3VZS
 
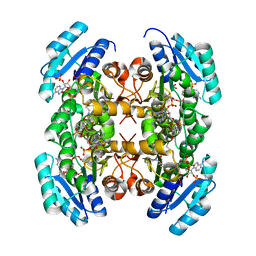 | | Crystal structure of PhaB from Ralstonia eutropha in complex with Acetoacetyl-CoA and NADP | | Descriptor: | ACETOACETYL-COENZYME A, Acetoacetyl-CoA reductase, NADP NICOTINAMIDE-ADENINE-DINUCLEOTIDE PHOSPHATE, ... | | Authors: | Ikeda, K, Tanaka, Y, Tanaka, I, Yao, M. | | Deposit date: | 2012-10-15 | | Release date: | 2013-08-28 | | Last modified: | 2023-11-08 | | Method: | X-RAY DIFFRACTION (2.14 Å) | | Cite: | Directed evolution and structural analysis of NADPH-dependent Acetoacetyl Coenzyme A (Acetoacetyl-CoA) reductase from Ralstonia eutropha reveals two mutations responsible for enhanced kinetics
Appl.Environ.Microbiol., 79, 2013
|
|
3W9V
 
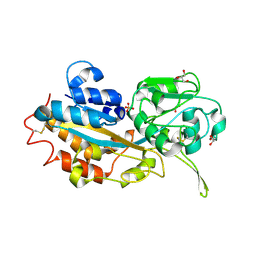 | | Crystal structure of refolded DING protein | | Descriptor: | GLYCEROL, PHOSPHATE ION, Phosphate-binding protein | | Authors: | Gai, Z.Q, Nakamura, A, Tanaka, Y, Hirano, N, Tanaka, I, Yao, M. | | Deposit date: | 2013-04-17 | | Release date: | 2013-10-30 | | Last modified: | 2023-11-08 | | Method: | X-RAY DIFFRACTION (1.031 Å) | | Cite: | Crystal structure analysis, overexpression and refolding behaviour of a DING protein with single mutation.
J.SYNCHROTRON RADIAT., 20, 2013
|
|
7EQE
 
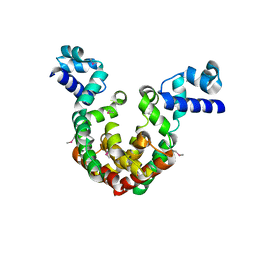 | | Crystal Structure of a transcription factor | | Descriptor: | TetR/AcrR family transcriptional regulator | | Authors: | Uehara, S, Tsugita, A, Matsui, T, Yokoyama, T, Ostash, I, Ostash, B, Tanaka, Y. | | Deposit date: | 2021-05-01 | | Release date: | 2022-04-27 | | Last modified: | 2022-10-19 | | Method: | X-RAY DIFFRACTION (2.399 Å) | | Cite: | The carbohydrate tail of landomycin A is responsible for its interaction with the repressor protein LanK.
Febs J., 289, 2022
|
|
3W9W
 
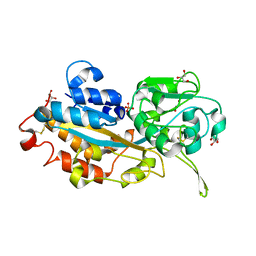 | | Crystal structure of DING protein | | Descriptor: | DING protein, GLYCEROL, PHOSPHATE ION | | Authors: | Gai, Z.Q, Nakamura, A, Tanaka, Y, Hirano, N, Tanaka, I, Yao, M. | | Deposit date: | 2013-04-17 | | Release date: | 2013-10-30 | | Last modified: | 2023-11-08 | | Method: | X-RAY DIFFRACTION (1.35 Å) | | Cite: | Crystal structure analysis, overexpression and refolding behaviour of a DING protein with single mutation.
J.SYNCHROTRON RADIAT., 20, 2013
|
|
2E8H
 
