutamate:sodium symporter activity / L-glutamate import / cellular response to mercury ion / Transport of inorganic cations/anions and amino acids/oligopeptides / zinc ion transmembrane transport / retina layer formation / L-glutamate transmembrane transport / L-glutamate transmembrane transporter activity / cellular response to bisphenol A / L-aspartate transmembrane transport / D-aspartate import across plasma membrane / cellular response to ammonium ion / glutathione biosynthetic process / righting reflex / monoatomic anion channel activity / L-aspartate transmembrane transporter activity / intracellular glutamate homeostasis / L-aspartate import across plasma membrane / grooming behavior / Glutamate Neurotransmitter Release Cycle / proximal dendrite / L-glutamate import across plasma membrane / transepithelial transport / apical dendrite / conditioned place preference / intracellular zinc ion homeostasis / glutamate receptor signaling pathway / cellular response to cocaine / blood vessel morphogenesis / neurotransmitter transport / motor behavior / motor neuron apoptotic process / chloride transmembrane transporter activity / response to morphine / glutamate binding / positive regulation of heart rate / G protein-coupled dopamine receptor signaling pathway / postsynaptic modulation of chemical synaptic transmission / adult behavior / maintenance of blood-brain barrier / heart contraction / superoxide metabolic process / dopamine metabolic process / perisynaptic space / transport across blood-brain barrier / glial cell projection / cellular response to organic cyclic compound / asymmetric synapse / response to axon injury / behavioral fear response / synaptic cleft / monoatomic ion transport / axon terminus / chloride transmembrane transport / response to amphetamine / neurogenesis / dendritic shaft / cell periphery / locomotory behavior / long-term synaptic potentiation / regulation of protein phosphorylation / synapse organization / Schaffer collateral - CA1 synapse / brain development / cytokine-mediated signaling pathway / memory / recycling endosome membrane / late endosome membrane / presynapse / cellular response to oxidative stress / gene expression / early endosome membrane / chemical synaptic transmission / perikaryon / negative regulation of neuron apoptotic process / dendritic spine / response to xenobiotic stimulus / apical plasma membrane / membrane raft / axon / neuronal cell body / dendrite / cell surface / endoplasmic reticulum / extracellular exosome / identical protein binding / membrane / metal ion binding / plasma membrane Similarity search - Function | |||||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | |||||||||
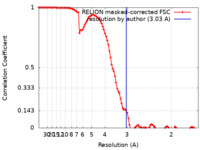
| Method | single particle reconstruction / cryo EM / Resolution: 3.03 Å | |||||||||
 Authors Authors | Qiu B / Matthies D | |||||||||
| Funding support |  United States, 1 items United States, 1 items
| |||||||||
 Citation Citation |  Journal: Sci Adv / Year: 2021 Journal: Sci Adv / Year: 2021Title: Cryo-EM structures of excitatory amino acid transporter 3 visualize coupled substrate, sodium, and proton binding and transport. Authors: Biao Qiu / Doreen Matthies / Eva Fortea / Zhiheng Yu / Olga Boudker /  Abstract: Human excitatory amino acid transporter 3 (hEAAT3) mediates glutamate uptake in neurons, intestine, and kidney. Here, we report cryo-EM structures of hEAAT3 in several functional states where the ...Human excitatory amino acid transporter 3 (hEAAT3) mediates glutamate uptake in neurons, intestine, and kidney. Here, we report cryo-EM structures of hEAAT3 in several functional states where the transporter is empty, bound to coupled sodium ions only, or fully loaded with three sodium ions, a proton, and the substrate aspartate. The structures suggest that hEAAT3 operates by an elevator mechanism involving three functionally independent subunits. When the substrate-binding site is near the cytoplasm, it has a remarkably low affinity for the substrate, perhaps facilitating its release and allowing the rapid transport turnover. The mechanism of the coupled uptake of the sodium ions and the substrate is conserved across evolutionarily distant families and is augmented by coupling to protons in EAATs. The structures further suggest a mechanism by which a conserved glutamate residue mediates proton symport. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_22014.map.gz emd_22014.map.gz | 85.3 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-22014-v30.xml emd-22014-v30.xml emd-22014.xml emd-22014.xml | 14.2 KB 14.2 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_22014_fsc.xml emd_22014_fsc.xml | 10.3 KB | Display |  FSC data file FSC data file |
