+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-0356 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cryo-EM structure of the 2:1 hPtch1-Shhp complex | |||||||||
 Map data Map data | em-volume_P1 | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | Receptor / RND family / PROTEIN BINDING | |||||||||
| Function / homology |  Function and homology information Function and homology informationregulation of nodal signaling pathway / trunk neural crest cell migration / neural plate axis specification / response to chlorate / cell differentiation involved in kidney development / positive regulation of skeletal muscle cell proliferation / right lung development / left lung development / primary prostatic bud elongation / : ...regulation of nodal signaling pathway / trunk neural crest cell migration / neural plate axis specification / response to chlorate / cell differentiation involved in kidney development / positive regulation of skeletal muscle cell proliferation / right lung development / left lung development / primary prostatic bud elongation / : / mesenchymal smoothened signaling pathway involved in prostate gland development / positive regulation of sclerotome development / tracheoesophageal septum formation / negative regulation of ureter smooth muscle cell differentiation / positive regulation of ureter smooth muscle cell differentiation / negative regulation of kidney smooth muscle cell differentiation / positive regulation of kidney smooth muscle cell differentiation / morphogen activity / regulation of odontogenesis / positive regulation of mesenchymal cell proliferation involved in ureter development / hedgehog receptor activity / Formation of lateral plate mesoderm / regulation of glial cell proliferation / hindgut morphogenesis / polarity specification of anterior/posterior axis / cell proliferation involved in metanephros development / negative regulation of alpha-beta T cell differentiation / regulation of prostatic bud formation / neural tube patterning / formation of anatomical boundary / smoothened binding / positive regulation of striated muscle cell differentiation / ventral midline development / metanephric mesenchymal cell proliferation involved in metanephros development / trachea morphogenesis / hedgehog family protein binding / cholesterol-protein transferase activity / HHAT G278V doesn't palmitoylate Hh-Np / telencephalon regionalization / bud outgrowth involved in lung branching / epithelial-mesenchymal cell signaling / Ligand-receptor interactions / hindlimb morphogenesis / laminin-1 binding / lung epithelium development / salivary gland cavitation / epidermal cell fate specification / negative regulation of mesenchymal cell apoptotic process / spinal cord dorsal/ventral patterning / determination of left/right asymmetry in lateral mesoderm / negative regulation of cholesterol efflux / establishment of epithelial cell polarity / skeletal muscle cell proliferation / spinal cord motor neuron differentiation / negative regulation of T cell differentiation in thymus / positive regulation of T cell differentiation in thymus / cell development / prostate gland development / intermediate filament organization / skeletal muscle fiber differentiation / embryonic skeletal system development / stem cell development / mesenchymal cell apoptotic process / positive regulation of cerebellar granule cell precursor proliferation / limb bud formation / animal organ formation / Developmental Lineage of Multipotent Pancreatic Progenitor Cells / patched binding / negative regulation of cell division / positive regulation of skeletal muscle tissue development / embryonic digestive tract morphogenesis / somite development / hindbrain development / ectoderm development / cerebellar granule cell precursor proliferation / embryonic foregut morphogenesis / limb morphogenesis / epithelial cell proliferation involved in salivary gland morphogenesis / mesenchymal cell proliferation involved in lung development / neuron fate commitment / negative regulation of dopaminergic neuron differentiation / Activation of