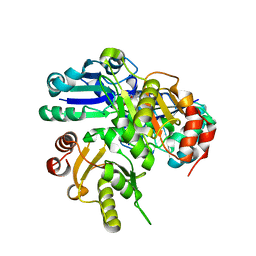 | | Crystal structure of PH0725 from Pyrococcus horikoshii OT3 | | Descriptor: | Probable diphthine synthase, S-ADENOSYL-L-HOMOCYSTEINE, SODIUM ION | | Authors: | Sugahara, M, Taketa, M, Tanaka, Y, Matsuura, Y, Kunishima, N, RIKEN Structural Genomics/Proteomics Initiative (RSGI) | | Deposit date: | 2007-01-19 | | Release date: | 2007-07-24 | | Last modified: | 2023-10-25 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Crystal structure of PH0725 from Pyrococcus horikoshii OT3
To be Published
|
|
2E27
 
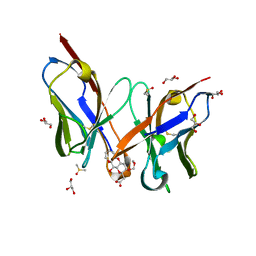 | | Crystal structure of Fv fragment of anti-ciguatoxin antibody complexed with ABC-ring of ciguatoxin | | Descriptor: | 3,7:6,10:9,14-TRIANHYDRO-2,5,11,12,13-PENTADEOXY-4-O-(METHOXYMETHYL)-L-ARABINO-L-ALLO-TETRADEC-12-ENONIC ACID, DIMETHYL SULFOXIDE, GLYCEROL, ... | | Authors: | Yokota, A, Tanaka, Y, Tsumoto, K. | | Deposit date: | 2006-11-08 | | Release date: | 2007-11-13 | | Last modified: | 2023-10-25 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | Critical contribution of aromatic rings to specific recognition of polyether rings. The case of ciguatoxin CTX3C-ABC and its specific antibody 1C49.
J.Biol.Chem., 283, 2008
|
|
2P6O
 
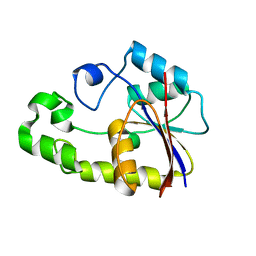 | | Crystal structure of TTHB049 from Thermus thermophilus HB8 | | Descriptor: | Alpha-ribazole-5'-phosphate phosphatase | | Authors: | Yamamoto, H, Taketa, M, Tanaka, Y, Matsuura, Y, Kunishima, N, RIKEN Structural Genomics/Proteomics Initiative (RSGI) | | Deposit date: | 2007-03-19 | | Release date: | 2007-09-25 | | Last modified: | 2023-10-25 | | Method: | X-RAY DIFFRACTION (1.65 Å) | | Cite: | Crystal structure of TTHB049 from Thermus thermophilus HB8
To be Published
|
|
2PCM
 
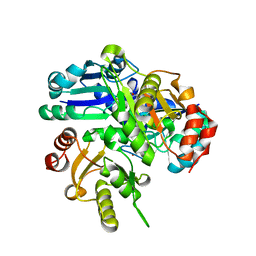 | | Crystal structure of PH0725 from Pyrococcus horikoshii OT3 | | Descriptor: | DIPHTHINE SYNTHASE, GLYCEROL, S-ADENOSYL-L-HOMOCYSTEINE | | Authors: | Yamamoto, H, Taketa, M, Tanaka, Y, Matsuura, Y, Kunishima, N, RIKEN Structural Genomics/Proteomics Initiative (RSGI) | | Deposit date: | 2007-03-30 | | Release date: | 2007-10-02 | | Last modified: | 2023-10-25 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | Crystal structure of PH0725 from Pyrococcus horikoshii OT3
To be Published
|
|
3VV6
 
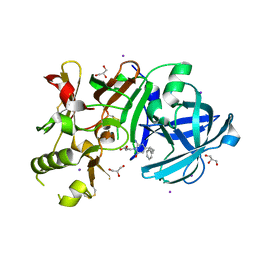 | | Crystal Structure of beta secetase in complex with 2-amino-3-methyl-6-((1S, 2R)-2-phenylcyclopropyl)pyrimidin-4(3H)-one | | Descriptor: | 2-amino-3-methyl-6-[(1S,2R)-2-phenylcyclopropyl]pyrimidin-4(3H)-one, Beta-secretase 1, GLYCEROL, ... | | Authors: | Yonezawa, S, Yamamoto, T, Yamakawa, H, Muto, C, Hosono, M, Hattori, K, Higashino, K, Sakagami, M, Togame, H, Tanaka, Y, Nakano, T, Takemoto, H, Arisawa, M, Shuto, S. | | Deposit date: | 2012-07-17 | | Release date: | 2012-10-24 | | Last modified: | 2017-11-22 | | Method: | X-RAY DIFFRACTION (2.05 Å) | | Cite: | Conformational restriction approach to beta-secretase (BACE1) inhibitors: effect of a cyclopropane ring to induce an alternative binding mode
J.Med.Chem., 55, 2012
|
|
3VV8
 