| Images |  emd_22014.png emd_22014.png | 22.5 KB | ||
| Filedesc metadata |  emd-22014.cif.gz emd-22014.cif.gz | 5.8 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-22014 http://ftp.pdbj.org/pub/emdb/structures/EMD-22014 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-22014 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-22014 | HTTPS FTP |
-Validation report
| Summary document |  emd_22014_validation.pdf.gz emd_22014_validation.pdf.gz | 582.2 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_22014_full_validation.pdf.gz emd_22014_full_validation.pdf.gz | 581.7 KB | Display | |
| Data in XML |  emd_22014_validation.xml.gz emd_22014_validation.xml.gz | 11.6 KB | Display | |
| Data in CIF |  emd_22014_validation.cif.gz emd_22014_validation.cif.gz | 15.3 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-22014 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-22014 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-22014 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-22014 | HTTPS FTP |
-Related structure data
| Related structure data |  6x2zMC  6x2lC  6x3eC  6x3fC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_22014.map.gz / Format: CCP4 / Size: 91.1 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_22014.map.gz / Format: CCP4 / Size: 91.1 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | outward facing hEAAT3 protomer with Asp and sodium bound | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.832 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : hEAAT3 trimer
| Entire | Name: hEAAT3 trimer |
|---|---|
| Components |
|
-Supramolecule #1: hEAAT3 trimer
| Supramolecule | Name: hEAAT3 trimer / type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1 |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
-Supramolecule #2: Out ward facing hEAAT3 protomer
| Supramolecule | Name: Out ward facing hEAAT3 protomer / type: complex / ID: 2 / Parent: 1 / Macromolecule list: #1 |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
-Macromolecule #1: Excitatory amino acid transporter 3
| Macromolecule | Name: Excitatory amino acid transporter 3 / type: protein_or_peptide / ID: 1 Details: The Glycine and Proline at the N terminal are the residues left after PreScission Protease treatment Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 57.275168 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: GPMGKPARKG CEWKRFLKNN WVLLSTVAAV VLGITTGVLV REHSNLSTLE KFYFAFPGEI LMRMLKLIIL PLIISSMITG VAALDSNVS GKIGLRAVVY YFCTTLIAVI LGIVLVVSIK PGVTQKVGEI ARTGSTPEVS TVDAMLDLIR NMFPENLVQA C FQQYKTKR ...String: GPMGKPARKG CEWKRFLKNN WVLLSTVAAV VLGITTGVLV REHSNLSTLE KFYFAFPGEI LMRMLKLIIL PLIISSMITG VAALDSNVS GKIGLRAVVY YFCTTLIAVI LGIVLVVSIK PGVTQKVGEI ARTGSTPEVS TVDAMLDLIR NMFPENLVQA C FQQYKTKR EEVKPPSDPE MTMTEESFTA VMTTAISKTK TKEYKIVGMY SDGINVLGLI VFCLVFGLVI GKMGEKGQIL VD FFNALSD ATMKIVQIIM CYMPLGILFL IAGKIIEVED WEIFRKLGLY MATVLTGLAI HSIVILPLIY FIVVRKNPFR FAM GMAQAL LTALMISSSS ATLPVTFRCA EENNQVDKRI TRFVLPVGAT INMDGTALYE AVAAVFIAQL NDLDLGIGQI ITIS ITATS ASIGAAGVPQ AGLVTMVIVL SAVGLPAEDV TLIIAVDWLL DRFRTMVNVL GDAFGTGIVE KLSKKELEQM DVSSE VNIV NPFALESTIL DNEDSDTKKS YVNGGFAVDK SDTISFTQTS QF UniProtKB: Excitatory amino acid transporter 3 |
-Macromolecule #2: ASPARTIC ACID
| Macromolecule | Name: ASPARTIC ACID / type: ligand / ID: 2 / Number of copies: 1 / Formula: ASP |
|---|---|
| Molecular weight | Theoretical: 133.103 Da |
| Chemical component information |  ChemComp-ASP: |
-Macromolecule #3: SODIUM ION
| Macromolecule | Name: SODIUM ION / type: ligand / ID: 3 / Number of copies: 3 |
|---|---|
| Molecular weight | Theoretical: 22.99 Da |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 6 mg/mL | ||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Buffer | pH: 8 Component:
| ||||||||||||||||||
| Grid | Model: Quantifoil R1.2/1.3 / Material: COPPER / Mesh: 400 / Support film - Material: CARBON / Support film - topology: HOLEY | ||||||||||||||||||
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 277 K / Instrument: FEI VITROBOT MARK IV / Details: blot 3s. | ||||||||||||||||||
| Details | This sample was mono disperse |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 BIOQUANTUM (6k x 4k) / Average electron dose: 60.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | C2 aperture diameter: 100.0 µm / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: 1.5 µm / Nominal defocus min: 0.5 µm / Nominal magnification: 105000 |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller




















 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)