6NFV
 
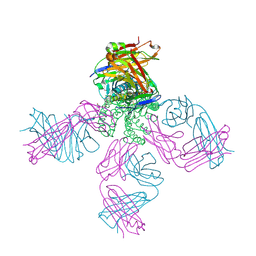 | | Structure of the KcsA-G77C mutant or the 2,4-ion bound configuration of a K+ channel selectivity filter. | | Descriptor: | (1S)-2-HYDROXY-1-[(NONANOYLOXY)METHYL]ETHYL MYRISTATE, NONAN-1-OL, POTASSIUM ION, ... | | Authors: | Tilegenova, C, Cortes, D.M, Jahovic, N, Hardy, E, Parameswaran, H, Guan, L, Cuello, L.G. | | Deposit date: | 2018-12-20 | | Release date: | 2019-08-07 | | Last modified: | 2019-12-18 | | Method: | X-RAY DIFFRACTION (2.13 Å) | | Cite: | Structure, function, and ion-binding properties of a K+channel stabilized in the 2,4-ion-bound configuration.
Proc.Natl.Acad.Sci.USA, 116, 2019
|
|
6NFU
 
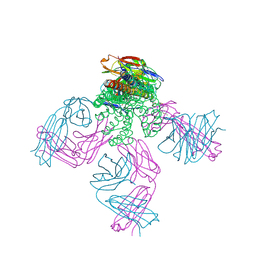 | | Structure of the KcsA-G77A mutant or the 2,4-ion bound configuration of a K+ channel selectivity filter. | | Descriptor: | (1S)-2-HYDROXY-1-[(NONANOYLOXY)METHYL]ETHYL MYRISTATE, NONAN-1-OL, POTASSIUM ION, ... | | Authors: | Tilegenova, C, Cortes, D.M, Jahovic, N, Hardy, E, Parameswaran, H, Guan, L, Cuello, L.G. | | Deposit date: | 2018-12-20 | | Release date: | 2019-08-07 | | Last modified: | 2023-10-11 | | Method: | X-RAY DIFFRACTION (2.09 Å) | | Cite: | Structure, function, and ion-binding properties of a K+channel stabilized in the 2,4-ion-bound configuration.
Proc.Natl.Acad.Sci.USA, 116, 2019
|
|
6CPV
 
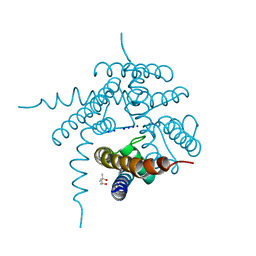 | |
6CQ6
 
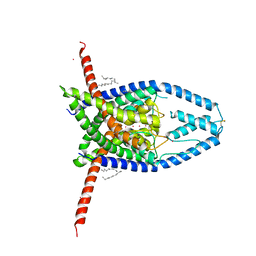 | | K2P2.1(TREK-1) apo structure | | Descriptor: | CADMIUM ION, DECANE, POTASSIUM ION, ... | | Authors: | Lolicato, M, Minor, D.L. | | Deposit date: | 2018-03-14 | | Release date: | 2018-03-28 | | Last modified: | 2023-10-04 | | Method: | X-RAY DIFFRACTION (3.1 Å) | | Cite: | K2P2.1 (TREK-1)-activator complexes reveal a cryptic selectivity filter binding site.
Nature, 547, 2017
|
|
6CQ9
 
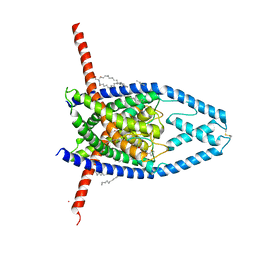 | | K2P2.1(TREK-1):ML402 complex | | Descriptor: | CADMIUM ION, HEXADECANE, N-((E,2S,3R)-1,3-DIHYDROXYOCTADEC-4-EN-2-YL)PALMITAMIDE, ... | | Authors: | Lolicato, M, Minor, D.L. | | Deposit date: | 2018-03-14 | | Release date: | 2018-04-11 | | Last modified: | 2023-10-04 | | Method: | X-RAY DIFFRACTION (2.8 Å) | | Cite: | K2P2.1 (TREK-1)-activator complexes reveal a cryptic selectivity filter binding site.
Nature, 547, 2017
|
|
6CQ8
 
