+ データを開く
データを開く
- 基本情報
基本情報
| 登録情報 | データベース: EMDB / ID: EMD-10574 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| タイトル | Beta-galactosidase in complex with PETG | |||||||||
 マップデータ マップデータ | None | |||||||||
 試料 試料 |
| |||||||||
 キーワード キーワード | Bgal / PETG / SUGAR BINDING PROTEIN | |||||||||
| 機能・相同性 |  機能・相同性情報 機能・相同性情報alkali metal ion binding / lactose catabolic process / beta-galactosidase complex / beta-galactosidase / beta-galactosidase activity / carbohydrate binding / magnesium ion binding / identical protein binding 類似検索 - 分子機能 | |||||||||
| 生物種 |  | |||||||||
| 手法 | 単粒子再構成法 / クライオ電子顕微鏡法 / 解像度: 2.2 Å | |||||||||
 データ登録者 データ登録者 | Saur M / Hartshorn MJ | |||||||||
 引用 引用 |  ジャーナル: Drug Discov Today / 年: 2020 ジャーナル: Drug Discov Today / 年: 2020タイトル: Fragment-based drug discovery using cryo-EM. 著者: Michael Saur / Michael J Hartshorn / Jing Dong / Judith Reeks / Gabor Bunkoczi / Harren Jhoti / Pamela A Williams /  要旨: Recent advances in electron cryo-microscopy (cryo-EM) structure determination have pushed the resolutions obtainable by the method into the range widely considered to be of utility for drug discovery. ...Recent advances in electron cryo-microscopy (cryo-EM) structure determination have pushed the resolutions obtainable by the method into the range widely considered to be of utility for drug discovery. Here, we review the use of cryo-EM in fragment-based drug discovery (FBDD) based on in-house method development. We demonstrate not only that cryo-EM can reveal details of the molecular interactions between fragments and a protein, but also that the current reproducibility, quality, and throughput are compatible with FBDD. We exemplify this using the test system β-galactosidase (Bgal) and the oncology target pyruvate kinase 2 (PKM2). | |||||||||
| 履歴 |
|
- 構造の表示
構造の表示
| ムービー |
 ムービービューア ムービービューア |
|---|---|
| 構造ビューア | EMマップ:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| 添付画像 |
- ダウンロードとリンク
ダウンロードとリンク
-EMDBアーカイブ
| マップデータ |  emd_10574.map.gz emd_10574.map.gz | 15.2 MB |  EMDBマップデータ形式 EMDBマップデータ形式 | |
|---|---|---|---|---|
| ヘッダ (付随情報) |  emd-10574-v30.xml emd-10574-v30.xml emd-10574.xml emd-10574.xml | 18.9 KB 18.9 KB | 表示 表示 |  EMDBヘッダ EMDBヘッダ |
| 画像 |  emd_10574.png emd_10574.png | 172.8 KB | ||
| Filedesc metadata |  emd-10574.cif.gz emd-10574.cif.gz | 6.2 KB | ||
| その他 |  emd_10574_additional.map.gz emd_10574_additional.map.gz emd_10574_additional_1.map.gz emd_10574_additional_1.map.gz emd_10574_half_map_1.map.gz emd_10574_half_map_1.map.gz emd_10574_half_map_2.map.gz emd_10574_half_map_2.map.gz | 308.8 MB 308.8 MB 261.2 MB 262.3 MB | ||
| アーカイブディレクトリ |  http://ftp.pdbj.org/pub/emdb/structures/EMD-10574 http://ftp.pdbj.org/pub/emdb/structures/EMD-10574 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-10574 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-10574 | HTTPS FTP |