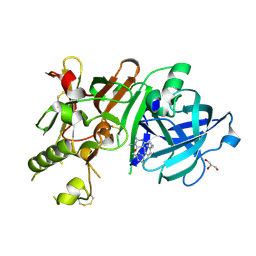 | | Crystal structure of beta secetase in complex with 2-amino-3-methyl-6-((1S,2R)-2-(3'-methylbiphenyl-4-yl)cyclopropyl)pyrimidin-4(3H)-one | | Descriptor: | 2-amino-3-methyl-6-[(1S,2R)-2-(3'-methylbiphenyl-4-yl)cyclopropyl]pyrimidin-4(3H)-one, Beta-secretase 1, GLYCEROL | | Authors: | Yonezawa, S, Yamamoto, T, Yamakawa, H, Muto, C, Hosono, M, Hattori, K, Higashino, K, Sakagami, M, Togame, H, Tanaka, Y, Nakano, T, Takemoto, H, Arisawa, M, Shuto, S. | | Deposit date: | 2012-07-17 | | Release date: | 2012-10-24 | | Last modified: | 2013-09-04 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | Conformational restriction approach to beta-secretase (BACE1) inhibitors: effect of a cyclopropane ring to induce an alternative binding mode.
J.Med.Chem., 55, 2012
|
|
3VV7
 
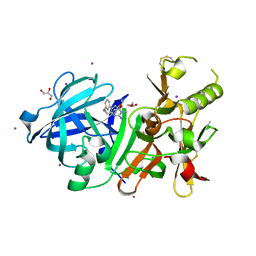 | | Crystal Structure of beta secetase in complex with 2-amino-6-((1S,2R)-2-(3'-methoxybiphenyl-3-yl)cyclopropyl)-3-methylpyrimidin-4(3H)-one | | Descriptor: | 2-amino-6-[(1S,2R)-2-(3'-methoxybiphenyl-3-yl)cyclopropyl]-3-methylpyrimidin-4(3H)-one, Beta-secretase 1, GLYCEROL, ... | | Authors: | Yonezawa, S, Yamamoto, T, Yamakawa, H, Muto, C, Hosono, M, Hattori, K, Higashino, K, Sakagami, M, Togame, H, Tanaka, Y, Nakano, T, Takemoto, H, Arisawa, M, Shuto, S. | | Deposit date: | 2012-07-17 | | Release date: | 2012-10-24 | | Last modified: | 2013-09-04 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Conformational restriction approach to beta-secretase (BACE1) inhibitors: effect of a cyclopropane ring to induce an alternative binding mode.
J.Med.Chem., 55, 2012
|
|
2CUO
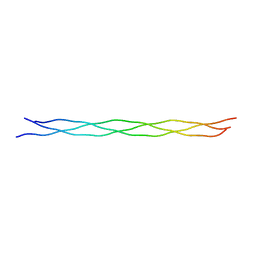 
 | | Collagen model peptide (PRO-PRO-GLY)9 | | Descriptor: | COLLAGEN MODEL PEPTIDE (PRO-PRO-GLY)9 | | Authors: | Hongo, C, Noguchi, K, Okuyama, K, Tanaka, Y, Nishino, N. | | Deposit date: | 2005-05-27 | | Release date: | 2005-06-14 | | Last modified: | 2023-10-25 | | Method: | X-RAY DIFFRACTION (1.33 Å) | | Cite: | Repetitive interactions observed in the crystal structure of a collagen-model peptide, [(Pro-Pro-Gly)9]3
J.Biochem.(Tokyo), 138, 2005
|
|
6AL3
 
