1V9P
 
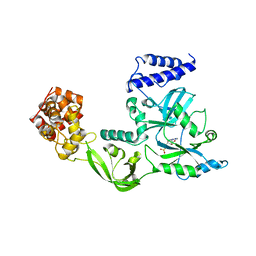 | | Crystal Structure Of Nad+-Dependent DNA Ligase | | Descriptor: | ADENOSINE MONOPHOSPHATE, DNA ligase, ZINC ION | | Authors: | Lee, J.Y, Chang, C, Song, H.K, Moon, J, Yang, J.K, Kim, H.K, Kwon, S.K, Suh, S.W. | | Deposit date: | 2004-01-27 | | Release date: | 2004-03-30 | | Last modified: | 2023-12-27 | | Method: | X-RAY DIFFRACTION (2.9 Å) | | Cite: | Crystal structure of NAD(+)-dependent DNA ligase: modular architecture and functional implications.
Embo J., 19, 2000
|
|
5JST
 | | MBP fused MDV1 coiled coil | | Descriptor: | ACETATE ION, GLYCEROL, Maltose-binding periplasmic protein,Mitochondrial division protein 1, ... | | Authors: | Kim, B.-W, Song, H.K. | | Deposit date: | 2016-05-09 | | Release date: | 2017-03-22 | | Last modified: | 2023-11-08 | | Method: | X-RAY DIFFRACTION (2.199 Å) | | Cite: | ACCORD: an assessment tool to determine the orientation of homodimeric coiled-coils.
Sci Rep, 7, 2017
|
|
6A9B
 
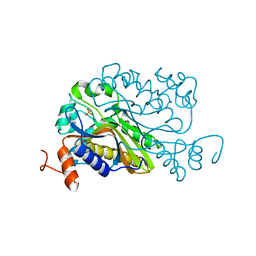 | | T4 dCMP hydroxymethylase structure solved by I-SAD using a home source | | Descriptor: | Deoxycytidylate 5-hydroxymethyltransferase, IODIDE ION, PHOSPHATE ION | | Authors: | Park, S.H, Song, H.K. | | Deposit date: | 2018-07-12 | | Release date: | 2019-01-02 | | Last modified: | 2024-03-27 | | Method: | X-RAY DIFFRACTION (2.01 Å) | | Cite: | A cytosine modification mechanism revealed by the structure of a ternary complex of deoxycytidylate hydroxymethylase from bacteriophage T4 with its cofactor and substrate.
Iucrj, 6, 2019
|
|
1OIL
 
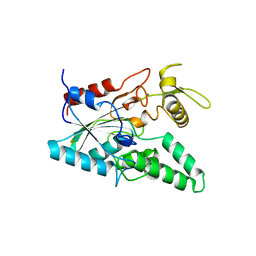 | | STRUCTURE OF LIPASE | | Descriptor: | CALCIUM ION, LIPASE | | Authors: | Kim, K.K, Song, H.K, Shin, D.H, Suh, S.W. | | Deposit date: | 1996-12-06 | | Release date: | 1997-05-15 | | Last modified: | 2018-04-18 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | The crystal structure of a triacylglycerol lipase from Pseudomonas cepacia reveals a highly open conformation in the absence of a bound inhibitor.
Structure, 5, 1997
|
|
6SJ0
 
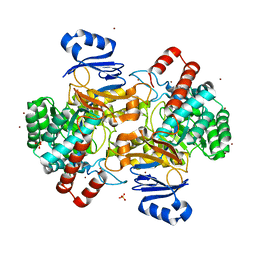 | | Amidohydrolase, AHS | | Descriptor: | Amidohydrolase, BICARBONATE ION, DI(HYDROXYETHYL)ETHER, ... | | Authors: | Naismith, J.H, Song, H. | | Deposit date: | 2019-08-12 | | Release date: | 2020-01-15 | | Last modified: | 2024-05-15 | | Method: | X-RAY DIFFRACTION (1.75 Å) | | Cite: | The Biosynthesis of the Benzoxazole in Nataxazole Proceeds via an Unstable Ester and has Synthetic Utility.
Angew.Chem.Int.Ed.Engl., 59, 2020
|
|
6ICO
 
