+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | human Connexin 26 dodecamer at 55mm Hg PCO2, pH7.4 | |||||||||
 Map data Map data | masked D6 .mrc file | |||||||||
 Sample Sample |
| |||||||||
| Function / homology |  Function and homology information Function and homology informationTransport of connexons to the plasma membrane / response to human chorionic gonadotropin / gap junction-mediated intercellular transport / epididymis development / gap junction channel activity involved in cell communication by electrical coupling / Oligomerization of connexins into connexons / Transport of connexins along the secretory pathway /  gap junction assembly / gap junction assembly /  connexin complex / connexin complex /  Gap junction assembly ...Transport of connexons to the plasma membrane / response to human chorionic gonadotropin / gap junction-mediated intercellular transport / epididymis development / gap junction channel activity involved in cell communication by electrical coupling / Oligomerization of connexins into connexons / Transport of connexins along the secretory pathway / Gap junction assembly ...Transport of connexons to the plasma membrane / response to human chorionic gonadotropin / gap junction-mediated intercellular transport / epididymis development / gap junction channel activity involved in cell communication by electrical coupling / Oligomerization of connexins into connexons / Transport of connexins along the secretory pathway /  gap junction assembly / gap junction assembly /  connexin complex / connexin complex /  Gap junction assembly / astrocyte projection / gap junction channel activity / Gap junction assembly / astrocyte projection / gap junction channel activity /  gap junction / cellular response to glucagon stimulus / inner ear development / gap junction / cellular response to glucagon stimulus / inner ear development /  decidualization / decidualization /  endoplasmic reticulum-Golgi intermediate compartment / lateral plasma membrane / response to retinoic acid / cellular response to dexamethasone stimulus / response to progesterone / response to ischemia / sensory perception of sound / transmembrane transport / response to estradiol / cell-cell signaling / cellular response to oxidative stress / endoplasmic reticulum-Golgi intermediate compartment / lateral plasma membrane / response to retinoic acid / cellular response to dexamethasone stimulus / response to progesterone / response to ischemia / sensory perception of sound / transmembrane transport / response to estradiol / cell-cell signaling / cellular response to oxidative stress /  cell body / response to lipopolysaccharide / cell body / response to lipopolysaccharide /  calcium ion binding / perinuclear region of cytoplasm / identical protein binding / calcium ion binding / perinuclear region of cytoplasm / identical protein binding /  plasma membrane plasma membraneSimilarity search - Function | |||||||||
| Biological species |   Homo sapiens (human) Homo sapiens (human) | |||||||||
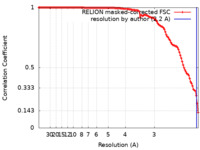
| Method |  single particle reconstruction / single particle reconstruction /  cryo EM / Resolution: 2.2 Å cryo EM / Resolution: 2.2 Å | |||||||||
 Authors Authors | Brotherton DH / Cameron AD / Savva CG / Ragan TJ | |||||||||
| Funding support |  United Kingdom, 2 items United Kingdom, 2 items
| |||||||||
 Citation Citation |  Journal: Structure / Year: 2022 Journal: Structure / Year: 2022Title: Conformational changes and CO-induced channel gating in connexin26. Authors: Deborah H Brotherton / Christos G Savva / Timothy J Ragan / Nicholas Dale / Alexander D Cameron /  Abstract: Connexins form large-pore channels that function either as dodecameric gap junctions or hexameric hemichannels to allow the regulated movement of small molecules and ions across cell membranes. ...Connexins form large-pore channels that function either as dodecameric gap junctions or hexameric hemichannels to allow the regulated movement of small molecules and ions across cell membranes. Opening or closing of the channels is controlled by a variety of stimuli, and dysregulation leads to multiple diseases. An increase in the partial pressure of carbon dioxide (PCO) has been shown to cause connexin26 (Cx26) gap junctions to close. Here, we use cryoelectron microscopy (cryo-EM) to determine the structure of human Cx26 gap junctions under increasing levels of PCO. We show a correlation between the level of PCO and the size of the aperture of the pore, governed by the N-terminal helices that line the pore. This indicates that CO alone is sufficient to cause conformational changes in the protein. Analysis of the conformational states shows that movements at the N terminus are linked to both subunit rotation and flexing of the transmembrane helices. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_13938.map.gz emd_13938.map.gz | 83.7 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-13938-v30.xml emd-13938-v30.xml emd-13938.xml emd-13938.xml | 20.3 KB 20.3 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_13938_fsc.xml emd_13938_fsc.xml | 11.9 KB | Display |  FSC data file FSC data file |
| Images |  emd_13938.png emd_13938.png | 91.4 KB | ||
| Others |  emd_13938_half_map_1.map.gz emd_13938_half_map_1.map.gz emd_13938_half_map_2.map.gz emd_13938_half_map_2.map.gz | 116.4 MB 116.4 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-13938 http://ftp.pdbj.org/pub/emdb/structures/EMD-13938 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-13938 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-13938 | HTTPS FTP |
-Related structure data
| Related structure data |  7qerMC  7qeoC  7qeqC  7qesC  7qetC  7qeuC  7qevC  7qewC  7qeyC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_13938.map.gz / Format: CCP4 / Size: 149.9 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_13938.map.gz / Format: CCP4 / Size: 149.9 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | masked D6 .mrc file | ||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.09 Å | ||||||||||||||||||||||||||||||||||||
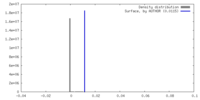
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Half map: human connexin 26 dodecamer at 55mmHg PCO2 half...
| File | emd_13938_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | human connexin 26 dodecamer at 55mmHg PCO2 half map 2 from full dataset | ||||||||||||
| Projections & Slices |
| ||||||||||||
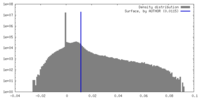
| Density Histograms |
-Half map: human connexin 26 dodecamer at 55mmHg PCO2 half...