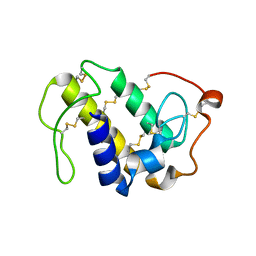 | | Lys49 PLA2 BPII derived from the venom of Protobothrops flavoviridis. | | Descriptor: | Basic phospholipase A2 BP-II, SULFATE ION | | Authors: | Matsui, T, Kamata, S, Suzuki, A, Oda-Ueda, N, Ogawa, T, Tanaka, Y. | | Deposit date: | 2018-09-05 | | Release date: | 2019-01-16 | | Last modified: | 2023-11-22 | | Method: | X-RAY DIFFRACTION (2.57 Å) | | Cite: | SDS-induced oligomerization of Lys49-phospholipase A2from snake venom.
Sci Rep, 9, 2019
|
|
2F08
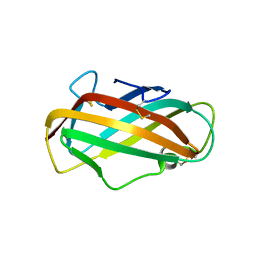 
 | | Crystal structure of a major house dust mite allergen, Derf 2 | | Descriptor: | O-ACETALDEHYDYL-HEXAETHYLENE GLYCOL, mite allergen Der f II | | Authors: | Mikami, B, Tanaka, Y, Minato, N, Suzuki, M, Korematsu, S. | | Deposit date: | 2005-11-12 | | Release date: | 2005-11-29 | | Last modified: | 2011-07-13 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Crystal structure and some properties of a major house dust mite allergen, Derf 2
Biochem.Biophys.Res.Commun., 339, 2006
|
|
5B5O
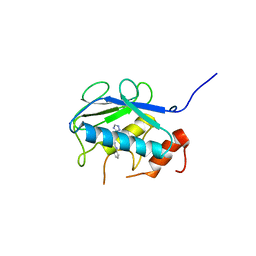 
 | | Crystal structure of the catalytic domain of MMP-13 complexed with N-phenyl-4-((4H-1,2,4-triazol-3-ylsulfanyl)methyl)-1,3-thiazol-2-amine | | Descriptor: | 1,2-ETHANEDIOL, CALCIUM ION, Collagenase 3, ... | | Authors: | Oki, H, Tanaka, Y. | | Deposit date: | 2016-05-13 | | Release date: | 2017-01-18 | | Last modified: | 2023-11-08 | | Method: | X-RAY DIFFRACTION (1.2 Å) | | Cite: | Discovery of Novel, Highly Potent, and Selective Matrix Metalloproteinase (MMP)-13 Inhibitors with a 1,2,4-Triazol-3-yl Moiety as a Zinc Binding Group Using a Structure-Based Design Approach
J. Med. Chem., 60, 2017
|
|
5B5P
 
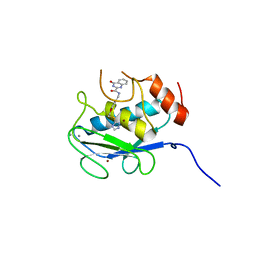 | | Crystal structure of the catalytic domain of MMP-13 complexed with 4-oxo-N-(3-(2-(1H-1,2,4-triazol-3-ylsulfanyl)ethoxy)benzyl)-3,4-dihydroquinazoline-2-carboxamide | | Descriptor: | 4-oxo-N-{3-[2-(1H-1,2,4-triazol-3-ylsulfanyl)ethoxy]benzyl}-3,4-dihydroquinazoline-2-carboxamide, CALCIUM ION, Collagenase 3, ... | | Authors: | Oki, H, Tanaka, Y. | | Deposit date: | 2016-05-13 | | Release date: | 2017-01-18 | | Last modified: | 2023-11-08 | | Method: | X-RAY DIFFRACTION (1.6 Å) | | Cite: | Discovery of Novel, Highly Potent, and Selective Matrix Metalloproteinase (MMP)-13 Inhibitors with a 1,2,4-Triazol-3-yl Moiety as a Zinc Binding Group Using a Structure-Based Design Approach
J. Med. Chem., 60, 2017
|
|
2YVX
 
