5NU7
 
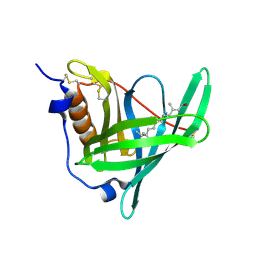 | | Structure of human holo plasma RBP4 | | Descriptor: | CHLORIDE ION, RETINOL, Retinol-binding protein 4 | | Authors: | Perduca, M, Monaco, H.L, Galliano, M. | | Deposit date: | 2017-04-28 | | Release date: | 2018-03-07 | | Last modified: | 2024-01-17 | | Method: | X-RAY DIFFRACTION (1.5 Å) | | Cite: | Human plasma retinol-binding protein (RBP4) is also a fatty acid-binding protein.
Biochim. Biophys. Acta, 1863, 2018
|
|
4IAW
 
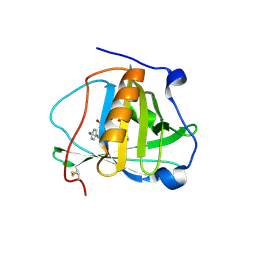 | | Engineered human lipocalin 2 (C26) in complex with Y-DTPA | | Descriptor: | N-{(1S,2S)-2-[bis(carboxymethyl)amino]cyclohexyl}-N-{(2R)-2-[bis(carboxymethyl)amino]-3-[4-({[2-hydroxy-1,1-bis(hydroxymethyl)ethyl]carbamothioyl}amino)phenyl]propyl}glycine, Neutrophil gelatinase-associated lipocalin, YTTRIUM (III) ION | | Authors: | Eichinger, A, Skerra, A. | | Deposit date: | 2012-12-07 | | Release date: | 2013-06-19 | | Last modified: | 2023-11-08 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | Structure-guided engineering of Anticalins with improved binding behavior and biochemical characteristics for application in radio-immuno imaging and/or therapy
J.Struct.Biol., 185, 2014
|
|
5NUK
 
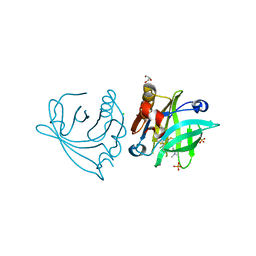 | | Engineered beta-lactoglobulin: variant I56F-L39A-M107F in complex with chlorpromazine (LG-FAF-CLP) | | Descriptor: | 3-(2-chloro-10H-phenothiazin-10-yl)-N,N-dimethylpropan-1-amine, Beta-lactoglobulin, CHLORIDE ION, ... | | Authors: | Loch, J.I, Bonarek, P, Tworzydlo, M, Lazinska, I, Szydlowska, J, Lewinski, K. | | Deposit date: | 2017-04-30 | | Release date: | 2018-04-04 | | Last modified: | 2024-01-31 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | The engineered beta-lactoglobulin with complementarity to the chlorpromazine chiral conformers.
Int. J. Biol. Macromol., 114, 2018
|
|
5OGB
 
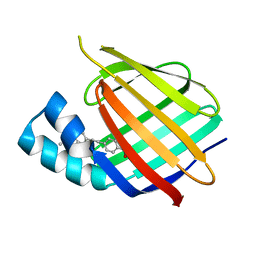 | | Human Cellular Retinoic Acid Binding Protein II (CRABPII) with bound synthetic retinoid DC360. | | Descriptor: | 4-[2-(4,4-dimethyl-1-propan-2-yl-quinolin-6-yl)ethynyl]benzoic acid, Cellular retinoic acid-binding protein 2 | | Authors: | Chisholm, D, Tomlinson, C, Whiting, A, Pohl, E. | | Deposit date: | 2017-07-12 | | Release date: | 2018-10-24 | | Last modified: | 2024-01-17 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Fluorescent Retinoic Acid Analogues as Probes for Biochemical and Intracellular Characterization of Retinoid Signaling Pathways.
Acs Chem.Biol., 14, 2019
|
|
4IAX
 