 | | K2P2.1(TREK-1):ML335 complex | | Descriptor: | CADMIUM ION, HEXADECANE, N-((E,2S,3R)-1,3-DIHYDROXYOCTADEC-4-EN-2-YL)PALMITAMIDE, ... | | Authors: | Lolicato, M, Minor, D.L. | | Deposit date: | 2018-03-14 | | Release date: | 2018-03-28 | | Last modified: | 2023-10-04 | | Method: | X-RAY DIFFRACTION (3 Å) | | Cite: | K2P2.1 (TREK-1)-activator complexes reveal a cryptic selectivity filter binding site.
Nature, 547, 2017
|
|
6CNM
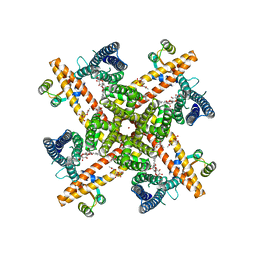 
 | | Cryo-EM structure of the human SK4/calmodulin channel complex | | Descriptor: | (2S)-3-(hexadecanoyloxy)-2-[(9Z)-octadec-9-enoyloxy]propyl 2-(trimethylammonio)ethyl phosphate, Calmodulin-1, DODECYL-BETA-D-MALTOSIDE, ... | | Authors: | Lee, C.H, MacKinnon, R. | | Deposit date: | 2018-03-08 | | Release date: | 2018-05-02 | | Last modified: | 2024-03-13 | | Method: | ELECTRON MICROSCOPY (3.4 Å) | | Cite: | Activation mechanism of a human SK-calmodulin channel complex elucidated by cryo-EM structures.
Science, 360, 2018
|
|
6CNN
 
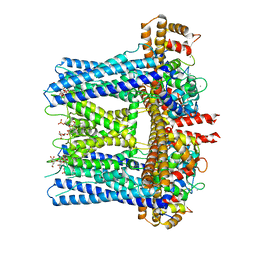 | |
6CNO
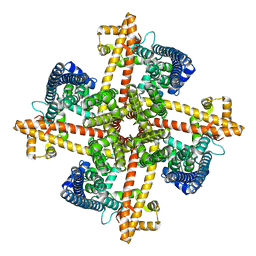 
 | |
6CJU
 
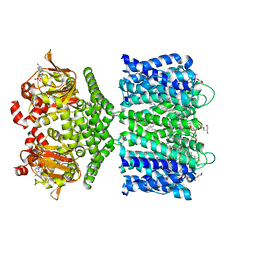 | | Structure of the SthK cyclic nucleotide-gated potassium channel in complex with cAMP | | Descriptor: | (1R)-2-{[(S)-{[(2S)-2,3-dihydroxypropyl]oxy}(hydroxy)phosphoryl]oxy}-1-[(hexadecanoyloxy)methyl]ethyl (9Z)-octadec-9-enoate, ADENOSINE-3',5'-CYCLIC-MONOPHOSPHATE, SthK cyclic nucleotide-gated potassium channel | | Authors: | Nimigean, C.M, Rheinberger, J. | | Deposit date: | 2018-02-26 | | Release date: | 2018-08-01 | | Last modified: | 2024-03-13 | | Method: | ELECTRON MICROSCOPY (3.35 Å) | | Cite: | Ligand discrimination and gating in cyclic nucleotide-gated ion channels from apo and partial agonist-bound cryo-EM structures.
Elife, 7, 2018
|
|
6CJQ
 
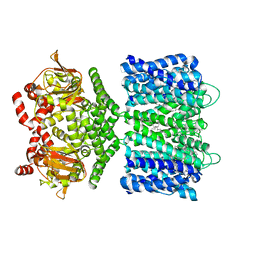 | | Structure of the SthK cyclic nucleotide-gated potassium channel | | Descriptor: | (1R)-2-{[(S)-{[(2S)-2,3-dihydroxypropyl]oxy}(hydroxy)phosphoryl]oxy}-1-[(hexadecanoyloxy)methyl]ethyl (9Z)-octadec-9-enoate, SthK cyclic nucleotide-gated potassium channel | | Authors: | Nimigean, C.M, Rheinberger, J. | | Deposit date: | 2018-02-26 | | Release date: | 2018-08-01 | | Last modified: | 2024-03-13 | | Method: | ELECTRON MICROSCOPY (3.42 Å) | | Cite: | Ligand discrimination and gating in cyclic nucleotide-gated ion channels from apo and partial agonist-bound cryo-EM structures.
Elife, 7, 2018
|
|
6CJT
 