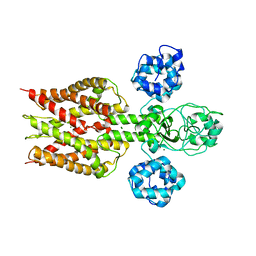 | | Crystal structure of magnesium transporter MgtE | | Descriptor: | MAGNESIUM ION, Mg2+ transporter MgtE | | Authors: | Hattori, M, Tanaka, Y, Fukai, S, Ishitani, R, Nureki, O. | | Deposit date: | 2007-04-18 | | Release date: | 2007-09-04 | | Last modified: | 2024-03-13 | | Method: | X-RAY DIFFRACTION (3.5 Å) | | Cite: | Crystal structure of the MgtE Mg(2+) transporter
Nature, 448, 2007
|
|
2YVZ
 
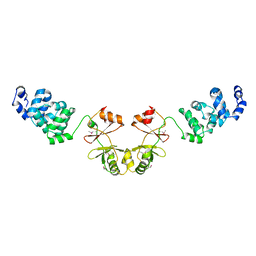 | | Crystal structure of magnesium transporter MgtE cytosolic domain, Mg2+-free form | | Descriptor: | Mg2+ transporter MgtE | | Authors: | Hattori, M, Tanaka, Y, Fukai, S, Ishitani, R, Nureki, O. | | Deposit date: | 2007-04-18 | | Release date: | 2007-09-04 | | Last modified: | 2011-07-13 | | Method: | X-RAY DIFFRACTION (3.9 Å) | | Cite: | Crystal structure of the MgtE Mg(2+) transporter
Nature, 448, 2007
|
|
2YVY
 
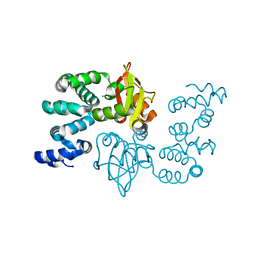 | | Crystal structure of magnesium transporter MgtE cytosolic domain, Mg2+ bound form | | Descriptor: | MAGNESIUM ION, Mg2+ transporter MgtE | | Authors: | Hattori, M, Tanaka, Y, Fukai, S, Ishitani, R, Nureki, O. | | Deposit date: | 2007-04-18 | | Release date: | 2007-09-04 | | Last modified: | 2024-03-13 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Crystal structure of the MgtE Mg(2+) transporter
Nature, 448, 2007
|
|
7WAX
 
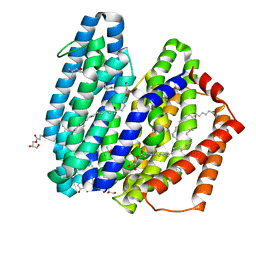 | | MurJ inward occluded form | | Descriptor: | (2R)-2,3-dihydroxypropyl (9Z)-octadec-9-enoate, (4S)-2-METHYL-2,4-PENTANEDIOL, lipid II flippase MurJ | | Authors: | Tsukazaki, T, Kohga, H, Tanaka, Y, Yoshikaie, K, Taniguchi, K, Fujimoto, K. | | Deposit date: | 2021-12-15 | | Release date: | 2022-06-01 | | Last modified: | 2022-08-17 | | Method: | X-RAY DIFFRACTION (2.35 Å) | | Cite: | Crystal structure of the lipid flippase MurJ in a "squeezed" form distinct from its inward- and outward-facing forms.
Structure, 30, 2022
|
|
7WAG
 
