[English] 日本語
 Yorodumi
Yorodumi- EMDB-13603: Asymmetric single particle reconstruction of the Haliangium ochra... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-13603 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Asymmetric single particle reconstruction of the Haliangium ochraceum encapsulin encapsulated ferritin complex calculated using Cryosparc | |||||||||
 Map data Map data | cryosparc map | |||||||||
 Sample Sample |
| |||||||||
| Function / homology |  Function and homology information Function and homology informationencapsulin nanocompartment / ferroxidase / ferroxidase activity / iron ion transport / intracellular iron ion homeostasis / metal ion binding Similarity search - Function | |||||||||
| Biological species |  Haliangium ochraceum (bacteria) Haliangium ochraceum (bacteria) | |||||||||
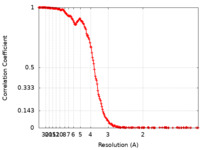
| Method | single particle reconstruction / cryo EM / Resolution: 3.24 Å | |||||||||
 Authors Authors | Marles-Wright J / Basle A / Ross J / Clarke DJ | |||||||||
| Funding support |  United Kingdom, 1 items United Kingdom, 1 items
| |||||||||
 Citation Citation |  Journal: Sci Adv / Year: 2022 Journal: Sci Adv / Year: 2022Title: Pore dynamics and asymmetric cargo loading in an encapsulin nanocompartment. Authors: Jennifer Ross / Zak McIver / Thomas Lambert / Cecilia Piergentili / Jasmine Emma Bird / Kelly J Gallagher / Faye L Cruickshank / Patrick James / Efrain Zarazúa-Arvizu / Louise E Horsfall / ...Authors: Jennifer Ross / Zak McIver / Thomas Lambert / Cecilia Piergentili / Jasmine Emma Bird / Kelly J Gallagher / Faye L Cruickshank / Patrick James / Efrain Zarazúa-Arvizu / Louise E Horsfall / Kevin J Waldron / Marcus D Wilson / C Logan Mackay / Arnaud Baslé / David J Clarke / Jon Marles-Wright /  Abstract: Encapsulins are protein nanocompartments that house various cargo enzymes, including a family of decameric ferritin-like proteins. Here, we study a recombinant encapsulin:encapsulated ferritin ...Encapsulins are protein nanocompartments that house various cargo enzymes, including a family of decameric ferritin-like proteins. Here, we study a recombinant encapsulin:encapsulated ferritin complex using cryo-electron microscopy and hydrogen/deuterium exchange mass spectrometry to gain insight into the structural relationship between the encapsulin shell and its protein cargo. An asymmetric single-particle reconstruction reveals four encapsulated ferritin decamers in a tetrahedral arrangement within the encapsulin nanocompartment. This leads to a symmetry mismatch between the protein cargo and the icosahedral encapsulin shell. The encapsulated ferritin decamers are offset from the interior face of the encapsulin shell. Using hydrogen/deuterium exchange mass spectrometry, we observed the dynamic behavior of the major fivefold pore in the encapsulin shell and show the pore opening via the movement of the encapsulin A-domain. These data will accelerate efforts to engineer the encapsulation of heterologous cargo proteins and to alter the permeability of the encapsulin shell via pore modifications. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_13603.map.gz emd_13603.map.gz | 364.1 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-13603-v30.xml emd-13603-v30.xml emd-13603.xml emd-13603.xml | 22.2 KB 22.2 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_13603_fsc.xml emd_13603_fsc.xml | 22 KB | Display |  FSC data file FSC data file |
| Images |  emd_13603.png emd_13603.png | 75.4 KB | ||
| Masks |  emd_13603_msk_1.map emd_13603_msk_1.map | 421.9 MB |  Mask map Mask map | |
| Others |  emd_13603_additional_1.map.gz emd_13603_additional_1.map.gz emd_13603_half_map_1.map.gz emd_13603_half_map_1.map.gz emd_13603_half_map_2.map.gz emd_13603_half_map_2.map.gz | 394.6 MB 390.9 MB 390.9 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-13603 http://ftp.pdbj.org/pub/emdb/structures/EMD-13603 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-13603 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-13603 | HTTPS FTP |
-Validation report
| Summary document |  emd_13603_validation.pdf.gz emd_13603_validation.pdf.gz | 682 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_13603_full_validation.pdf.gz emd_13603_full_validation.pdf.gz | 681.6 KB | Display | |
| Data in XML |  emd_13603_validation.xml.gz emd_13603_validation.xml.gz | 25.4 KB | Display | |
| Data in CIF |  emd_13603_validation.cif.gz emd_13603_validation.cif.gz | 33 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-13603 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-13603 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-13603 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-13603 | HTTPS FTP |
-Related structure data
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_13603.map.gz / Format: CCP4 / Size: 421.9 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_13603.map.gz / Format: CCP4 / Size: 421.9 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | cryosparc map | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.652 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
-Mask #1
| File |  emd_13603_msk_1.map emd_13603_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
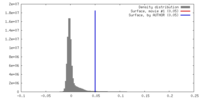
| Density Histograms |
-Additional map: sharpened map
| File | emd_13603_additional_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | sharpened map | ||||||||||||
| Projections & Slices |
| ||||||||||||
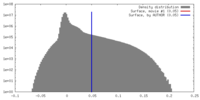
| Density Histograms |
-Half map: Half map A
| File | emd_13603_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Half map A | ||||||||||||
| Projections & Slices |
| ||||||||||||
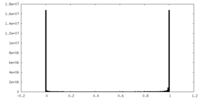
| Density Histograms |
-Half map: Half map B
| File | emd_13603_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Half map B | ||||||||||||
| Projections & Slices |
| ||||||||||||
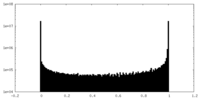
| Density Histograms |
- Sample components
Sample components
-Entire : Ternary complex of Haliangium ochraceum encapsulin and encapsulat...