| File | emd_13938_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | human connexin 26 dodecamer at 55mmHg PCO2 half map 1 from full dataset | ||||||||||||
| Projections & Slices |
| ||||||||||||
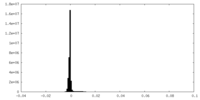
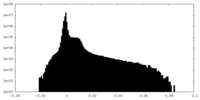
| Density Histograms |
- Sample components
Sample components
-Entire : dodecameric assembly of human connexin 26
| Entire | Name: dodecameric assembly of human connexin 26 |
|---|---|
| Components |
|
-Supramolecule #1: dodecameric assembly of human connexin 26
| Supramolecule | Name: dodecameric assembly of human connexin 26 / type: organelle_or_cellular_component / ID: 1 / Parent: 0 / Macromolecule list: #1 |
|---|---|
| Source (natural) | Organism:   Homo sapiens (human) Homo sapiens (human) |
| Recombinant expression | Organism:   Spodoptera frugiperda (fall armyworm) Spodoptera frugiperda (fall armyworm) |
-Macromolecule #1: Gap junction beta-2 protein
| Macromolecule | Name: Gap junction beta-2 protein / type: protein_or_peptide / ID: 1 / Number of copies: 12 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:   Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 26.713674 KDa |
| Recombinant expression | Organism:   Spodoptera frugiperda (fall armyworm) Spodoptera frugiperda (fall armyworm) |
| Sequence | String: MDWGTLQTIL GGVNKHSTSI GKIWLTVLFI FRIMILVVAA KEVWGDEQAD FVCNTLQPGC KNVCYDHYFP ISHIRLWALQ LIFVSTPAL LVAMHVAYRR HEKKRKFIKG EIKSEFKDIE EIKTQKVRIE GSLWWTYTSS IFFRVIFEAA FMYVFYVMYD G FSMQRLVK ...String: MDWGTLQTIL GGVNKHSTSI GKIWLTVLFI FRIMILVVAA KEVWGDEQAD FVCNTLQPGC KNVCYDHYFP ISHIRLWALQ LIFVSTPAL LVAMHVAYRR HEKKRKFIKG EIKSEFKDIE EIKTQKVRIE GSLWWTYTSS IFFRVIFEAA FMYVFYVMYD G FSMQRLVK CNAWPCPNTV DCFVSRPTEK TVFTVFMIAV SGICILLNVT ELCYLLIRYC SGKSKKPVLV PR |
-Macromolecule #2: DODECYL-BETA-D-MALTOSIDE
| Macromolecule | Name: DODECYL-BETA-D-MALTOSIDE / type: ligand / ID: 2 / Number of copies: 48 / Formula: LMT |
|---|---|
| Molecular weight | Theoretical: 510.615 Da |
| Chemical component information |  ChemComp-LMT: |
-Macromolecule #3: PHOSPHATIDYLETHANOLAMINE
| Macromolecule | Name: PHOSPHATIDYLETHANOLAMINE / type: ligand / ID: 3 / Number of copies: 12 / Formula: PTY |
|---|---|
| Molecular weight | Theoretical: 734.039 Da |
| Chemical component information |  ChemComp-PTY: |
-Macromolecule #4: water
| Macromolecule | Name: water / type: ligand / ID: 4 / Number of copies: 305 / Formula: HOH |
|---|---|
| Molecular weight | Theoretical: 18.015 Da |
| Chemical component information |  ChemComp-HOH: |
-Experimental details
-Structure determination
| Method |  cryo EM cryo EM |
|---|---|
 Processing Processing |  single particle reconstruction single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 3.5 mg/mL | ||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Buffer | pH: 7.4 Component:
Details: The buffer, except DDM and DTT, was prepared fresh from 10x stock on day of use. The basal buffer was filtered and de-gassed, and DDM and DTT added. The buffer was pH corrected at point of ...Details: The buffer, except DDM and DTT, was prepared fresh from 10x stock on day of use. The basal buffer was filtered and de-gassed, and DDM and DTT added. The buffer was pH corrected at point of use to 7.4 using 10% CO2 in 90% N2. | ||||||||||||||||||||||||||||||
| Grid | Model: UltrAuFoil R1.2/1.3 / Material: GOLD / Mesh: 300 / Support film - Material: GOLD / Support film - topology: HOLEY / Support film - Film thickness: 50.0 nm / Pretreatment - Type: GLOW DISCHARGE / Pretreatment - Atmosphere: AIR | ||||||||||||||||||||||||||||||
| Vitrification | Cryogen name: ETHANE-PROPANE / Chamber humidity: 95 % / Chamber temperature: 277 K / Instrument: LEICA EM GP Details: 3 microlitres protein applied to grid, blot time 6 seconds, in 10% CO2/90%N2 atmosphere. | ||||||||||||||||||||||||||||||
| Details | This sample monodisperse |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | C2 aperture diameter: 50.0 µm / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD Bright-field microscopy / Cs: 2.7 mm / Nominal defocus max: 2.0 µm / Nominal defocus min: 1.0 µm / Nominal magnification: 81000 Bright-field microscopy / Cs: 2.7 mm / Nominal defocus max: 2.0 µm / Nominal defocus min: 1.0 µm / Nominal magnification: 81000 |
| Specialist optics | Energy filter - Name: GIF Bioquantum / Energy filter - Slit width: 20 eV |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER |
| Image recording | Film or detector model: GATAN K3 (6k x 4k) / Number grids imaged: 1 / Number real images: 4003 / Average exposure time: 5.0 sec. / Average electron dose: 45.0 e/Å2 |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller











 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)