+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-11213 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cryo-EM structure of DNA-PKcs (State 3) | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | Kinase / DNA-PKcs / NHEJ / DNA-repair / DNA-PK / DNA BINDING PROTEIN | |||||||||
| Function / homology |  Function and homology information Function and homology informationMHC class II antigen presentation / Neutrophil degranulation / entry into host cell by a symbiont-containing vacuole / Hydrolases; Acting on peptide bonds (peptidases); Aspartic endopeptidases / protein autoprocessing / transport vesicle / protein processing / aspartic-type endopeptidase activity / proteolysis Similarity search - Function | |||||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | |||||||||
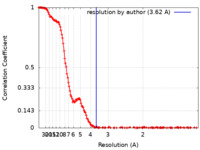
| Method | single particle reconstruction / cryo EM / Resolution: 3.62 Å | |||||||||
 Authors Authors | Chaplin AK / Hardwick SW | |||||||||
| Funding support |  United Kingdom, 1 items United Kingdom, 1 items
| |||||||||
 Citation Citation |  Journal: Nat Struct Mol Biol / Year: 2021 Journal: Nat Struct Mol Biol / Year: 2021Title: Dimers of DNA-PK create a stage for DNA double-strand break repair. Authors: Amanda K Chaplin / Steven W Hardwick / Shikang Liang / Antonia Kefala Stavridi / Ales Hnizda / Lee R Cooper / Taiana Maia De Oliveira / Dimitri Y Chirgadze / Tom L Blundell /  Abstract: DNA double-strand breaks are the most dangerous type of DNA damage and, if not repaired correctly, can lead to cancer. In humans, Ku70/80 recognizes DNA broken ends and recruits the DNA-dependent ...DNA double-strand breaks are the most dangerous type of DNA damage and, if not repaired correctly, can lead to cancer. In humans, Ku70/80 recognizes DNA broken ends and recruits the DNA-dependent protein kinase catalytic subunit (DNA-PKcs) to form DNA-dependent protein kinase holoenzyme (DNA-PK) in the process of non-homologous end joining (NHEJ). We present a 2.8-Å-resolution cryo-EM structure of DNA-PKcs, allowing precise amino acid sequence registration in regions uninterpreted in previous 4.3-Å X-ray maps. We also report a cryo-EM structure of DNA-PK at 3.5-Å resolution and reveal a dimer mediated by the Ku80 C terminus. Central to dimer formation is a domain swap of the conserved C-terminal helix of Ku80. Our results suggest a new mechanism for NHEJ utilizing a DNA-PK dimer to bring broken DNA ends together. Furthermore, drug inhibition of NHEJ in combination with chemo- and radiotherapy has proved successful, making these models central to structure-based drug targeting efforts. #1:  Journal: Nat.Struct.Mol.Biol. / Year: 2020 Journal: Nat.Struct.Mol.Biol. / Year: 2020Title: Dimers of DNA-PK create a stage for DNA-double strand break repair Authors: Chaplin