3KTU
 
 | |
3UBT
 
 | |
3IH7
 
 | |
1CW9
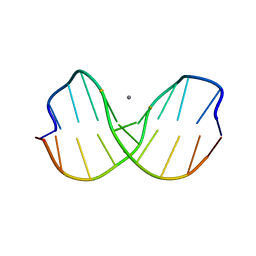 
 | | DNA DECAMER WITH AN ENGINEERED CROSSLINK IN THE MINOR GROOVE | | Descriptor: | 5'-D(*CP*CP*AP*GP*(G47)P*CP*CP*TP*GP*G)-3', CALCIUM ION | | Authors: | van Aalten, D.M.F, Erlanson, D.A, Verdine, G.L, Joshua-Tor, L. | | Deposit date: | 1999-08-26 | | Release date: | 1999-10-14 | | Last modified: | 2024-04-03 | | Method: | X-RAY DIFFRACTION (1.55 Å) | | Cite: | A structural snapshot of base-pair opening in DNA.
Proc.Natl.Acad.Sci.USA, 96, 1999
|
|
5KN9
 
 | | MutY N-terminal domain in complex with DNA containing an intrahelical oxoG:A base-pair | | Descriptor: | Adenine DNA glycosylase, CALCIUM ION, DNA (5'-D(*AP*GP*CP*AP*CP*AP*GP*GP*AP*T)-3'), ... | | Authors: | Wang, L, Chakravarthy, S, Verdine, G.L. | | Deposit date: | 2016-06-27 | | Release date: | 2017-02-08 | | Last modified: | 2023-09-27 | | Method: | X-RAY DIFFRACTION (1.93 Å) | | Cite: | Structural Basis for the Lesion-scanning Mechanism of the MutY DNA Glycosylase.
J. Biol. Chem., 292, 2017
|
|
5KN8
 
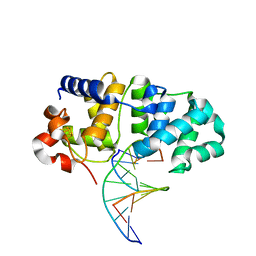 | | MutY N-terminal domain in complex with undamaged DNA | | Descriptor: | Adenine DNA glycosylase, CALCIUM ION, DNA (5'-D(*AP*GP*CP*AP*CP*AP*GP*GP*AP*T)-3'), ... | | Authors: | Wang, L, Chakravarthy, S, Verdine, G.L. | | Deposit date: | 2016-06-27 | | Release date: | 2017-02-08 | | Last modified: | 2023-09-27 | | Method: | X-RAY DIFFRACTION (1.81 Å) | | Cite: | Structural Basis for the Lesion-scanning Mechanism of the MutY DNA Glycosylase.
J. Biol. Chem., 292, 2017
|
|
4G4Q
 
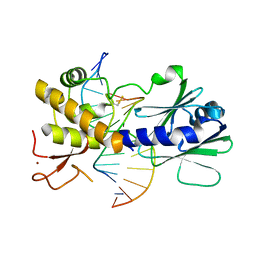 | | MutM containing F114A mutation bound to undamaged DNA | | Descriptor: | DNA (5'-D(*AP*GP*GP*TP*AP*GP*AP*CP*TP*CP*GP*GP*AP*CP*GP*C)-3'), DNA (5'-D(*TP*GP*CP*GP*TP*CP*CP*GP*AP*GP*(TX2)P*CP*TP*AP*CP*C)-3'), Formamidopyrimidine-DNA glycosylase, ... | | Authors: | Sung, R.J, Zhang, M, Qi, Y, Verdine, G.L. | | Deposit date: | 2012-07-16 | | Release date: | 2013-02-20 | | Last modified: | 2013-04-24 | | Method: | X-RAY DIFFRACTION (1.86 Å) | | Cite: | Structural and Biochemical Analysis of DNA Helix Invasion by the Bacterial 8-Oxoguanine DNA Glycosylase MutM.
J.Biol.Chem., 288, 2013
|
|
1ADN
 