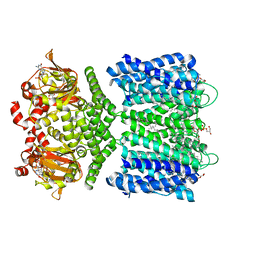 | | Structure of the SthK cyclic nucleotide-gated potassium channel in complex with cGMP | | Descriptor: | (1R)-2-{[(S)-{[(2S)-2,3-dihydroxypropyl]oxy}(hydroxy)phosphoryl]oxy}-1-[(hexadecanoyloxy)methyl]ethyl (9Z)-octadec-9-enoate, CYCLIC GUANOSINE MONOPHOSPHATE, SthK cyclic nucleotide-gated potassium channel | | Authors: | Nimigean, C.M, Rheinberger, J. | | Deposit date: | 2018-02-26 | | Release date: | 2018-08-01 | | Last modified: | 2024-03-13 | | Method: | ELECTRON MICROSCOPY (3.46 Å) | | Cite: | Ligand discrimination and gating in cyclic nucleotide-gated ion channels from apo and partial agonist-bound cryo-EM structures.
Elife, 7, 2018
|
|
6FIZ
 
 | | Crystal Structure of CNG mimicking NaK-EAPP mutant (T67A) cocrystallized with K+ | | Descriptor: | (4S)-2-METHYL-2,4-PENTANEDIOL, GLYCINE, POTASSIUM ION, ... | | Authors: | Napolitano, L.M.R, De March, M, Steiner, R.A, Onesti, S. | | Deposit date: | 2018-01-19 | | Release date: | 2019-03-06 | | Last modified: | 2024-01-17 | | Method: | X-RAY DIFFRACTION (2.63 Å) | | Cite: | Crystal Structure of CNG mimicking NaK-EAPP mutant (T67A) cocrystallized with K+
To Be Published
|
|
6BY3
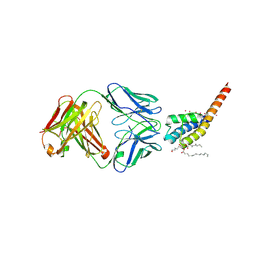 
 | | Open and conductive conformation of KcsA-T75A mutant | | Descriptor: | Antibody Heavy Chain, Antibody Light Chain, DIACYL GLYCEROL, ... | | Authors: | Labro, A.J, Cortes, D.M, Tilegenova, C, Cuello, L.G. | | Deposit date: | 2017-12-19 | | Release date: | 2018-05-09 | | Last modified: | 2020-01-01 | | Method: | X-RAY DIFFRACTION (2.37 Å) | | Cite: | Inverted allosteric coupling between activation and inactivation gates in K+channels.
Proc. Natl. Acad. Sci. U.S.A., 115, 2018
|
|
6BY2
 
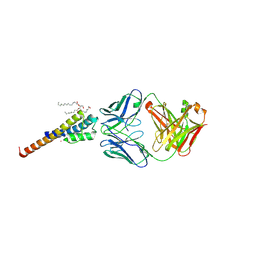 | | Closed and deep-inactivated conformation of KcsA-T75A mutant | | Descriptor: | Antibody Heavy Chain, Antibody Light Chain, DIACYL GLYCEROL, ... | | Authors: | Labro, A.J, Cortes, D.M, Tilegenova, C, Cuello, L.G. | | Deposit date: | 2017-12-19 | | Release date: | 2018-05-09 | | Last modified: | 2020-01-01 | | Method: | X-RAY DIFFRACTION (2.35 Å) | | Cite: | Inverted allosteric coupling between activation and inactivation gates in K+channels.
Proc. Natl. Acad. Sci. U.S.A., 115, 2018
|
|
5VKH
 
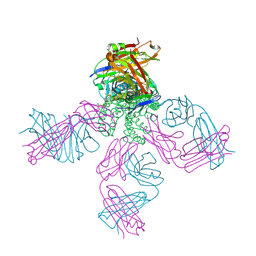 | | Closed conformation of KcsA Y82A-F103A mutant | | Descriptor: | (1S)-2-HYDROXY-1-[(NONANOYLOXY)METHYL]ETHYL MYRISTATE, Antibody Heavy Chain, Antibody Light Chain, ... | | Authors: | Cuello, L.G, Perozo, E, Cortes, D.M. | | Deposit date: | 2017-04-21 | | Release date: | 2017-12-06 | | Last modified: | 2020-01-01 | | Method: | X-RAY DIFFRACTION (2.25 Å) | | Cite: | The gating cycle of a K+ channel at atomic resolution.
Elife, 6, 2017
|
|
5VK6
 
