+ データを開く
データを開く
- 基本情報
基本情報
| 登録情報 | データベース: EMDB / ID: EMD-20221 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| タイトル | nhTMEM16 L302A +Ca2+ in nanodiscs | |||||||||
 マップデータ マップデータ | Final EM map for nhTMEM16 L302A/nanodisc complex with C2 symmetry. Used for model buidling. | |||||||||
 試料 試料 |
| |||||||||
 キーワード キーワード | TMEM16 / scramblase / lipid transport / anoactamin | |||||||||
| 機能・相同性 |  機能・相同性情報 機能・相同性情報cortical endoplasmic reticulum / chloride channel activity / membrane / metal ion binding / identical protein binding 類似検索 - 分子機能 | |||||||||
| 生物種 |  Nectria haematococca mpVI (菌類) / Nectria haematococca mpVI (菌類) /  Nectria haematococca (strain 77-13-4 / ATCC MYA-4622 / FGSC 9596 / MPVI) (菌類) Nectria haematococca (strain 77-13-4 / ATCC MYA-4622 / FGSC 9596 / MPVI) (菌類) | |||||||||
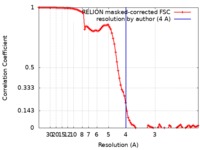
| 手法 | 単粒子再構成法 / クライオ電子顕微鏡法 / 解像度: 4.0 Å | |||||||||
 データ登録者 データ登録者 | Falzone M / Lee BC | |||||||||
| 資金援助 |  米国, 1件 米国, 1件
| |||||||||
 引用 引用 |  ジャーナル: Nat Commun / 年: 2019 ジャーナル: Nat Commun / 年: 2019タイトル: Dynamic modulation of the lipid translocation groove generates a conductive ion channel in Ca-bound nhTMEM16. 著者: George Khelashvili / Maria E Falzone / Xiaolu Cheng / Byoung-Cheol Lee / Alessio Accardi / Harel Weinstein /   要旨: Both lipid and ion translocation by Ca-regulated TMEM16 transmembrane proteins utilizes a membrane-exposed hydrophilic groove. Several conformations of the groove are observed in TMEM16 protein ...Both lipid and ion translocation by Ca-regulated TMEM16 transmembrane proteins utilizes a membrane-exposed hydrophilic groove. Several conformations of the groove are observed in TMEM16 protein structures, but how these conformations form, and what functions they support, remains unknown. From analyses of atomistic molecular dynamics simulations of Ca-bound nhTMEM16 we find that the mechanism of a conformational transition of the groove from membrane-exposed to occluded from the membrane involves the repositioning of transmembrane helix 4 (TM4) following its disengagement from a TM3/TM4 interaction interface. Residue L302 is a key element in the hydrophobic TM3/TM4 interaction patch that braces the open-groove conformation, which should be changed by an L302A mutation. The structure of the L302A mutant determined by cryogenic electron microscopy (cryo-EM) reveals a partially closed groove that could translocate ions, but not lipids. This is corroborated with functional assays showing severely impaired lipid scrambling, but robust channel activity by L302A. | |||||||||
| 履歴 |
|
- 構造の表示
構造の表示
| ムービー |
 ムービービューア ムービービューア |
|---|---|
| 構造ビューア | EMマップ:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| 添付画像 |
