[English] 日本語
 Yorodumi
Yorodumi- EMDB-1264: Following the signal sequence from ribosomal tunnel exit to signa... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-1264 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Following the signal sequence from ribosomal tunnel exit to signal recognition particle. | |||||||||
 Map data Map data | Mammalian SRP-RNC complex | |||||||||
 Sample Sample |
| |||||||||
| Function / homology |  Function and homology information Function and homology informationSRP-dependent cotranslational protein targeting to membrane / SRP-dependent cotranslational protein targeting to membrane, signal sequence recognition / endoplasmic reticulum signal peptide binding / signal recognition particle, endoplasmic reticulum targeting / granulocyte differentiation / signal recognition particle / cotranslational protein targeting to membrane / protein targeting to ER / signal-recognition-particle GTPase / exocrine pancreas development ...SRP-dependent cotranslational protein targeting to membrane / SRP-dependent cotranslational protein targeting to membrane, signal sequence recognition / endoplasmic reticulum signal peptide binding / signal recognition particle, endoplasmic reticulum targeting / granulocyte differentiation / signal recognition particle / cotranslational protein targeting to membrane / protein targeting to ER / signal-recognition-particle GTPase / exocrine pancreas development / SRP-dependent cotranslational protein targeting to membrane, translocation / 7S RNA binding / phototransduction / SRP-dependent cotranslational protein targeting to membrane / neutrophil chemotaxis / maturation of LSU-rRNA from tricistronic rRNA transcript (SSU-rRNA, 5.8S rRNA, LSU-rRNA) / visual perception / chloroplast / G protein-coupled receptor activity / GDP binding / cytoplasmic translation / cytosolic large ribosomal subunit / nuclear body / rRNA binding / ribosome / structural constituent of ribosome / nuclear speck / ribonucleoprotein complex / translation / GTPase activity / mRNA binding / GTP binding / nucleolus / endoplasmic reticulum / ATP hydrolysis activity / RNA binding / nucleus / plasma membrane / cytosol Similarity search - Function | |||||||||
| Biological species |  | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 8.7 Å | |||||||||
 Authors Authors | Halic M / Blau M / Becker T / Mielke T / Pool MR / Wild K / Sinning I / Beckmann R | |||||||||
 Citation Citation |  Journal: Nature / Year: 2006 Journal: Nature / Year: 2006Title: Following the signal sequence from ribosomal tunnel exit to signal recognition particle. Authors: Mario Halic / Michael Blau / Thomas Becker / Thorsten Mielke / Martin R Pool / Klemens Wild / Irmgard Sinning / Roland Beckmann /  Abstract: Membrane and secretory proteins can be co-translationally inserted into or translocated across the membrane. This process is dependent on signal sequence recognition on the ribosome by the signal ...Membrane and secretory proteins can be co-translationally inserted into or translocated across the membrane. This process is dependent on signal sequence recognition on the ribosome by the signal recognition particle (SRP), which results in targeting of the ribosome-nascent-chain complex to the protein-conducting channel at the membrane. Here we present an ensemble of structures at subnanometre resolution, revealing the signal sequence both at the ribosomal tunnel exit and in the bacterial and eukaryotic ribosome-SRP complexes. Molecular details of signal sequence interaction in both prokaryotic and eukaryotic complexes were obtained by fitting high-resolution molecular models. The signal sequence is presented at the ribosomal tunnel exit in an exposed position ready for accommodation in the hydrophobic groove of the rearranged SRP54 M domain. Upon ribosome binding, the SRP54 NG domain also undergoes a conformational rearrangement, priming it for the subsequent docking reaction with the NG domain of the SRP receptor. These findings provide the structural basis for improving our understanding of the early steps of co-translational protein sorting. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_1264.map.gz emd_1264.map.gz | 19.1 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-1264-v30.xml emd-1264-v30.xml emd-1264.xml emd-1264.xml | 7.9 KB 7.9 KB | Display Display |  EMDB header EMDB header |
| Images |  1264.gif 1264.gif | 39.5 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-1264 http://ftp.pdbj.org/pub/emdb/structures/EMD-1264 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-1264 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-1264 | HTTPS FTP |
-Validation report
| Summary document |  emd_1264_validation.pdf.gz emd_1264_validation.pdf.gz | 269.4 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_1264_full_validation.pdf.gz emd_1264_full_validation.pdf.gz | 268.5 KB | Display | |
| Data in XML |  emd_1264_validation.xml.gz emd_1264_validation.xml.gz | 7.1 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-1264 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-1264 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-1264 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-1264 | HTTPS FTP |
-Related structure data
| Related structure data |  2j37MC  1261C  1262C  1263C  2j28C M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_1264.map.gz / Format: CCP4 / Size: 185.7 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_1264.map.gz / Format: CCP4 / Size: 185.7 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Mammalian SRP-RNC complex | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.23 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
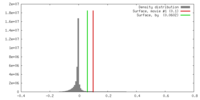
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : mammalian signal recognition particle bound to ribosome nascent c...
| Entire | Name: mammalian signal recognition particle bound to ribosome nascent chain complex |
|---|---|
| Components |
|
-Supramolecule #1000: mammalian signal recognition particle bound to ribosome nascent c...
| Supramolecule | Name: mammalian signal recognition particle bound to ribosome nascent chain complex type: sample / ID: 1000 / Number unique components: 2 |
|---|
-Supramolecule #1: ribosome
| Supramolecule | Name: ribosome / type: complex / ID: 1 / Recombinant expression: No / Ribosome-details: ribosome-prokaryote: ALL |
|---|---|
| Source (natural) | Organism:  |
-Macromolecule #1: signal recognition particle
| Macromolecule | Name: signal recognition particle / type: protein_or_peptide / ID: 1 / Name.synonym: srp / Recombinant expression: No |
|---|---|
| Source (natural) | Organism:  |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Vitrification | Cryogen name: ETHANE |
|---|
- Electron microscopy
Electron microscopy
| Microscope | FEI TECNAI F30 |
|---|---|
| Image recording | Category: FILM / Film or detector model: KODAK SO-163 FILM / Digitization - Scanner: PRIMESCAN / Average electron dose: 20 e/Å2 / Bits/pixel: 16 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 3.0 µm / Nominal defocus min: 0.9 µm |
| Sample stage | Specimen holder: f / Specimen holder model: OTHER |
| Experimental equipment |  Model: Tecnai F30 / Image courtesy: FEI Company |
- Image processing
Image processing
| Final reconstruction | Applied symmetry - Point group: C1 (asymmetric) / Resolution.type: BY AUTHOR / Resolution: 8.7 Å / Resolution method: FSC 0.5 CUT-OFF / Software - Name: spider |
|---|
 Movie
Movie Controller
Controller



























 Z (Sec.)
Z (Sec.) X (Row.)
X (Row.) Y (Col.)
Y (Col.)