-関連構造データ
| 関連構造データ |  6tteMC  6tshC  6tskC  6ttfC  6tthC  6ttiC  6ttqC C: 同じ文献を引用 ( M: このマップから作成された原子モデル |
|---|---|
| 類似構造データ | |
| 電子顕微鏡画像生データ |  EMPIAR-10644 (タイトル: Beta-galactosidase in complex with PETG / Data size: 720.1 EMPIAR-10644 (タイトル: Beta-galactosidase in complex with PETG / Data size: 720.1 Data #1: Data from EPU (movies have been converted to compressed TIF) [micrographs - multiframe]) |
- リンク
リンク
| EMDBのページ |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| 「今月の分子」の関連する項目 |
- マップ
マップ
| ファイル |  ダウンロード / ファイル: emd_10574.map.gz / 形式: CCP4 / 大きさ: 35.7 MB / タイプ: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) ダウンロード / ファイル: emd_10574.map.gz / 形式: CCP4 / 大きさ: 35.7 MB / タイプ: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 注釈 | None | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| 投影像・断面図 | 画像のコントロール
画像は Spider により作成 これらの図は立方格子座標系で作成されたものです | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| ボクセルのサイズ | X=Y=Z: 0.68 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| 密度 |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| 対称性 | 空間群: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| 詳細 | EMDB XML:
CCP4マップ ヘッダ情報:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-添付データ
-追加マップ: Relion post-process unmasked map
| ファイル | emd_10574_additional.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 注釈 | Relion post-process unmasked map | ||||||||||||
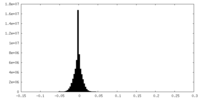
| 投影像・断面図 |
| ||||||||||||
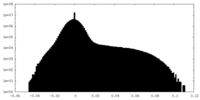
| 密度ヒストグラム |
-追加マップ: Relion post-process unmasked map
| ファイル | emd_10574_additional_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 注釈 | Relion post-process unmasked map | ||||||||||||
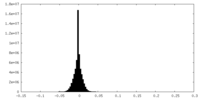
| 投影像・断面図 |
| ||||||||||||
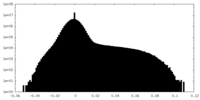
| 密度ヒストグラム |
-ハーフマップ: Relion auto-refine halfmap 1
| ファイル | emd_10574_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 注釈 | Relion auto-refine halfmap 1 | ||||||||||||
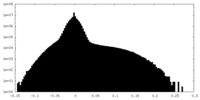
| 投影像・断面図 |
| ||||||||||||
| 密度ヒストグラム |
-ハーフマップ: Relion auto-refine halfmap 2
| ファイル | emd_10574_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 注釈 | Relion auto-refine halfmap 2 | ||||||||||||
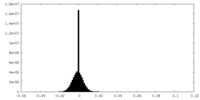
| 投影像・断面図 |
| ||||||||||||
| 密度ヒストグラム |
- 試料の構成要素
試料の構成要素
-全体 : Beta-galactosidase
| 全体 | 名称: Beta-galactosidase |
|---|---|
| 要素 |
|
-超分子 #1: Beta-galactosidase
| 超分子 | 名称: Beta-galactosidase / タイプ: complex / ID: 1 / 親要素: 0 / 含まれる分子: #1 |
|---|---|
| 由来(天然) | 生物種:  |
| 分子量 | 理論値: 464 KDa |
-分子 #1: Beta-galactosidase
| 分子 | 名称: Beta-galactosidase / タイプ: protein_or_peptide / ID: 1 / コピー数: 4 / 光学異性体: LEVO / EC番号: beta-galactosidase |
|---|---|