SMO / self proteolysis / positive regulation of immature T cell proliferation in thymus / regulation of stem cell proliferation / lung lobe morphogenesis / positive regulation of astrocyte differentiation / smooth muscle tissue development / artery development / CD4-positive or CD8-positive, alpha-beta T cell lineage commitment / pharyngeal system development / lymphoid progenitor cell differentiation / mammary gland duct morphogenesis / epithelial cell proliferation involved in prostate gland development / cellular response to cholesterol / mammary gland epithelial cell differentiation / positive regulation of epithelial cell proliferation involved in prostate gland development / negative thymic T cell selection / male genitalia development / pattern specification process Similarity search - Function | |||||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 3.6 Å | |||||||||
 Authors Authors | Yan N / Gong X | |||||||||
| Funding support |  United States, 1 items United States, 1 items
| |||||||||
 Citation Citation |  Journal: Nat Commun / Year: 2019 Journal: Nat Commun / Year: 2019Title: Inhibition of tetrameric Patched1 by Sonic Hedgehog through an asymmetric paradigm. Authors: Hongwu Qian / Pingping Cao / Miaohui Hu / Shuai Gao / Nieng Yan / Xin Gong /   Abstract: The Hedgehog (Hh) pathway controls embryonic development and postnatal tissue maintenance and regeneration. Inhibition of Hh receptor Patched (Ptch) by the Hh ligands relieves suppression of ...The Hedgehog (Hh) pathway controls embryonic development and postnatal tissue maintenance and regeneration. Inhibition of Hh receptor Patched (Ptch) by the Hh ligands relieves suppression of signaling cascades. Here, we report the cryo-EM structure of tetrameric Ptch1 in complex with the palmitoylated N-terminal signaling domain of human Sonic hedgehog (ShhN) at a 4:2 stoichiometric ratio. The structure shows that four Ptch1 protomers are organized as a loose dimer of dimers. Each dimer binds to one ShhN through two distinct inhibitory interfaces, one mainly through the N-terminal peptide and the palmitoyl moiety of ShhN and the other through the Ca-mediated interface on ShhN. Map comparison reveals that the cholesteryl moiety of native ShhN occupies a recently identified extracellular steroid binding pocket in Ptch1. Our structure elucidates the tetrameric assembly of Ptch1 and suggests an asymmetric mode of action of the Hh ligands for inhibiting the potential cholesterol transport activity of Ptch1. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_0356.map.gz emd_0356.map.gz | 77.2 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-0356-v30.xml emd-0356-v30.xml emd-0356.xml emd-0356.xml | 22.5 KB 22.5 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_0356.png emd_0356.png | 67.6 KB | ||
| Filedesc metadata |  emd-0356.cif.gz emd-0356.cif.gz | 8.1 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-0356 http://ftp.pdbj.org/pub/emdb/structures/EMD-0356 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-0356 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-0356 | HTTPS FTP |
-Related structure data
| Related structure data |  6n7hMC  0355C  0358C  6n7gC  6n7kC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | |
| EM raw data |  EMPIAR-10456 (Title: Tetrameric Ptch1 complexed with ShhNp / Data size: 8.9 TB EMPIAR-10456 (Title: Tetrameric Ptch1 complexed with ShhNp / Data size: 8.9 TBData #1: Oligomeric complex of Ptch1 and Shhp [micrographs - multiframe]) |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_0356.map.gz / Format: CCP4 / Size: 83.7 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_0356.map.gz / Format: CCP4 / Size: 83.7 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | em-volume_P1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.114 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
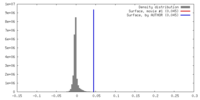
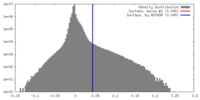
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : Patch1
| Entire | Name: Patch1 |
|---|---|
| Components |
|
-Supramolecule #1: Patch1