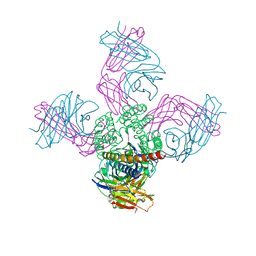 | | Open conformation of KcsA non-inactivating E71A mutant | | Descriptor: | (1S)-2-HYDROXY-1-[(NONANOYLOXY)METHYL]ETHYL MYRISTATE, Antibody Heavy Chain, Antibody Light Chain, ... | | Authors: | Cuello, L.G, Perozo, E. | | Deposit date: | 2017-04-21 | | Release date: | 2017-12-06 | | Last modified: | 2020-01-01 | | Method: | X-RAY DIFFRACTION (2.25 Å) | | Cite: | The gating cycle of a K+ channel at atomic resolution.
Elife, 6, 2017
|
|
5VKE
 
 | | Open conformation of KcsA deep-inactivated | | Descriptor: | (1S)-2-HYDROXY-1-[(NONANOYLOXY)METHYL]ETHYL MYRISTATE, Antibody Heavy Chain, Antibody Light Chain, ... | | Authors: | Cuello, L.G, Perozo, E, Cortes, D.M. | | Deposit date: | 2017-04-21 | | Release date: | 2017-12-06 | | Last modified: | 2024-04-03 | | Method: | X-RAY DIFFRACTION (2.37 Å) | | Cite: | The gating cycle of a K+ channel at atomic resolution.
Elife, 6, 2017
|
|
5U76
 
 | |
5U70
 
 | |
5J9P
 
 | | KcsA in vitro | | Descriptor: | Fab, POTASSIUM ION, pH-gated potassium channel KcsA | | Authors: | Matulef, K, Valiyaveetil, F.I. | | Deposit date: | 2016-04-10 | | Release date: | 2016-07-20 | | Last modified: | 2019-12-04 | | Method: | X-RAY DIFFRACTION (2.85 Å) | | Cite: | Combining in Vitro Folding with Cell Free Protein Synthesis for Membrane Protein Expression.
Biochemistry, 55, 2016
|
|
5EC2
 
 | | KcsA with V76ester+G77dA mutations | | Descriptor: | Antibody Fab Fragment Light Chain, DIACYL GLYCEROL, NONAN-1-OL, ... | | Authors: | Matulef, K, Valiyaveetil, F.I. | | Deposit date: | 2015-10-20 | | Release date: | 2016-04-20 | | Last modified: | 2023-11-15 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Individual Ion Binding Sites in the K(+) Channel Play Distinct Roles in C-type Inactivation and in Recovery from Inactivation.
Structure, 24, 2016
|
|
5EC1
 
 | | KcsA with V76ester mutation | | Descriptor: | Antibody Fab Fragment Light Chain, DIACYL GLYCEROL, NONAN-1-OL, ... | | Authors: | Matulef, K, Valiyaveetil, F.I. | | Deposit date: | 2015-10-20 | | Release date: | 2016-04-20 | | Last modified: | 2023-11-15 | | Method: | X-RAY DIFFRACTION (2.75 Å) | | Cite: | Individual Ion Binding Sites in the K(+) Channel Play Distinct Roles in C-type Inactivation and in Recovery from Inactivation.
Structure, 24, 2016
|
|
5EBW
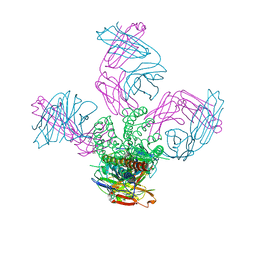 
 | | KcsA with G77ester mutation | | Descriptor: | Antibody Fab Fragment Light Chain, DIACYL GLYCEROL, NONAN-1-OL, ... | | Authors: | Matulef, K, Valiyaveetil, F.I. | | Deposit date: | 2015-10-19 | | Release date: | 2016-04-20 | | Last modified: | 2023-11-15 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Individual Ion Binding Sites in the K(+) Channel Play Distinct Roles in C-type Inactivation and in Recovery from Inactivation.
Structure, 24, 2016
|
|
5EBM
 
 | | KcsA T75G mutant in the nonconductive state | | Descriptor: | Antibody Fab Fragment Light Chain, DIACYL GLYCEROL, NONAN-1-OL, ... | | Authors: | Matulef, K, Valiyaveetil, F.I. | | Deposit date: | 2015-10-19 | | Release date: | 2016-04-20 | | Last modified: | 2023-09-27 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | Individual Ion Binding Sites in the K(+) Channel Play Distinct Roles in C-type Inactivation and in Recovery from Inactivation.
Structure, 24, 2016
|
|