 | | Engineered human lipocalin 2 (CL31) in complex with Y-DTPA | | Descriptor: | N-{(1S,2S)-2-[bis(carboxymethyl)amino]cyclohexyl}-N-{(2R)-2-[bis(carboxymethyl)amino]-3-[4-({[2-hydroxy-1,1-bis(hydroxymethyl)ethyl]carbamothioyl}amino)phenyl]propyl}glycine, Neutrophil gelatinase-associated lipocalin, YTTRIUM (III) ION | | Authors: | Eichinger, A, Skerra, A. | | Deposit date: | 2012-12-07 | | Release date: | 2013-06-19 | | Last modified: | 2023-11-08 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Structure-guided engineering of Anticalins with improved binding behavior and biochemical characteristics for application in radio-immuno imaging and/or therapy
J.Struct.Biol., 185, 2014
|
|
4I3D
 | | Crystal structure of fluorescent protein UnaG N57A mutant | | Descriptor: | 3-[5-[(Z)-(4-ethenyl-3-methyl-5-oxidanylidene-pyrrol-2-ylidene)methyl]-2-[[5-[(Z)-(3-ethenyl-4-methyl-5-oxidanylidene-pyrrol-2-ylidene)methyl]-3-(3-hydroxy-3-oxopropyl)-4-methyl-1H-pyrrol-2-yl]methyl]-4-methyl-1H-pyrrol-3-yl]propanoic acid, Bilirubin-inducible fluorescent protein UnaG | | Authors: | Kumagai, A, Ando, R, Miyatake, H, Miyawaki, A. | | Deposit date: | 2012-11-26 | | Release date: | 2013-06-19 | | Last modified: | 2023-11-08 | | Method: | X-RAY DIFFRACTION (2.298 Å) | | Cite: | A bilirubin-inducible fluorescent protein from eel muscle
Cell(Cambridge,Mass.), 153, 2013
|
|
4I9S
 | |
4I9R
 | |
4IB7
 
 | | Bovine beta-lactoglobulin (isoform A) in complex with dodecyltrimethylammonium (DTAC) | | Descriptor: | DODECANE-TRIMETHYLAMINE, beta-lactoglobulin | | Authors: | Loch, J.I, Bonarek, P, Polit, A, Swiatek, S, Dziedzicka-Wasylewska, M, Lewinski, K. | | Deposit date: | 2012-12-08 | | Release date: | 2013-06-26 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | The differences in binding 12-carbon aliphatic ligands by bovine beta-lactoglobulin isoform A and B studied by isothermal titration calorimetry and X-ray crystallography
J.Mol.Recognit., 26, 2013
|
|
4IBA
 
 | | Bovine beta-lactoglobulin (isoform B) in complex with dodecyl sulphate (SDS) | | Descriptor: | DODECYL SULFATE, GLYCEROL, beta-lactoglobulin | | Authors: | Loch, J.I, Bonarek, P, Polit, A, Swiatek, S, Dziedzicka-Wasylewska, M, Lewinski, K. | | Deposit date: | 2012-12-08 | | Release date: | 2013-06-26 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | The differences in binding 12-carbon aliphatic ligands by bovine beta-lactoglobulin isoform A and B studied by isothermal titration calorimetry and X-ray crystallography
J.Mol.Recognit., 26, 2013
|
|
4IB6
 
 | | Bovine beta-lactoglobulin (isoform A) in complex with lauric acid (C12) | | Descriptor: | Beta-lactoglobulin, LAURIC ACID | | Authors: | Loch, J.I, Bonarek, P, Polit, A, Swiatek, S, Dziedzicka-Wasylewska, M, Lewinski, K. | | Deposit date: | 2012-12-08 | | Release date: | 2013-06-26 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | The differences in binding 12-carbon aliphatic ligands by bovine beta-lactoglobulin isoform A and B studied by isothermal titration calorimetry and X-ray crystallography
J.Mol.Recognit., 26, 2013
|
|
4IB9
 
