[English] 日本語
 Yorodumi
Yorodumi- EMDB-1036: Structure of a fast kinesin: implications for ATPase mechanism an... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-1036 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Structure of a fast kinesin: implications for ATPase mechanism and interactions with microtubules. | |||||||||
 Map data Map data | Neurospora crassa kinesin monomer (nK355) AMP-PNP state | |||||||||
 Sample Sample |
| |||||||||
| Biological species |  Neurospora crassa (fungus) Neurospora crassa (fungus) | |||||||||
| Method | helical reconstruction / cryo EM / negative staining / Resolution: 25.0 Å | |||||||||
 Authors Authors | Krebs A | |||||||||
 Citation Citation |  Journal: EMBO J / Year: 2001 Journal: EMBO J / Year: 2001Title: Structure of a fast kinesin: implications for ATPase mechanism and interactions with microtubules. Authors: Y H Song / A Marx / J Müller / G Woehlke / M Schliwa / A Krebs / A Hoenger / E Mandelkow /  Abstract: We determined the crystal structure of the motor domain of the fast fungal kinesin from Neurospora crassa (NcKin). The structure has several unique features. (i) Loop 11 in the switch 2 region is ...We determined the crystal structure of the motor domain of the fast fungal kinesin from Neurospora crassa (NcKin). The structure has several unique features. (i) Loop 11 in the switch 2 region is ordered and enables one to describe the complete nucleotide-binding pocket, including three inter-switch salt bridges between switch 1 and 2. (ii) Loop 9 in the switch 1 region bends outwards, making the nucleotide-binding pocket very wide. The displacement in switch 1 resembles that of the G-protein ras complexed with its guanosine nucleotide exchange factor. (iii) Loop 5 in the entrance to the nucleotide-binding pocket is remarkably long and interacts with the ribose of ATP. (iv) The linker and neck region is not well defined, indicating that it is mobile. (v) Image reconstructions of ice-embedded microtubules decorated with NcKin show that it interacts with several tubulin subunits, including a central beta-tubulin monomer and the two flanking alpha-tubulin monomers within the microtubule protofilament. Comparison of NcKin with other kinesins, myosin and G-proteins suggests that the rate-limiting step of ADP release is accelerated in the fungal kinesin and accounts for the unusually high velocity and ATPase activity. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_1036.map.gz emd_1036.map.gz | 2.8 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-1036-v30.xml emd-1036-v30.xml emd-1036.xml emd-1036.xml | 9.7 KB 9.7 KB | Display Display |  EMDB header EMDB header |
| Images |  1036.gif 1036.gif | 64.9 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-1036 http://ftp.pdbj.org/pub/emdb/structures/EMD-1036 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-1036 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-1036 | HTTPS FTP |
-Validation report
| Summary document |  emd_1036_validation.pdf.gz emd_1036_validation.pdf.gz | 250.5 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_1036_full_validation.pdf.gz emd_1036_full_validation.pdf.gz | 249.6 KB | Display | |
| Data in XML |  emd_1036_validation.xml.gz emd_1036_validation.xml.gz | 5 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-1036 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-1036 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-1036 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-1036 | HTTPS FTP |
-Related structure data
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_1036.map.gz / Format: CCP4 / Size: 3.7 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_1036.map.gz / Format: CCP4 / Size: 3.7 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Neurospora crassa kinesin monomer (nK355) AMP-PNP state | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 5.526 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
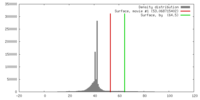
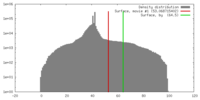
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : neurospora crassa kinesin monomer
| Entire | Name: neurospora crassa kinesin monomer |
|---|---|
| Components |
|
-Supramolecule #1000: neurospora crassa kinesin monomer
| Supramolecule | Name: neurospora crassa kinesin monomer / type: sample / ID: 1000 / Oligomeric state: monomer / Number unique components: 2 |
|---|
-Macromolecule #1: neurospora crassakinesin
| Macromolecule | Name: neurospora crassakinesin / type: protein_or_peptide / ID: 1 / Name.synonym: molecular motor / Number of copies: 1 / Oligomeric state: monomer / Recombinant expression: Yes / Database: NCBI |
|---|---|
| Source (natural) | Organism:  Neurospora crassa (fungus) / synonym: neurospora Neurospora crassa (fungus) / synonym: neurospora |
-Macromolecule #2: tubulin
| Macromolecule | Name: tubulin / type: protein_or_peptide / ID: 2 / Name.synonym: microtubules / Number of copies: 1 / Oligomeric state: hetero dimer / Recombinant expression: No / Database: NCBI |
|---|---|
| Source (natural) | Organism:  Neurospora crassa (fungus) / synonym: neurospora / Tissue: brain / Cell: neuronal cells / Location in cell: cytoplasm Neurospora crassa (fungus) / synonym: neurospora / Tissue: brain / Cell: neuronal cells / Location in cell: cytoplasm |
-Experimental details
-Structure determination
| Method | negative staining, cryo EM |
|---|---|
 Processing Processing | helical reconstruction |
| Aggregation state | filament |
- Sample preparation
Sample preparation
| Concentration | 0.5 mg/mL |
|---|---|
| Buffer | pH: 6.9 Details: Pipes 20mM, 50 mM NaCl, 5mM mgcl,1mM Mg-ATP, 20um taxol. |
| Staining | Type: NEGATIVE / Details: ice-embedded |
| Grid | Details: holey grids |
| Vitrification | Cryogen name: PROPANE / Chamber temperature: 93 K / Instrument: HOMEMADE PLUNGER / Details: Vitrification instrument: self made |
- Electron microscopy
Electron microscopy
| Microscope | FEI/PHILIPS CM120T |
|---|---|
| Image recording | Category: FILM / Film or detector model: KODAK SO-163 FILM / Digitization - Scanner: ZEISS SCAI / Digitization - Sampling interval: 21 µm / Number real images: 18 / Average electron dose: 5 e/Å2 / Bits/pixel: 16 |
| Tilt angle min | 0 |
| Tilt angle max | 0 |
| Electron beam | Acceleration voltage: 100 kV / Electron source: LAB6 |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 2.0 µm / Nominal defocus min: 1.5 µm / Nominal magnification: 37000 |
| Sample stage | Specimen holder: side entry / Specimen holder model: GATAN LIQUID NITROGEN |
- Image processing
Image processing
| Final reconstruction | Applied symmetry - Helical parameters - Axial symmetry: C1 (asymmetric) Algorithm: OTHER / Resolution.type: BY AUTHOR / Resolution: 25.0 Å / Resolution method: FSC 0.5 CUT-OFF / Software - Name: Phoelix, Suprim Details: Final maps from 36 averaged datasets = 18 helical tubes |
|---|
 Movie
Movie Controller
Controller


 UCSF Chimera
UCSF Chimera








 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)