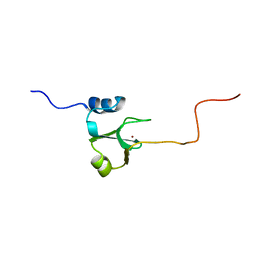 | |
6O8E
 
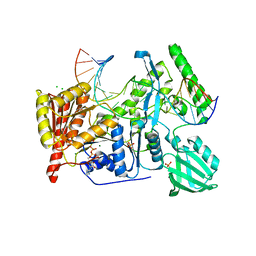 | | Crystal structure of UvrB bound to duplex DNA with ADP | | Descriptor: | ADENOSINE-5'-DIPHOSPHATE, CHLORIDE ION, DNA (5'-D(*GP*CP*CP*GP*TP*AP*TP*GP*CP*CP*AP*AP*TP*CP*TP*AP*GP*AP*GP*C)-3'), ... | | Authors: | Lee, S.-J, Verdine, G.L. | | Deposit date: | 2019-03-10 | | Release date: | 2020-01-22 | | Last modified: | 2024-03-13 | | Method: | X-RAY DIFFRACTION (2.61 Å) | | Cite: | Mechanism of DNA Lesion Homing and Recognition by the Uvr Nucleotide Excision Repair System.
Res, 2019, 2019
|
|
6O8G
 
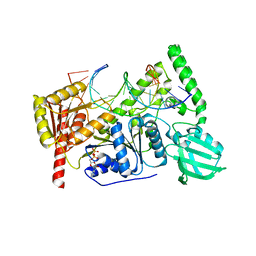 | | Crystal structure of UvrB bound to fully duplex DNA | | Descriptor: | ADENOSINE-5'-DIPHOSPHATE, CHLORIDE ION, DNA (5'-D(*GP*GP*TP*AP*GP*CP*GP*CP*GP*AP*TP*GP*GP*AP*GP*A)-3'), ... | | Authors: | Lee, S.-J, Sung, R.-J, Verdine, G.L. | | Deposit date: | 2019-03-10 | | Release date: | 2020-01-22 | | Last modified: | 2024-03-13 | | Method: | X-RAY DIFFRACTION (2.64 Å) | | Cite: | Mechanism of DNA Lesion Homing and Recognition by the Uvr Nucleotide Excision Repair System.
Res, 2019, 2019
|
|
6O8H
 
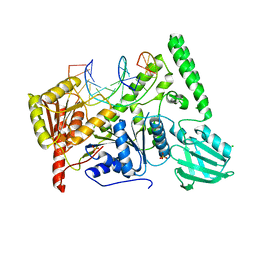 | | Crystal structure of UvrB mutant bound to duplex DNA | | Descriptor: | CHLORIDE ION, DNA (5'-D(P*AP*GP*CP*GP*CP*GP*AP*TP*GP*GP*AP*GP*A)-3'), DNA (5'-D(P*CP*CP*AP*TP*CP*GP*CP*GP*CP*TP*AP*CP*C)-3'), ... | | Authors: | Lee, S.-J, Verdine, G.L. | | Deposit date: | 2019-03-10 | | Release date: | 2020-01-22 | | Last modified: | 2024-03-13 | | Method: | X-RAY DIFFRACTION (2.39 Å) | | Cite: | Mechanism of DNA Lesion Homing and Recognition by the Uvr Nucleotide Excision Repair System.
Res, 2019, 2019
|
|
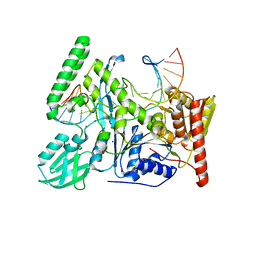
6O8F
 
 | | Crystal structure of UvrB bound to duplex DNA | | Descriptor: | ACETATE ION, CHLORIDE ION, DNA (5'-D(*GP*CP*CP*GP*TP*AP*TP*GP*CP*CP*AP*AP*TP*CP*TP*AP*GP*AP*GP*C)-3'), ... | | Authors: | Lee, S.-J, Verdine, G.L. | | Deposit date: | 2019-03-10 | | Release date: | 2020-01-22 | | Last modified: | 2024-03-13 | | Method: | X-RAY DIFFRACTION (2.81 Å) | | Cite: | Mechanism of DNA Lesion Homing and Recognition by the Uvr Nucleotide Excision Repair System.
Res, 2019, 2019
|
|
6OQA
 