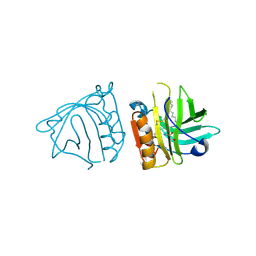 | | Bovine beta-lactoglobulin (isoform B) in complex with dodecyltrimethylammonium (DTAC) | | Descriptor: | DODECANE-TRIMETHYLAMINE, GLYCEROL, beta-lactoglobulin | | Authors: | Loch, J.I, Bonarek, P, Polit, A, Swiatek, S, Dziedzicka-Wasylewska, M, Lewinski, K. | | Deposit date: | 2012-12-08 | | Release date: | 2013-06-26 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | The differences in binding 12-carbon aliphatic ligands by bovine beta-lactoglobulin isoform A and B studied by isothermal titration calorimetry and X-ray crystallography
J.Mol.Recognit., 26, 2013
|
|
4IB8
 
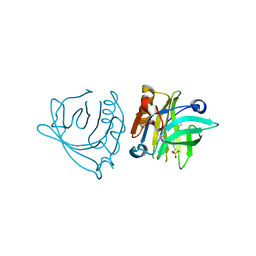 | | Bovine beta-lactoglobulin (isoform A) in complex with dodecyl sulphate (SDS) | | Descriptor: | DODECYL SULFATE, beta-lactoglobulin | | Authors: | Loch, J.I, Bonarek, P, Polit, A, Swiatek, S, Dziedzicka-Wasylewska, M, Lewinski, K. | | Deposit date: | 2012-12-08 | | Release date: | 2013-06-26 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | The differences in binding 12-carbon aliphatic ligands by bovine beta-lactoglobulin isoform A and B studied by isothermal titration calorimetry and X-ray crystallography
J.Mol.Recognit., 26, 2013
|
|
4IMO
 
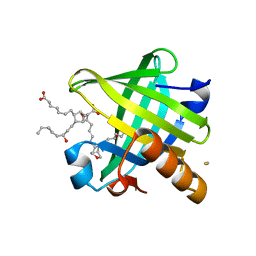 | | Crystal structure of wild type human Lipocalin PGDS in complex with substrate analog U44069 | | Descriptor: | (5E)-7-{(1R,4S,5S,6R)-5-[(1E,3S)-3-hydroxyoct-1-en-1-yl]-2-oxabicyclo[2.2.1]hept-6-yl}hept-5-enoic acid, Lipocalin-type prostaglandin-D synthase, THIOCYANATE ION | | Authors: | Lim, S.M, Chen, D, Teo, H, Roos, A, Nyman, T, Tresaugues, L, Pervushin, K, Nordlund, P. | | Deposit date: | 2013-01-03 | | Release date: | 2013-03-20 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (1.88 Å) | | Cite: | Structural and dynamic insights into substrate binding and catalysis of human lipocalin prostaglandin D synthase.
J.Lipid Res., 54, 2013
|
|
4IMN
 | | Crystal structure of wild type human Lipocalin PGDS bound with PEG MME 2000 | | Descriptor: | 2-(2-{2-[2-(2-METHOXY-ETHOXY)-ETHOXY]-ETHOXY}-ETHOXY)-ETHANOL, Lipocalin-type prostaglandin-D synthase | | Authors: | Lim, S.M, Chen, D, Teo, H, Roos, A, Nyman, T, Tresaugues, L, Pervushin, K, Nordlund, P. | | Deposit date: | 2013-01-03 | | Release date: | 2013-03-20 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (2.09 Å) | | Cite: | Structural and dynamic insights into substrate binding and catalysis of human lipocalin prostaglandin D synthase.
J.Lipid Res., 54, 2013
|
|
4K19
 
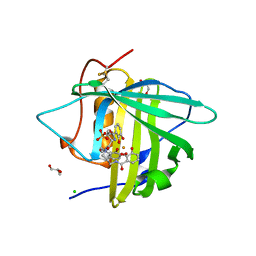 | | The structure of Human Siderocalin bound to the bacterial siderophore fluvibactin | | Descriptor: | (4S,5R)-N,N-bis{3-[(2,3-dihydroxybenzoyl)amino]propyl}-2-(2,3-dihydroxyphenyl)-5-methyl-4,5-dihydro-1,3-oxazole-4-carboxamide, CHLORIDE ION, FE (III) ION, ... | | Authors: | Correnti, C, Clifton, M.C, Strong, R.K. | | Deposit date: | 2013-04-04 | | Release date: | 2013-07-31 | | Last modified: | 2017-11-15 | | Method: | X-RAY DIFFRACTION (2.74 Å) | | Cite: | Siderocalin Outwits the Coordination Chemistry of Vibriobactin, a Siderophore of Vibrio cholerae.
Acs Chem.Biol., 8, 2013
|
|
4KII
 