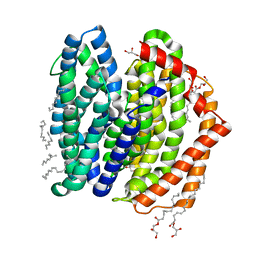 | | Crystal structure of MurJ squeezed form | | Descriptor: | (2R)-2,3-dihydroxypropyl (9Z)-octadec-9-enoate, DI(HYDROXYETHYL)ETHER, Lipid II flippase MurJ | | Authors: | Tsukazaki, T, Kohga, H, Tanaka, Y, Yoshikaie, K, Taniguchi, K, Fujimoto, K. | | Deposit date: | 2021-12-14 | | Release date: | 2022-06-01 | | Last modified: | 2023-11-29 | | Method: | X-RAY DIFFRACTION (2.55 Å) | | Cite: | Crystal structure of the lipid flippase MurJ in a "squeezed" form distinct from its inward- and outward-facing forms.
Structure, 30, 2022
|
|
7WAW
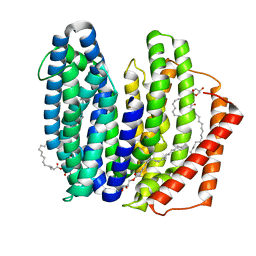 
 | | MurJ inward closed form | | Descriptor: | (2R)-2,3-dihydroxypropyl (9Z)-octadec-9-enoate, lipid II flippase MurJ | | Authors: | Tsukazaki, T, Kohga, H, Tanaka, Y, Yoshikaie, K, Taniguchi, K, Fujimoto, K. | | Deposit date: | 2021-12-15 | | Release date: | 2022-06-01 | | Last modified: | 2022-08-17 | | Method: | X-RAY DIFFRACTION (2.8 Å) | | Cite: | Crystal structure of the lipid flippase MurJ in a "squeezed" form distinct from its inward- and outward-facing forms.
Structure, 30, 2022
|
|
2Z92
 
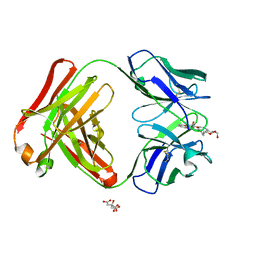 | | Crystal structure of the Fab fragment of anti-ciguatoxin antibody 10C9 in complex with CTX3C_ABCDE | | Descriptor: | (4Z)-2,8:7,12:11,15:14,18:17,22-PENTAANHYDRO-4,5,6,9,10,13,19,20,21-NONADEOXY-D-ARABINO-D-ALLO-D-ALLO-DOCOSA-4,9,20-TRIENITOL, Anti-ciguatoxin antibody 10C9 Fab heavy chain, Anti-ciguatoxin antibody 10C9 Fab light chain, ... | | Authors: | Ui, M, Tanaka, Y, Tsumoto, K. | | Deposit date: | 2007-09-14 | | Release date: | 2008-05-06 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | How Protein Recognizes Ladder-like Polycyclic Ethers: INTERACTIONS BETWEEN CIGUATOXIN (CTX3C) FRAGMENTS AND ITS SPECIFIC ANTIBODY 10C9
J.Biol.Chem., 283, 2008
|
|
2Z91
 
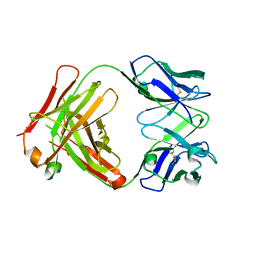 | | Crystal structure of the Fab fragment of anti-ciguatoxin antibody 10C9 | | Descriptor: | Anti-ciguatoxin antibody 10C9 FAB heavy chain, Anti-ciguatoxin antibody 10C9 FAB light chain | | Authors: | Ui, M, Tanaka, Y, Tsumoto, K. | | Deposit date: | 2007-09-14 | | Release date: | 2008-05-06 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (2.6 Å) | | Cite: | How Protein Recognizes Ladder-like Polycyclic Ethers: INTERACTIONS BETWEEN CIGUATOXIN (CTX3C) FRAGMENTS AND ITS SPECIFIC ANTIBODY 10C9
J.Biol.Chem., 283, 2008
|
|
2Z93
 
 | | Crystal structure of Fab fragment of anti-ciguatoxin antibody 10C9 in complex with CTX3C-ABCD | | Descriptor: | 1,6:5,9:8,12:11,16-TETRAANHYDRO-2,3,4,10,13,14-HEXADEOXY-D-GLYCERO-D-ALLO-D-GULO-HEPTADECA-2,13-DIENITOL, Anti-ciguatoxin antibody 10C9 Fab heavy chain, Anti-ciguatoxin antibody 10C9 Fab light chain | | Authors: | Ui, M, Tanaka, Y, Tsumoto, K. | | Deposit date: | 2007-09-14 | | Release date: | 2008-05-06 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | How Protein Recognizes Ladder-like Polycyclic Ethers: INTERACTIONS BETWEEN CIGUATOXIN (CTX3C) FRAGMENTS AND ITS SPECIFIC ANTIBODY 10C9
J.Biol.Chem., 283, 2008
|
|