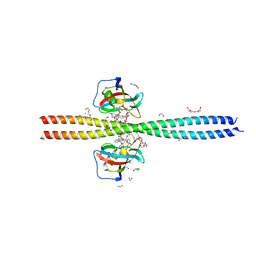 | | Crystal structure of CEP250 bound to FKBP12 in the presence of FK506-like novel natural product | | Descriptor: | (3R,4E,7E,10R,11S,12R,13S,16R,17R,24aS)-11,17-dihydroxy-10,12,16-trimethyl-3-[(2R)-1-phenylbutan-2-yl]-6,9,10,11,12,13,14,15,16,17,22,23,24,24a-tetradecahydro-3H-13,17-epoxypyrido[2,1-c][1,4]oxazacyclohenicosine-1,18,19(21H)-trione, 1,2-ETHANEDIOL, 3,6,9,12,15,18,21-HEPTAOXATRICOSANE-1,23-DIOL, ... | | Authors: | Lee, S.-J, Shigdel, U.K, Townson, S.A, Verdine, G.L. | | Deposit date: | 2019-04-26 | | Release date: | 2020-04-29 | | Last modified: | 2024-03-13 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Genomic discovery of an evolutionarily programmed modality for small-molecule targeting of an intractable protein surface.
Proc.Natl.Acad.Sci.USA, 117, 2020
|
|
4YPR
 
 | | Crystal Structure of D144N MutY bound to its anti-substrate | | Descriptor: | A/G-specific adenine glycosylase, DNA (5'-D(*AP*AP*GP*AP*CP*(8OG)P*TP*GP*GP*AP*C)-3'), DNA (5'-D(*T*GP*TP*CP*CP*AP*CP*GP*TP*CP*T)-3'), ... | | Authors: | Wang, L, Lee, S, Verdine, G.L. | | Deposit date: | 2015-03-13 | | Release date: | 2015-05-27 | | Last modified: | 2023-09-27 | | Method: | X-RAY DIFFRACTION (2.59 Å) | | Cite: | Structural Basis for Avoidance of Promutagenic DNA Repair by MutY Adenine DNA Glycosylase.
J.Biol.Chem., 290, 2015
|
|
4YOQ
 
 | | Crystal Structure of MutY bound to its anti-substrate | | Descriptor: | A/G-specific adenine glycosylase, DNA (5'-D(*AP*AP*GP*AP*CP*(8OG)P*TP*GP*GP*AP*C)-3'), DNA (5'-D(*T*GP*TP*CP*CP*AP*CP*GP*TP*CP*T)-3'), ... | | Authors: | Wang, L, Lee, S, Verdine, G.L. | | Deposit date: | 2015-03-11 | | Release date: | 2015-05-27 | | Last modified: | 2023-09-27 | | Method: | X-RAY DIFFRACTION (2.21 Å) | | Cite: | Structural Basis for Avoidance of Promutagenic DNA Repair by MutY Adenine DNA Glycosylase.
J.Biol.Chem., 290, 2015
|
|
4YPH
 
 | | Crystal Structure of MutY bound to its anti-substrate with the disulfide cross-linker reduced | | Descriptor: | A/G-specific adenine glycosylase, DNA (5'-D(*AP*AP*GP*AP*CP*(8OG)P*TP*GP*GP*AP*C)-3'), DNA (5'-D(TP*GP*TP*CP*CP*AP*CP*GP*TP*CP*T)-3'), ... | | Authors: | Wang, L, Lee, S, Verdine, G.L. | | Deposit date: | 2015-03-12 | | Release date: | 2015-05-27 | | Last modified: | 2023-09-27 | | Method: | X-RAY DIFFRACTION (2.32 Å) | | Cite: | Structural Basis for Avoidance of Promutagenic DNA Repair by MutY Adenine DNA Glycosylase.
J.Biol.Chem., 290, 2015
|
|
4DJS
 
