[English] 日本語
 Yorodumi
Yorodumi- EMDB-21589: Cryo-EM structure of human Pannexin 1 channel with its C-terminal... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-21589 | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cryo-EM structure of human Pannexin 1 channel with its C-terminal tail cleaved by caspase-7 | ||||||||||||
 Map data Map data | Unsharpened map of %u0394CTT-hsPANX1 | ||||||||||||
 Sample Sample |
| ||||||||||||
 Keywords Keywords | ion channel / TRANSPORT PROTEIN | ||||||||||||
| Function / homology |  Function and homology information Function and homology informationATP transmembrane transporter activity / ATP transport / leak channel activity / Electric Transmission Across Gap Junctions / positive regulation of interleukin-1 alpha production / monoatomic anion transmembrane transport / wide pore channel activity / bleb / monoatomic anion channel activity / gap junction ...ATP transmembrane transporter activity / ATP transport / leak channel activity / Electric Transmission Across Gap Junctions / positive regulation of interleukin-1 alpha production / monoatomic anion transmembrane transport / wide pore channel activity / bleb / monoatomic anion channel activity / gap junction / gap junction channel activity / positive regulation of macrophage cytokine production / Mechanical load activates signaling by PIEZO1 and integrins in osteocytes / oogenesis / response to ATP / The NLRP3 inflammasome / monoatomic cation transport / response to ischemia / positive regulation of interleukin-1 beta production / calcium channel activity / actin filament binding / calcium ion transport / cell-cell signaling / protease binding / scaffold protein binding / High laminar flow shear stress activates signaling by PIEZO1 and PECAM1:CDH5:KDR in endothelial cells / transmembrane transporter binding / signaling receptor binding / endoplasmic reticulum membrane / structural molecule activity / endoplasmic reticulum / protein-containing complex / membrane / identical protein binding / plasma membrane Similarity search - Function | ||||||||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | ||||||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 2.97 Å | ||||||||||||
 Authors Authors | Lu W / Du J | ||||||||||||
| Funding support |  United States, 3 items United States, 3 items
| ||||||||||||
 Citation Citation |  Journal: Nature / Year: 2020 Journal: Nature / Year: 2020Title: Structures of human pannexin 1 reveal ion pathways and mechanism of gating. Authors: Zheng Ruan / Ian J Orozco / Juan Du / Wei Lü /  Abstract: Pannexin 1 (PANX1) is an ATP-permeable channel with critical roles in a variety of physiological functions such as blood pressure regulation, apoptotic cell clearance and human oocyte development. ...Pannexin 1 (PANX1) is an ATP-permeable channel with critical roles in a variety of physiological functions such as blood pressure regulation, apoptotic cell clearance and human oocyte development. Here we present several structures of human PANX1 in a heptameric assembly at resolutions of up to 2.8 angström, including an apo state, a caspase-7-cleaved state and a carbenoxolone-bound state. We reveal a gating mechanism that involves two ion-conducting pathways. Under normal cellular conditions, the intracellular entry of the wide main pore is physically plugged by the C-terminal tail. Small anions are conducted through narrow tunnels in the intracellular domain. These tunnels connect to the main pore and are gated by a long linker between the N-terminal helix and the first transmembrane helix. During apoptosis, the C-terminal tail is cleaved by caspase, allowing the release of ATP through the main pore. We identified a carbenoxolone-binding site embraced by W74 in the extracellular entrance and a role for carbenoxolone as a channel blocker. We identified a gap-junction-like structure using a glycosylation-deficient mutant, N255A. Our studies provide a solid foundation for understanding the molecular mechanisms underlying the channel gating and inhibition of PANX1 and related large-pore channels. | ||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_21589.map.gz emd_21589.map.gz | 40.1 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-21589-v30.xml emd-21589-v30.xml emd-21589.xml emd-21589.xml | 17.8 KB 17.8 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_21589.png emd_21589.png | 169.5 KB | ||
| Filedesc metadata |  emd-21589.cif.gz emd-21589.cif.gz | 5.9 KB | ||