| Supramolecule | Name: Patch1 / type: organelle_or_cellular_component / ID: 1 / Parent: 0 / Macromolecule list: #1 |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
-Macromolecule #1: Protein patched homolog 1
| Macromolecule | Name: Protein patched homolog 1 / type: protein_or_peptide / ID: 1 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 150.189578 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: MADYKDDDDK SGPDEVDASG RMASAGNAAE PQDRGGGGSG CIGAPGRPAG GGRRRRTGGL RRAAAPDRDY LHRPSYCDAA FALEQISKG KATGRKAPLW LRAKFQRLLF KLGCYIQKNC GKFLVVGLLI FGAFAVGLKA ANLETNVEEL WVEVGGRVSR E LNYTRQKI ...String: MADYKDDDDK SGPDEVDASG RMASAGNAAE PQDRGGGGSG CIGAPGRPAG GGRRRRTGGL RRAAAPDRDY LHRPSYCDAA FALEQISKG KATGRKAPLW LRAKFQRLLF KLGCYIQKNC GKFLVVGLLI FGAFAVGLKA ANLETNVEEL WVEVGGRVSR E LNYTRQKI GEEAMFNPQL MIQTPKEEGA NVLTTEALLQ HLDSALQASR VHVYMYNRQW KLEHLCYKSG ELITETGYMD QI IEYLYPC LIITPLDCFW EGAKLQSGTA YLLGKPPLRW TNFDPLEFLE ELKKINYQVD SWEEMLNKAE VGHGYMDRPC LNP ADPDCP ATAPNKNSTK PLDMALVLNG GCHGLSRKYM HWQEELIVGG TVKNSTGKLV SAHALQTMFQ LMTPKQMYEH FKGY EYVSH INWNEDKAAA ILEAWQRTYV EVVHQSVAQN STQKVLSFTT TTLDDILKSF SDVSVIRVAS GYLLMLAYAC LTMLR WDCS KSQGAVGLAG VLLVALSVAA GLGLCSLIGI SFNAATTQVL PFLALGVGVD DVFLLAHAFS ETGQNKRIPF EDRTGE CLK RTGASVALTS ISNVTAFFMA ALIPIPALRA FSLQAAVVVV FNFAMVLLIF PAILSMDLYR REDRRLDIFC CFTSPCV SR VIQVEPQAYT DTHDNTRYSP PPPYSSHSFA HETQITMQST VQLRTEYDPH THVYYTTAEP RSEISVQPVT VTQDTLSC Q SPESTSSTRD LLSQFSDSSL HCLEPPCTKW TLSSFAEKHY APFLLKPKAK VVVIFLFLGL LGVSLYGTTR VRDGLDLTD IVPRETREYD FIAAQFKYFS FYNMYIVTQK ADYPNIQHLL YDLHRSFSNV KYVMLEENKQ LPKMWLHYFR DWLQGLQDAF DSDWETGKI MPNNYKNGSD DGVLAYKLLV QTGSRDKPID ISQLTKQRLV DADGIINPSA FYIYLTAWVS NDPVAYAASQ A NIRPHRPE WVHDKADYMP ETRLRIPAAE PIEYAQFPFY LNGLRDTSDF VEAIEKVRTI CSNYTSLGLS SYPNGYPFLF WE QYIGLRH WLLLFISVVL ACTFLVCAVF LLNPWTAGII VMVLALMTVE LFGMMGLIGI KLSAVPVVIL IASVGIGVEF TVH VALAFL TAIGDKNRRA VLALEHMFAP VLDGAVSTLL GVLMLAGSEF DFIVRYFFAV LAILTILGVL NGLVLLPVLL SFFG PYPEV SPANGLNRLP TPSPEPPPSV VRFAMPPGHT HSGSDSSDSE YSSQTTVSGL SEELRHYEAQ QGAGGPAHQV IVEAT ENPV FAHSTVVHPE SRHHPPSNPR QQPHLDSGSL PPGRQGQQPR RDLEGSDEVD AVEGSHHHHH HHHHH UniProtKB: Protein patched homolog 1 |
-Macromolecule #2: Sonic hedgehog protein
| Macromolecule | Name: Sonic hedgehog protein / type: protein_or_peptide / ID: 2 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 19.594039 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: CGPGRGFGKR RHPKKLTPLA YKQFIPNVAE KTLGASGRYE GKISRNSERF KELTPNYNPD IIFKDEENTG ADRLMTQRCK DKLNALAIS VMNQWPGVKL RVTEGWDEDG HHSEESLHYE GRAVDITTSD RDRSKYGMLA RLAVEAGFDW VYYESKAHIH C SVKAENSV AAKSGG UniProtKB: Sonic hedgehog protein |
-Macromolecule #4: 2-acetamido-2-deoxy-beta-D-glucopyranose
| Macromolecule | Name: 2-acetamido-2-deoxy-beta-D-glucopyranose / type: ligand / ID: 4 / Number of copies: 10 / Formula: NAG |
|---|---|
| Molecular weight | Theoretical: 221.208 Da |
| Chemical component information |  ChemComp-NAG: |
-Macromolecule #5: CHOLESTEROL
| Macromolecule | Name: CHOLESTEROL / type: ligand / ID: 5 / Number of copies: 5 / Formula: CLR |
|---|---|
| Molecular weight | Theoretical: 386.654 Da |
| Chemical component information |  ChemComp-CLR: |
-Macromolecule #6: ZINC ION
| Macromolecule | Name: ZINC ION / type: ligand / ID: 6 / Number of copies: 1 / Formula: ZN |
|---|---|
| Molecular weight | Theoretical: 65.409 Da |
-Macromolecule #7: CALCIUM ION
| Macromolecule | Name: CALCIUM ION / type: ligand / ID: 7 / Number of copies: 2 / Formula: CA |
|---|---|
| Molecular weight | Theoretical: 40.078 Da |
-Macromolecule #8: PALMITIC ACID
| Macromolecule | Name: PALMITIC ACID / type: ligand / ID: 8 / Number of copies: 1 / Formula: PLM |
|---|---|
| Molecular weight | Theoretical: 256.424 Da |
| Chemical component information |  ChemComp-PLM: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 15 mg/mL |
|---|---|
| Buffer | pH: 8 |
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Detector mode: SUPER-RESOLUTION / Average electron dose: 50.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller


















 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)