- ダウンロードとリンク
ダウンロードとリンク
-EMDBアーカイブ
| マップデータ |  emd_20221.map.gz emd_20221.map.gz | 58 MB |  EMDBマップデータ形式 EMDBマップデータ形式 | |
|---|---|---|---|---|
| ヘッダ (付随情報) |  emd-20221-v30.xml emd-20221-v30.xml emd-20221.xml emd-20221.xml | 15.5 KB 15.5 KB | 表示 表示 |  EMDBヘッダ EMDBヘッダ |
| FSC (解像度算出) |  emd_20221_fsc.xml emd_20221_fsc.xml | 9.2 KB | 表示 |  FSCデータファイル FSCデータファイル |
| 画像 |  emd_20221.png emd_20221.png | 224.1 KB | ||
| Filedesc metadata |  emd-20221.cif.gz emd-20221.cif.gz | 5.9 KB | ||
| その他 |  emd_20221_additional_1.map.gz emd_20221_additional_1.map.gz emd_20221_additional_2.map.gz emd_20221_additional_2.map.gz | 59.8 MB 55.3 MB | ||
| アーカイブディレクトリ |  http://ftp.pdbj.org/pub/emdb/structures/EMD-20221 http://ftp.pdbj.org/pub/emdb/structures/EMD-20221 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-20221 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-20221 | HTTPS FTP |
-関連構造データ
- リンク
リンク
| EMDBのページ |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- マップ
マップ
| ファイル |  ダウンロード / ファイル: emd_20221.map.gz / 形式: CCP4 / 大きさ: 64 MB / タイプ: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) ダウンロード / ファイル: emd_20221.map.gz / 形式: CCP4 / 大きさ: 64 MB / タイプ: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 注釈 | Final EM map for nhTMEM16 L302A/nanodisc complex with C2 symmetry. Used for model buidling. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
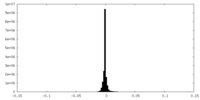
| 投影像・断面図 | 画像のコントロール
画像は Spider により作成 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| ボクセルのサイズ | X=Y=Z: 1.0961 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| 密度 |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| 対称性 | 空間群: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| 詳細 | EMDB XML:
CCP4マップ ヘッダ情報:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-添付データ
-追加マップ: Final EM map for nhTMEM16 L302A/nanodisc complex with...
| ファイル | emd_20221_additional_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 注釈 | Final EM map for nhTMEM16 L302A/nanodisc complex with C1 symmetry. Used to analyze effects of nhTMEM16 | ||||||||||||
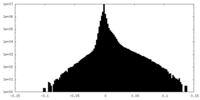
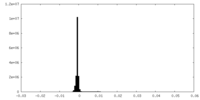
| 投影像・断面図 |
| ||||||||||||
| 密度ヒストグラム |
-追加マップ: Unfiltered map from refinement of the nhTMEM16 L302A/nanodisc....
| ファイル | emd_20221_additional_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 注釈 | Unfiltered map from refinement of the nhTMEM16 L302A/nanodisc. Used to assist model building. | ||||||||||||
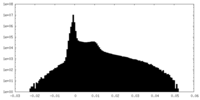
| 投影像・断面図 |
| ||||||||||||
| 密度ヒストグラム |
- 試料の構成要素
試料の構成要素
-全体 : nhTMEM16 L302A reconstituted in nanodiscs in the presence of Ca2+