AK / Hardwick SW / Liang S / Stavridi AK / Hnizda A / Cooper LR / De Oliveira TM / Chirgadze DY / Blundell TL | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_11213.map.gz emd_11213.map.gz | 286.7 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-11213-v30.xml emd-11213-v30.xml emd-11213.xml emd-11213.xml | 20.8 KB 20.8 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_11213_fsc.xml emd_11213_fsc.xml | 19.8 KB | Display |  FSC data file FSC data file |
| Images |  emd_11213.png emd_11213.png | 42 KB | ||
| Filedesc metadata |  emd-11213.cif.gz emd-11213.cif.gz | 7.9 KB | ||
| Others |  emd_11213_additional_1.map.gz emd_11213_additional_1.map.gz emd_11213_half_map_1.map.gz emd_11213_half_map_1.map.gz emd_11213_half_map_2.map.gz emd_11213_half_map_2.map.gz | 69.8 MB 281.3 MB 281.3 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-11213 http://ftp.pdbj.org/pub/emdb/structures/EMD-11213 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-11213 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-11213 | HTTPS FTP |
-Validation report
| Summary document |  emd_11213_validation.pdf.gz emd_11213_validation.pdf.gz | 1.2 MB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_11213_full_validation.pdf.gz emd_11213_full_validation.pdf.gz | 1.2 MB | Display | |
| Data in XML |  emd_11213_validation.xml.gz emd_11213_validation.xml.gz | 22.9 KB | Display | |
| Data in CIF |  emd_11213_validation.cif.gz emd_11213_validation.cif.gz | 30.2 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-11213 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-11213 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-11213 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-11213 | HTTPS FTP |
-Related structure data
| Related structure data |  6zh4MC  6zfpC  6zh2C  6zh6C  6zh8C  6zhaC  6zheC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_11213.map.gz / Format: CCP4 / Size: 303.3 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_11213.map.gz / Format: CCP4 / Size: 303.3 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.652 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
-Additional map: Resolve cryoEM density modified map at 3.48 angstrom resolution
| File | emd_11213_additional_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Resolve cryoEM density modified map at 3.48 angstrom resolution | ||||||||||||
| Projections & Slices |
| ||||||||||||
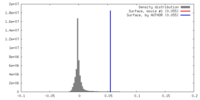
| Density Histograms |
-Half map: #1
| File | emd_11213_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #2
| File | emd_11213_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : DNA-PKcs
| Entire | Name: DNA-PKcs |
|---|---|
| Components |
|
-Supramolecule #1: DNA-PKcs