| 由来(天然) | 生物種:  |
| 分子量 | 理論値: 118.395336 KDa |
| 組換発現 | 生物種:  |
| 配列 | 文字列: MGSSHHHHHH SSGLVPRGSH MLEDPVVLQR RDWENPGVTQ LNRLAAHPPF ASWRNSEEAR TDRPSQQLRS LNGEWRFAWF PAPEAVPES WLECDLPEAD TVVVPSNWQM HGYDAPIYTN VTYPITVNPP FVPTENPTGC YSLTFNVDES WLQEGQTRII F DGVNSAFH ...文字列: MGSSHHHHHH SSGLVPRGSH MLEDPVVLQR RDWENPGVTQ LNRLAAHPPF ASWRNSEEAR TDRPSQQLRS LNGEWRFAWF PAPEAVPES WLECDLPEAD TVVVPSNWQM HGYDAPIYTN VTYPITVNPP FVPTENPTGC YSLTFNVDES WLQEGQTRII F DGVNSAFH LWCNGRWVGY GQDSRLPSEF DLSAFLRAGE NRLAVMVLRW SDGSYLEDQD MWRMSGIFRD VSLLHKPTTQ IS DFHVATR FNDDFSRAVL EAEVQMCGEL RDYLRVTVSL WQGETQVASG TAPFGGEIID ERGGYADRVT LRLNVENPKL WSA EIPNLY RAVVELHTAD GTLIEAEACD VGFREVRIEN GLLLLNGKPL LIRGVNRHEH HPLHGQVMDE QTMVQDILLM KQNN FNAVR CSHYPNHPLW YTLCDRYGLY VVDEANIETH GMVPMNRLTD DPRWLPAMSE RVTRMVQRDR NHPSVIIWSL GNESG HGAN HDALYRWIKS VDPSRPVQYE GGGADTTATD IICPMYARVD EDQPFPAVPK WSIKKWLSLP GETRPLILCE YAHAMG NSL GGFAKYWQAF RQYPRLQGGF VWDWVDQSLI KYDENGNPWS AYGGDFGDTP NDRQFCMNGL VFADRTPHPA LTEAKHQ QQ FFQFRLSGQT IEVTSEYLFR HSDNELLHWM VALDGKPLAS GEVPLDVAPQ GKQLIELPEL PQPESAGQLW LTVRVVQP N ATAWSEAGHI SAWQQWRLAE NLSVTLPAAS HAIPHLTTSE MDFCIELGNK RWQFNRQSGF LSQMWIGDKK QLLTPLRDQ FTRAPLDNDI GVSEATRIDP NAWVERWKAA GHYQAEAALL QCTADTLADA VLITTAHAWQ HQGKTLFISR KTYRIDGSGQ MAITVDVEV ASDTPHPARI GLNCQLAQVA ERVNWLGLGP QENYPDRLTA ACFDRWDLPL SDMYTPYVFP SENGLRCGTR E LNYGPHQW RGDFQFNISR YSQQQLMETS HRHLLHAEEG TWLNIDGFHM GIGGDDSWSP SVSAEFQLSA GRYHYQLVWC QK UniProtKB: Beta-galactosidase |
-分子 #2: MAGNESIUM ION
| 分子 | 名称: MAGNESIUM ION / タイプ: ligand / ID: 2 / コピー数: 4 / 式: MG |
|---|---|
| 分子量 | 理論値: 24.305 Da |
-分子 #3: 2-phenylethyl 1-thio-beta-D-galactopyranoside
| 分子 | 名称: 2-phenylethyl 1-thio-beta-D-galactopyranoside / タイプ: ligand / ID: 3 / コピー数: 4 / 式: PTQ |
|---|---|
| 分子量 | 理論値: 300.371 Da |
| Chemical component information |  ChemComp-PTQ: |
-分子 #4: water
| 分子 | 名称: water / タイプ: ligand / ID: 4 / コピー数: 1224 / 式: HOH |
|---|---|
| 分子量 | 理論値: 18.015 Da |
| Chemical component information |  ChemComp-HOH: |
-実験情報
-構造解析
| 手法 | クライオ電子顕微鏡法 |
|---|---|
 解析 解析 | 単粒子再構成法 |
| 試料の集合状態 | particle |
- 試料調製
試料調製
| 濃度 | 0.17 mg/mL |
|---|---|
| 緩衝液 | pH: 6.8 |
| グリッド | モデル: Quantifoil R1.2/1.3 / 材質: COPPER / メッシュ: 300 / 支持フィルム - 材質: GRAPHENE OXIDE / 支持フィルム - トポロジー: CONTINUOUS |
| 凍結 | 凍結剤: ETHANE / チャンバー内湿度: 100 % / チャンバー内温度: 277 K / 装置: FEI VITROBOT MARK IV |
- 電子顕微鏡法
電子顕微鏡法
| 顕微鏡 | FEI TITAN KRIOS |
|---|---|
| 撮影 | フィルム・検出器のモデル: FEI FALCON III (4k x 4k) 検出モード: COUNTING / 撮影したグリッド数: 1 / 実像数: 562 / 平均露光時間: 59.98 sec. / 平均電子線量: 65.53 e/Å2 |
| 電子線 | 加速電圧: 300 kV / 電子線源:  FIELD EMISSION GUN FIELD EMISSION GUN |
| 電子光学系 | 照射モード: FLOOD BEAM / 撮影モード: BRIGHT FIELD |
| 実験機器 |  モデル: Titan Krios / 画像提供: FEI Company |
 ムービー
ムービー コントローラー
コントローラー






















 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)