 | | Beta-lactoglobulin in complex with Cp*Rh(III)Cl N,N-di(pyridin-2-yl)dodecanamide | | Descriptor: | Beta-lactoglobulin, [N,N-di(pyridin-2-yl-kappaN)dodecanamide]rhodium | | Authors: | Cherrier, M.V, Amara, P, Engilberge, S, Fontecilla-Camps, J.C. | | Deposit date: | 2013-05-02 | | Release date: | 2013-09-04 | | Last modified: | 2023-11-08 | | Method: | X-RAY DIFFRACTION (1.85 Å) | | Cite: | Structural Basis for Enantioselectivity in the Transfer Hydrogenation of a Ketone Catalyzed by an Artificial Metalloenzyme
Eur.J.Inorg.Chem., 21, 2013
|
|
4LKP
 
 | |
4LKT
 | |
1TVQ
 | | Crystal Structure of Apo Chicken Liver Basic Fatty Acid Binding Protein (or Bile Acid Binding Protein) | | Descriptor: | Fatty acid-binding protein | | Authors: | Nichesola, D, Perduca, M, Capaldi, S, Carrizo, M.E, Righetti, P.G, Monaco, H.L. | | Deposit date: | 2004-06-30 | | Release date: | 2004-11-23 | | Last modified: | 2024-03-13 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Crystal structure of chicken liver basic Fatty Acid-binding protein complexed with cholic acid
Biochemistry, 43, 2004
|
|
1UZ2
 
 | | The Cys121Ser Mutant of Beta-Lactoglobulin | | Descriptor: | BETA-LACTOGLOBULIN | | Authors: | McNae, I, Jayat, D, Haertle, T, Holt, C, Sawyer, L. | | Deposit date: | 2004-03-03 | | Release date: | 2004-05-20 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (1.95 Å) | | Cite: | A recombinant C121S mutant of bovine beta-lactoglobulin is more susceptible to peptic digestion and to denaturation by reducing agents and heating.
Biochemistry, 43, 2004
|
|
1URE
 | |
1VYF
 | | schistosoma mansoni fatty acid binding protein in complex with oleic acid | | Descriptor: | 14 KDA FATTY ACID BINDING PROTEIN, OLEIC ACID | | Authors: | Angelucci, F, Johnson, K.A, Baiocco, P, Miele, A.E, Bellelli, A. | | Deposit date: | 2004-04-29 | | Release date: | 2004-09-23 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (1.85 Å) | | Cite: | Schistosoma Mansoni Fatty Acid Binding Protein: Specificity and Functional Control as Revealed by Crystallographic Structure
Biochemistry, 43, 2004
|
|
1VYG
 | | schistosoma mansoni fatty acid binding protein in complex with arachidonic acid | | Descriptor: | ARACHIDONIC ACID, FATTY ACID BINDING PROTEIN | | Authors: | Angelucci, F, Johnson, K.A, Baiocco, P, Miele, A.E, Bellelli, A. | | Deposit date: | 2004-04-29 | | Release date: | 2004-09-23 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | Schistosoma Mansoni Fatty Acid Binding Protein: Specificity and Functional Control as Revealed by Crystallographic Structure
Biochemistry, 43, 2004
|
|
1X8U
 | | Crystal structure of Siderocalin (NGAL, Lipocalin 2) complexed with Carboxymycobactin T | | Descriptor: | CARBOXYMYCOBACTIN T, Neutrophil gelatinase-associated lipocalin | | Authors: | Holmes, M.A, Paulsene, W, Jide, X, Ratledge, C, Strong, R.K. | | Deposit date: | 2004-08-18 | | Release date: | 2005-01-25 | | Last modified: | 2024-04-03 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Siderocalin (Lcn 2) Also Binds Carboxymycobactins, Potentially Defending against Mycobacterial Infections through Iron Sequestration
Structure, 13, 2005
|
|