| Others |  emd_21589_additional.map.gz emd_21589_additional.map.gz | 7 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-21589 http://ftp.pdbj.org/pub/emdb/structures/EMD-21589 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-21589 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-21589 | HTTPS FTP |
-Related structure data
| Related structure data |  6wbgMC  6wbfC  6wbiC  6wbkC  6wblC  6wbmC  6wbnC C: citing same article ( M: atomic model generated by this map |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_21589.map.gz / Format: CCP4 / Size: 52.7 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_21589.map.gz / Format: CCP4 / Size: 52.7 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Unsharpened map of %u0394CTT-hsPANX1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.026 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
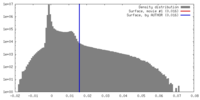
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
-Additional map: Sharpened map of %u0394CTT-hsPANX1
| File | emd_21589_additional.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Sharpened map of %u0394CTT-hsPANX1 | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Wild type human Pannexin 1 channel
| Entire | Name: Wild type human Pannexin 1 channel |
|---|---|
| Components |
|
-Supramolecule #1: Wild type human Pannexin 1 channel
| Supramolecule | Name: Wild type human Pannexin 1 channel / type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1 |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
-Macromolecule #1: Pannexin-1
| Macromolecule | Name: Pannexin-1 / type: protein_or_peptide / ID: 1 / Number of copies: 7 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 42.247441 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: MAIAQLATEY VFSDFLLKEP TEPKFKGLRL ELAVDKMVTC IAVGLPLLLI SLAFAQEISI GTQISCFSPS SFSWRQAAFV DSYCWAAVQ QKNSLQSESG NLPLWLHKFF PYILLLFAIL LYLPPLFWRF AAAPHICSDL KFIMEELDKV YNRAIKAAKS A RDLDMRDG ...String: MAIAQLATEY VFSDFLLKEP TEPKFKGLRL ELAVDKMVTC IAVGLPLLLI SLAFAQEISI GTQISCFSPS SFSWRQAAFV DSYCWAAVQ QKNSLQSESG NLPLWLHKFF PYILLLFAIL LYLPPLFWRF AAAPHICSDL KFIMEELDKV YNRAIKAAKS A RDLDMRDG ACSVPGVTEN LGQSLWEVSE SHFKYPIVEQ YLKTKKNSNN LIIKYISCRL LTLIIILLAC IYLGYYFSLS SL SDEFVCS IKSGILRNDS TVPDQFQCKL IAVGIFQLLS VINLVVYVLL APVVVYTLFV PFRQKTDVLK VYEILPTFDV LHF KSEGYN DLSLYNLFLE ENISEVKSYK CLKVLENIKS SGQGIDPMLL LTNLGMI UniProtKB: Pannexin-1 |
-Macromolecule #2: PHOSPHATIDYLETHANOLAMINE
| Macromolecule | Name: PHOSPHATIDYLETHANOLAMINE / type: ligand / ID: 2 / Number of copies: 14 / Formula: PTY |
|---|---|
| Molecular weight | Theoretical: 734.039 Da |
| Chemical component information |  ChemComp-PTY: |
-Macromolecule #3: 1,2-Distearoyl-sn-glycerophosphoethanolamine
| Macromolecule | Name: 1,2-Distearoyl-sn-glycerophosphoethanolamine / type: ligand / ID: 3 / Number of copies: 7 / Formula: 3PE |
|---|---|
| Molecular weight | Theoretical: 748.065 Da |
| Chemical component information |  ChemComp-3PE: |
-Macromolecule #4: DIACYL GLYCEROL
| Macromolecule | Name: DIACYL GLYCEROL / type: ligand / ID: 4 / Number of copies: 7 / Formula: DGA |
|---|---|
| Molecular weight | Theoretical: 625.018 Da |
| Chemical component information |  ChemComp-DGA: |
-Macromolecule #5: CHOLESTEROL
| Macromolecule | Name: CHOLESTEROL / type: ligand / ID: 5 / Number of copies: 14 / Formula: CLR |
|---|---|
| Molecular weight | Theoretical: 386.654 Da |
| Chemical component information |  ChemComp-CLR: |
-Macromolecule #6: 2-acetamido-2-deoxy-beta-D-glucopyranose
| Macromolecule | Name: 2-acetamido-2-deoxy-beta-D-glucopyranose / type: ligand / ID: 6 / Number of copies: 7 / Formula: NAG |
|---|---|
| Molecular weight | Theoretical: 221.208 Da |
| Chemical component information |  ChemComp-NAG: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 7 mg/mL |
|---|---|
| Buffer | pH: 8 |
| Grid | Details: unspecified |
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 291.15 K / Instrument: FEI VITROBOT MARK III |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Detector mode: SUPER-RESOLUTION / Average exposure time: 8.0 sec. / Average electron dose: 49.6 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
+ Image processing
Image processing
-Atomic model buiding 1
| Refinement | Space: REAL / Protocol: AB INITIO MODEL |
|---|---|
| Output model |  PDB-6wbg: |
 Movie
Movie Controller
Controller
























 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)