| Supramolecule | Name: DNA-PKcs / type: organelle_or_cellular_component / ID: 1 / Parent: 0 / Macromolecule list: all |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 470 KDa |
-Macromolecule #1: DNA-dependent protein kinase catalytic subunit,DNA-PKcs
| Macromolecule | Name: DNA-dependent protein kinase catalytic subunit,DNA-PKcs type: protein_or_peptide / ID: 1 / Number of copies: 1 / Enantiomer: LEVO / EC number: non-specific serine/threonine protein kinase |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 472.056281 KDa |
| Sequence | String: MAGSGAGVRC SLLRLQETLS AADRCGAALA GHQLIRGLGQ ECVLSSSPAV LALQTSLVFS RDFGLLVFVR KSLNSIEFRE CREEILKFL CIFLEKMGQK IAPYSVEIKN TCTSVYTKDR AAKCKIPALD LLIKLLQTFR SSRLMDEFKI GELFSKFYGE L ALKKKIPD ...String: MAGSGAGVRC SLLRLQETLS AADRCGAALA GHQLIRGLGQ ECVLSSSPAV LALQTSLVFS RDFGLLVFVR KSLNSIEFRE CREEILKFL CIFLEKMGQK IAPYSVEIKN TCTSVYTKDR AAKCKIPALD LLIKLLQTFR SSRLMDEFKI GELFSKFYGE L ALKKKIPD TVLEKVYELL GLLGEVHPSE MINNAENLFR AFLGELKTQM TSAVREPKLP VLAGCLKGLS SLLCNFTKSM EE DPQTSRE IFNFVLKAIR PQIDLKRYAV PSAGLRLFAL HASQFSTCLL DNYVSLFEVL LKWCAHTNVE LKKAALSALE SFL KQVSNM VAKNAEMHKN KLQYFMEQFY GIIRNVDSNN KELSIAIRGY GLFAGPCKVI NAKDVDFMYV ELIQRCKQMF LTQT DTGDD RVYQMPSFLQ SVASVLLYLD TVPEVYTPVL EHLVVMQIDS FPQYSPKMQL VCCRAIVKVF LALAAKGPVL RNCIS TVVH QGLIRICSKP VVLPKGPESE SEDHRASGEV RTGKWKVPTY KDYVDLFRHL LSSDQMMDSI LADEAFFSVN SSSESL NHL LYDEFVKSVL KIVEKLDLTL EIQTVGEQEN GDEAPGVWMI PTSDPAANLH PAKPKDFSAF INLVEFCREI LPEKQAE FF EPWVYSFSYE LILQSTRLPL ISGFYKLLSI TVRNAKKIKY FEGVSPKSLK HSPEDPEKYS CFALFVKFGK EVAVKMKQ Y KDELLASCLT FLLSLPHNII ELDVRAYVPA LQMAFKLGLS YTPLAEVGLN ALEEWSIYID RHVMQPYYKD ILPCLDGYL KTSALSDETK NNWEVSALSR AAQKGFNKVV LKHLKKTKNL SSNEAISLEE IRIRVVQMLG SLGGQINKNL LTVTSSDEMM KSYVAWDRE KRLSFAVPFR EMKPVIFLDV FLPRVTELAL TASDRQTKVA ACELLHSMVM FMLGKATQMP EGGQGAPPMY Q LYKRTFPV LLRLACDVDQ VTRQLYEPLV MQLIHWFTNN KKFESQDTVA LLEAILDGIV DPVDSTLRDF CGRCIREFLK WS IKQITPQ QQEKSPVNTK SLFKRLYSLA LHPNAFKRLG ASLAFNNIYR EFREEESLVE QFVFEALVIY MESLALAHAD EKS LGTIQQ CCDAIDHLCR IIEKKHVSLN KAKKRRLPRG FPPSASLCLL DLVKWLLAHC GRPQTECRHK SIELFYKFVP LLPG NRSPN LWLKDVLKEE GVSFLINTFE GGGCGQPSGI LAQPTLLYLR GPFSLQATLC WLDLLLAALE CYNTFIGERT VGALQ VLGT EAQSSLLKAV AFFLESIAMH DIIAAEKCFG TGAAGNRTSP QEGERYNYSK CTVVVRIMEF TTTLLNTSPE GWKLLK KDL CNTHLMRVLV QTLCEPASIG FNIGDVQVMA HLPDVCVNLM KALKMSPYKD ILETHLREKI TAQSIEELCA VNLYGPD AQ VDRSRLAAVV SACKQLHRAG LLHNILPSQS TDLHHSVGTE LLSLVYKGIA PGDERQCLPS LDLSCKQLAS GLLELAFA F GGLCERLVSL LLNPAVLSTA SLGSSQGSVI HFSHGEYFYS LFSETINTEL LKNLDLAVLE LMQSSVDNTK MVSAVLNGM LDQSFRERAN QKHQGLKLAT TILQHWKKCD SWWAKDSPLE TKMAVLALLA KILQIDSSVS FNTSHGSFPE VFTTYISLLA DTKLDLHLK GQAVTLLPFF TSLTGGSLEE LRRVLEQLIV AHFPMQSREF PPGTPRFNNY VDCMKKFLDA LELSQSPMLL E LMTEVLCR EQQHVMEELF QSSFRRIARR GSCVTQVGLL ESVYEMFRKD DPRLSFTRQS FVDRSLLTLL WHCSLDALRE FF STIVVDA IDVLKSRFTK LNESTFDTQI TKKMGYYKIL DVMYSRLPKD DVHAKESKIN QVFHGSCITE GNELTKTLIK LCY DAFTEN