| Entire | Name: Ternary complex of Haliangium ochraceum encapsulin and encapsulated ferritin proteins |
|---|---|
| Components |
|
-Supramolecule #1: Ternary complex of Haliangium ochraceum encapsulin and encapsulat...
| Supramolecule | Name: Ternary complex of Haliangium ochraceum encapsulin and encapsulated ferritin proteins type: complex / ID: 1 / Parent: 0 / Macromolecule list: all / Details: Complex produced by co-expression in E. coli |
|---|---|
| Source (natural) | Organism:  Haliangium ochraceum (bacteria) Haliangium ochraceum (bacteria) |
| Recombinant expression | Organism:  |
| Molecular weight | Theoretical: 1.8 MDa |
-Macromolecule #1: Haliangium ochraceum encapsulin
| Macromolecule | Name: Haliangium ochraceum encapsulin / type: protein_or_peptide / ID: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Haliangium ochraceum (bacteria) Haliangium ochraceum (bacteria) |
| Recombinant expression | Organism:  |
| Sequence | String: MDLLKRHLAP IVPDAWSAID EEAKEIFQGH LAGRKLVDFR GPFGWEYAAV NTGELRPIDD TPEDVDMKL RQVQPLAEVR VPFTLDVTEL DSVARGATNP DLDDVARAAE RMVEAEDSAI F HGWAQAGI KGIVDSTPHE ALAVASVSDF PRAVLSAADT LRKAGVTGPY ...String: MDLLKRHLAP IVPDAWSAID EEAKEIFQGH LAGRKLVDFR GPFGWEYAAV NTGELRPIDD TPEDVDMKL RQVQPLAEVR VPFTLDVTEL DSVARGATNP DLDDVARAAE RMVEAEDSAI F HGWAQAGI KGIVDSTPHE ALAVASVSDF PRAVLSAADT LRKAGVTGPY ALVLGPKAYD DL FAATQDG YPVAKQVQRL VVDGPLVRAN ALAGALVMSM RGGDYELTVG QDLSIGYAFH DRS KVELFV AESFTFRVLE PGAAVHLRYA |
-Macromolecule #2: Haliangium ochraceum encapsulated ferritin
| Macromolecule | Name: Haliangium ochraceum encapsulated ferritin / type: protein_or_peptide / ID: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Haliangium ochraceum (bacteria) Haliangium ochraceum (bacteria) |
| Recombinant expression | Organism:  |
| Sequence | String: MSSEQLHEPA ELLSEETKNM HRALVTLIEE LEAVDWYQQR ADACSEPGLH DVLIHNKNEE VEHAMMTLE WIRRRSPVFD AHMRTYLFTE RPILELEEED TGSSSSVAAS PTSAPSHGSL G IGSLRQEG KED |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 3 mg/mL | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Buffer | pH: 8 Component:
Details: Solutions were prepared with MilliQ water and filtered using a 0.22 um filter. | |||||||||
| Grid | Model: Quantifoil / Material: GOLD / Mesh: 200 / Support film - Material: CARBON / Support film - topology: HOLEY / Pretreatment - Type: GLOW DISCHARGE / Pretreatment - Atmosphere: AIR | |||||||||
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 281 K / Instrument: FEI VITROBOT MARK IV Details: blot force -5, wait time 10 seconds and blot time of 3 seconds. | |||||||||
| Details | Sample mono disperse as determined by SEC |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 (6k x 4k) / Digitization - Dimensions - Width: 1150 pixel / Digitization - Dimensions - Height: 8184 pixel / Number grids imaged: 1 / Number real images: 8109 / Average exposure time: 1.0 sec. / Average electron dose: 40.509 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: OTHER / Imaging mode: BRIGHT FIELD |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller






















 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)