 | | Structure of beta-catenin in complex with a stapled peptide inhibitor | | Descriptor: | Catenin beta-1, stapled peptide RRWPQ(MK8)ILD(MK8)HVRRVWR | | Authors: | Bowman, B.R, Grossmann, T.N, Yeh, J.T.-H, Verdine, G.L. | | Deposit date: | 2012-02-02 | | Release date: | 2012-10-17 | | Last modified: | 2019-07-17 | | Method: | X-RAY DIFFRACTION (3.029 Å) | | Cite: | Inhibition of oncogenic Wnt signaling through direct targeting of beta-catenin.
Proc.Natl.Acad.Sci.USA, 109, 2012
|
|
4G4O
 
 | | MutM containing M77A mutation bound to oxoG-containing DNA | | Descriptor: | DNA (5'-D(*TP*GP*CP*GP*TP*CP*CP*(8OG)P*AP*GP*(TX2)P*CP*TP*AP*CP*C)-3'), DNA (5'-D(P*AP*GP*GP*TP*AP*GP*AP*CP*TP*CP*GP*GP*AP*CP*GP*C)-3'), Formamidopyrimidine-DNA glycosylase, ... | | Authors: | Sung, R.J, Zhang, M, Qi, Y, Verdine, G.L. | | Deposit date: | 2012-07-16 | | Release date: | 2013-02-20 | | Last modified: | 2013-04-24 | | Method: | X-RAY DIFFRACTION (1.95 Å) | | Cite: | Structural and Biochemical Analysis of DNA Helix Invasion by the Bacterial 8-Oxoguanine DNA Glycosylase MutM.
J.Biol.Chem., 288, 2013
|
|
4G4R
 
 | | MutM containing F114A mutation bound to oxoG-containing DNA | | Descriptor: | DNA (5'-D(*TP*GP*CP*GP*TP*CP*CP*(8OG)P*AP*GP*(TX2)P*CP*TP*AP*CP*C)-3'), DNA (5'-D(P*AP*GP*GP*TP*AP*GP*AP*CP*TP*CP*GP*GP*AP*CP*GP*C)-3'), Formamidopyrimidine-DNA glycosylase, ... | | Authors: | Sung, R.J, Zhang, M, Qi, Y, Verdine, G.L. | | Deposit date: | 2012-07-16 | | Release date: | 2013-02-20 | | Last modified: | 2013-04-24 | | Method: | X-RAY DIFFRACTION (1.95 Å) | | Cite: | Structural and Biochemical Analysis of DNA Helix Invasion by the Bacterial 8-Oxoguanine DNA Glycosylase MutM.
J.Biol.Chem., 288, 2013
|
|
4G4N
 
 | | MutM containing M77A mutation bound to undamaged DNA | | Descriptor: | DNA (5'-D(*TP*GP*CP*GP*TP*CP*CP*GP*AP*GP*(TX2)P*CP*TP*AP*CP*C)-3'), DNA (5'-D(P*AP*GP*GP*TP*AP*GP*AP*CP*TP*CP*GP*GP*AP*CP*GP*C)-3'), Formamidopyrimidine-DNA glycosylase, ... | | Authors: | Sung, R.J, Zhang, M, Qi, Y, Verdine, G.L. | | Deposit date: | 2012-07-16 | | Release date: | 2013-02-20 | | Last modified: | 2013-11-20 | | Method: | X-RAY DIFFRACTION (1.85 Å) | | Cite: | Structural and Biochemical Analysis of DNA Helix Invasion by the Bacterial 8-Oxoguanine DNA Glycosylase MutM.
J.Biol.Chem., 288, 2013
|
|
1DCT
 
 | | DNA (CYTOSINE-5) METHYLASE FROM HAEIII COVALENTLY BOUND TO DNA | | Descriptor: | CALCIUM ION, DNA (5'-D(*AP*CP*CP*AP*GP*CP*AP*GP*GP*(C49)P*CP*AP*CP*CP*AP*GP*TP*G)-3'), DNA (5'-D(*TP*CP*AP*CP*TP*GP*GP*TP*GP*GP*(C5M)P*CP*TP*GP*CP*TP*GP*G)-3'), ... | | Authors: | Reinisch, K.M, Chen, L, Verdine, G.L, Lipscomb, W.N. | | Deposit date: | 1995-05-17 | | Release date: | 1995-09-15 | | Last modified: | 2019-11-20 | | Method: | X-RAY DIFFRACTION (2.8 Å) | | Cite: | The crystal structure of HaeIII methyltransferase convalently complexed to DNA: an extrahelical cytosine and rearranged base pairing.
Cell(Cambridge,Mass.), 82, 1995
|
|
5WHD
 