| 全体 | 名称: nhTMEM16 L302A reconstituted in nanodiscs in the presence of Ca2+ |
|---|---|
| 要素 |
|
-超分子 #1: nhTMEM16 L302A reconstituted in nanodiscs in the presence of Ca2+
| 超分子 | 名称: nhTMEM16 L302A reconstituted in nanodiscs in the presence of Ca2+ タイプ: complex / ID: 1 / 親要素: 0 / 含まれる分子: #1 |
|---|---|
| 由来(天然) | 生物種:  Nectria haematococca mpVI (菌類) Nectria haematococca mpVI (菌類) |
| 分子量 | 理論値: 166 kDa/nm |
-分子 #1: nhTMEM16
| 分子 | 名称: nhTMEM16 / タイプ: protein_or_peptide / ID: 1 / コピー数: 2 / 光学異性体: LEVO |
|---|---|
| 由来(天然) | 生物種:  Nectria haematococca (strain 77-13-4 / ATCC MYA-4622 / FGSC 9596 / MPVI) (菌類) Nectria haematococca (strain 77-13-4 / ATCC MYA-4622 / FGSC 9596 / MPVI) (菌類)株: 77-13-4 / ATCC MYA-4622 / FGSC 9596 / MPVI |
| 分子量 | 理論値: 83.15793 KDa |
| 組換発現 | 生物種:  |
| 配列 | 文字列: MSNLKDFSQP GSGQESNFGV DFVIHYKVPA AERDEAEAGF VQLIRALTTV GLATEVRHGE NESLLVFVKV ASPDLFAKQV YRARLGDWL HGVRVSAPHN DIAQALQDEP VVEAERLRLI YLMITKPHNE GGAGVTPTNA KWKHVESIFP LHSHSFNKEW I KKWSSKYT ...文字列: MSNLKDFSQP GSGQESNFGV DFVIHYKVPA AERDEAEAGF VQLIRALTTV GLATEVRHGE NESLLVFVKV ASPDLFAKQV YRARLGDWL HGVRVSAPHN DIAQALQDEP VVEAERLRLI YLMITKPHNE GGAGVTPTNA KWKHVESIFP LHSHSFNKEW I KKWSSKYT LEQTDIDNIR DKFGESVAFY FAFLRSYFRF LVIPSAFGFG AWLLLGQFSY LYALLCGLWS VVFFEYWKKQ EV DLAVQWG VRGVSSIQQS RPEFEWEHEA EDPITGEPVK VYPPMKRVKT QLLQIPFALA CVVAAGALIV TCNSLEVFIN EVY SGPGKQ YLGFLPTIFL VIGTPTISGV LMGAAEKLNA MENYATVDAH DAALIQKQFV LNFMTSYMAL FFTAFVYIPF GHIL HPFLN FWRATAQTLT FSEKELPTRE FQINPARISN QMFYFTVTAQ IVNFATEVVV PYIKQQAFQK AKQLKSGSKV QEDHE EEAE FLQRVREECT LEEYDVSGDY REMVMQFGYV AMFSVAWPLA ACCFLVNNWV ELRSDALKIA ISSRRPIPWR TDSIGP WLT ALSFLSWLGS ITSSAIVYLC SNSKNGTQGE ASPLKAWGLL LSILFAEHFY LVVQLAVRFV LSKLDSPGLQ KERKERF QT KKRLLQENLG QDAAEEAAAP GIEHSEKITR EALEEEARQA SIRGHGTPEE MFWQRQRGMQ ETIEIGRRMI EQQLAAGK N GKKSAPAVPS EKAS UniProtKB: Plasma membrane channel protein |
-分子 #2: CALCIUM ION
| 分子 | 名称: CALCIUM ION / タイプ: ligand / ID: 2 / コピー数: 4 / 式: CA |
|---|---|
| 分子量 | 理論値: 40.078 Da |
-実験情報
-構造解析
| 手法 | クライオ電子顕微鏡法 |
|---|---|
 解析 解析 | 単粒子再構成法 |
| 試料の集合状態 | particle |
- 試料調製
試料調製
| 濃度 | 6.5 mg/mL |
|---|---|
| 緩衝液 | pH: 8 |
| 凍結 | 凍結剤: ETHANE / チャンバー内湿度: 100 % / チャンバー内温度: 288 K / 装置: FEI VITROBOT MARK IV |
| 詳細 | nhTMEM16 L302A reconstituted in nanodiscs in the absence of Ca2+ |
- 電子顕微鏡法
電子顕微鏡法
| 顕微鏡 | FEI TITAN KRIOS |
|---|---|
| 撮影 | フィルム・検出器のモデル: GATAN K2 SUMMIT (4k x 4k) 検出モード: COUNTING / 撮影したグリッド数: 1 / 平均電子線量: 70.7 e/Å2 |
| 電子線 | 加速電圧: 300 kV / 電子線源:  FIELD EMISSION GUN FIELD EMISSION GUN |
| 電子光学系 | 照射モード: OTHER / 撮影モード: BRIGHT FIELD |
| 実験機器 |  モデル: Titan Krios / 画像提供: FEI Company |
 ムービー
ムービー コントローラー
コントローラー















 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)