MAGENQLLER RRLYHCAAYN CAISVICCVF NELKFYQGFL FSEKPEKNLL IFENLIDLKR RYNFPVEVEV PMER KKKYI EIRKEAREAA NGDSDGPSYM SSLSYLADST LSEEMSQFDF STGVQSYSYS SQDPRPATGR FRRREQRDPT VHDDV LELE MDELNRHECM APLTALVKHM HRSLGPPQGE EDSVPRDLPS WMKFLHGKLG NPIVPLNIRL FLAKLVINTE EVFRPY AKH WLSPLLQLAA SENNGGEGIH YMVVEIVATI LSWTGLATPT GVPKDEVLAN RLLNFLMKHV FHPKRAVFRH NLEIIKT LV ECWKDCLSIP YRLIFEKFSG KDPNSKDNSV GIQLLGIVMA NDLPPYDPQC GIQSSEYFQA LVNNMSFVRY KEVYAAAA E VLGLILRYVM ERKNILEESL CELVAKQLKQ HQNTMEDKFI VCLNKVTKSF PPLADRFMNA VFFLLPKFHG VLKTLCLEV VLCRVEGMTE LYFQLKSKDF VQVMRHRDDE RQKVCLDIIY KMMPKLKPVE LRELLNPVVE FVSHPSTTCR EQMYNILMWI HDNYRDPES ETDNDSQEIF KLAKDVLIQG LIDENPGLQL IIRNFWSHET RLPSNTLDRL LALNSLYSPK IEVHFLSLAT N FLLEMTSM SPDYPNPMFE HPLSECEFQE YTIDSDWRFR STVLTPMFVE TQASQGTLQT RTQEGSLSAR WPVAGQIRAT QQ QHDFTLT QTADGRSSFD WLTGSSTDPL VDHTSPSSDS LLFAHKRSER LQRAPLKSVG PDFGKKRLGL PGDEVDNKVK GAA GRTDLL RLRRRFMRDQ EKLSLMYARK GVAEQKREKE IKSELKMKQD AQVVLYRSYR HGDLPDIQIK HSSLITPLQA VAQR DPIIA KQLFSSLFSG ILKEMDKFKT LSEKNNITQK LLQDFNRFLN TTFSFFPPFV SCIQDISCQH AALLSLDPAA VSAGC LASL QQPVGIRLLE EALLRLLPAE LPAKRVRGKA RLPPDVLRWV ELAKLYRSIG EYDVLRGIFT SEIGTKQITQ SALLAE ARS DYSEAAKQYD EALNKQDWVD GEPTEAEKDF WELASLDCYN HLAEWKSLEY CSTASIDSEN PPDLNKIWSE PFYQETY LP YMIRSKLKLL LQGEADQSLL TFIDKAMHGE LQKAILELHY SQELSLLYLL QDDVDRAKYY IQNGIQSFMQ NYSSIDVL L HQSRLTKLQS VQALTEIQEF ISFISKQGNL SSQVPLKRLL NTWTNRYPDA KMDPMNIWDD IITNRCFFLS KIEEKLTPL PEDNSMNVDQ DGDPSDRMEV QEQEEDISSL IRSCKFSMKM KMIDSARKQN NFSLAMKLLK ELHKESKTRD DWLVSWVQSY CRLSHCRSR SQGCSEQVLT VLKTVSLLDE NNVSSYLSKN ILAFRDQNIL LGTTYRIIAN ALSSEPACLA EIEEDKARRI L ELSGSSSE DSEKVIAGLY QRAFQHLSEA VQAAEEEAQP PSWSCGPAAG VIDAYMTLAD FCDQQLRKEE ENASVIDSAE LQ AYPALVV EKMLKALKLN SNEARLKFPR LLQIIERYPE ETLSLMTKEI SSVPCWQFIS WISHMVALLD KDQAVAVQHS VEE ITDNYP QAIVYPFIIS SESYSFKDTS TGHKNKEFVA RIKSKLDQGG VIQDFINALD QLSNPELLFK DWSNDVRAEL AKTP VNKKN IEKMYERMYA ALGDPKAPGL GAFRRKFIQT FGKEFDKHFG KGGSKLLRMK LSDFNDITNM LLLKMNKDSK PPGNL KECS PWMSDFKVEF LRNELEIPGQ YDGRGKPLPE YHVRIAGFDE RVTVMASLRR PKRIIIRGHD EREHPFLVKG GEDLRQ DQR VEQLFQVMNG ILAQDSACSQ RALQLRTYSV VPMTSRLGLI EWLENTVTLK DLLLNTMSQE EKAAYLSDPR APPCEYK DW LTKMSGKHDV GAYMLMYKGA NRTETVTSFR KRESKVPADL LKRAFVRMST SPEAFLALRS HFASSHALIC ISHWILGI G DRHLNNFMVA METGGVIGID FGHAFGSATQ FLPVPELMPF RLTRQFINLM LPMKETGLMY SIMVHALRAF RSDPGLLTN TMDVFVKEPS FDWKNFEQKM LKKGGSWIQE INVAEKNWYP RQKICYAKRK LAGANPAVIT CDELLLGHEK APAFRDYVAV ARGSKDHNI RAQEPESGLS EETQVKCLMD QATDPNILGR TWEGWEPWM(UNK) (UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK) UniProtKB: Plasmepsin X |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.4 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 (6k x 4k) / Average electron dose: 53.95 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller

















 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)