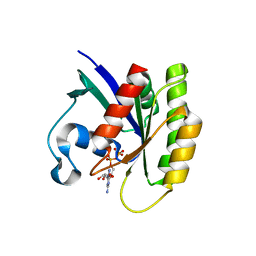 | | Crystal structure of KRas G12V/D38P, bound to GDP | | Descriptor: | GTPase KRas, GUANOSINE-5'-DIPHOSPHATE | | Authors: | Shim, S.Y, McGee, J.H, Lee, S.-J, Verdine, G.L. | | Deposit date: | 2017-07-16 | | Release date: | 2018-01-03 | | Last modified: | 2023-10-04 | | Method: | X-RAY DIFFRACTION (1.641 Å) | | Cite: | Exceptionally high-affinity Ras binders that remodel its effector domain.
J. Biol. Chem., 293, 2018
|
|
5WPM
 
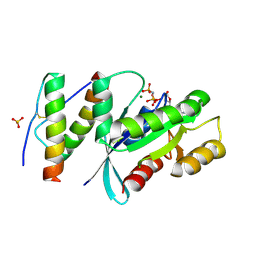 | | KRas G12V, bound to GppNHp and miniprotein 225-11(A30R) | | Descriptor: | GTPase KRas, MAGNESIUM ION, PHOSPHOAMINOPHOSPHONIC ACID-GUANYLATE ESTER, ... | | Authors: | Lee, S.-J, Shim, S.Y, McGee, J.H, Verdine, G.L. | | Deposit date: | 2017-08-05 | | Release date: | 2018-01-03 | | Last modified: | 2018-03-14 | | Method: | X-RAY DIFFRACTION (1.72 Å) | | Cite: | Exceptionally high-affinity Ras binders that remodel its effector domain.
J. Biol. Chem., 293, 2018
|
|
5WHA
 
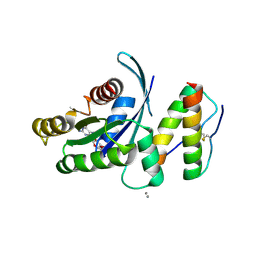 | | KRas G12V, bound to GDP and miniprotein 225-11 | | Descriptor: | CALCIUM ION, GTPase KRas, GUANOSINE-5'-DIPHOSPHATE, ... | | Authors: | Shim, S.Y, McGee, J.H, Lee, S.-J, Verdine, G.L. | | Deposit date: | 2017-07-16 | | Release date: | 2018-01-03 | | Last modified: | 2023-10-04 | | Method: | X-RAY DIFFRACTION (2.04 Å) | | Cite: | Exceptionally high-affinity Ras binders that remodel its effector domain.
J. Biol. Chem., 293, 2018
|
|
5WHB
 
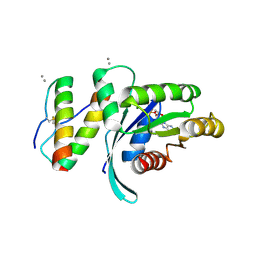 | | KRas G12V, bound to GDP and miniprotein 225-11(A30R) | | Descriptor: | CALCIUM ION, GTPase KRas, GUANOSINE-5'-DIPHOSPHATE, ... | | Authors: | Shim, S.Y, McGee, J.H, Lee, S.-J, Verdine, G.L. | | Deposit date: | 2017-07-16 | | Release date: | 2018-01-03 | | Last modified: | 2024-04-03 | | Method: | X-RAY DIFFRACTION (2.18 Å) | | Cite: | Exceptionally high-affinity Ras binders that remodel its effector domain.
J. Biol. Chem., 293, 2018
|
|