 | | Pseudomonas putida CBB5 NdmA with theophylline | | Descriptor: | COBALT (II) ION, FE2/S2 (INORGANIC) CLUSTER, Methylxanthine N1-demethylase NdmA, ... | | Authors: | Kim, J.H, Kim, B.H, Kang, S.Y, Song, H.K. | | Deposit date: | 2018-09-06 | | Release date: | 2019-09-04 | | Last modified: | 2024-03-27 | | Method: | X-RAY DIFFRACTION (1.85 Å) | | Cite: | Structural and Mechanistic Insights into Caffeine Degradation by the Bacterial N-Demethylase Complex.
J.Mol.Biol., 431, 2019
|
|
6ICK
 
 | | Pseudomonas putida CBB5 NdmA | | Descriptor: | FE (III) ION, FE2/S2 (INORGANIC) CLUSTER, Methylxanthine N1-demethylase NdmA | | Authors: | Kim, J.H, Kim, B.H, Kang, S.Y, Song, H.K. | | Deposit date: | 2018-09-06 | | Release date: | 2019-09-04 | | Last modified: | 2024-03-27 | | Method: | X-RAY DIFFRACTION (1.952 Å) | | Cite: | Structural and Mechanistic Insights into Caffeine Degradation by the Bacterial N-Demethylase Complex.
J.Mol.Biol., 431, 2019
|
|
6ICM
 
 | | Pseudomonas putida CBB5 NdmA with ferredoxin domain of NdmD | | Descriptor: | FE (III) ION, FE2/S2 (INORGANIC) CLUSTER, Methylxanthine N1-demethylase NdmA, ... | | Authors: | Kim, J.H, Kim, B.H, Kang, S.Y, Song, H.K. | | Deposit date: | 2018-09-06 | | Release date: | 2019-09-04 | | Last modified: | 2024-03-27 | | Method: | X-RAY DIFFRACTION (2.961 Å) | | Cite: | Structural and Mechanistic Insights into Caffeine Degradation by the Bacterial N-Demethylase Complex.
J.Mol.Biol., 431, 2019
|
|
6ICP
 
 | | Pseudomonas putida CBB5 NdmA QL mutant with caffeine | | Descriptor: | CAFFEINE, FE (III) ION, FE2/S2 (INORGANIC) CLUSTER, ... | | Authors: | Kim, J.H, Kim, B.H, Kang, S.Y, Song, H.K. | | Deposit date: | 2018-09-06 | | Release date: | 2019-09-04 | | Last modified: | 2024-03-27 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Structural and Mechanistic Insights into Caffeine Degradation by the Bacterial N-Demethylase Complex.
J.Mol.Biol., 431, 2019
|
|
6ICN
 
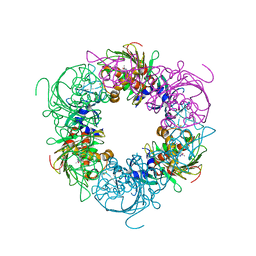 | | Pseudomonas putida CBB5 NdmA with caffeine | | Descriptor: | CAFFEINE, COBALT (II) ION, FE2/S2 (INORGANIC) CLUSTER, ... | | Authors: | Kim, J.H, Kim, B.H, Kang, S.Y, Song, H.K. | | Deposit date: | 2018-09-06 | | Release date: | 2019-09-04 | | Last modified: | 2024-03-27 | | Method: | X-RAY DIFFRACTION (1.65 Å) | | Cite: | Structural and Mechanistic Insights into Caffeine Degradation by the Bacterial N-Demethylase Complex.
J.Mol.Biol., 431, 2019
|
|
6ICQ
 
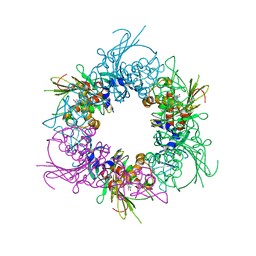 | | Pseudomonas putida CBB5 NdmA QL mutant with theobromine | | Descriptor: | FE (III) ION, FE2/S2 (INORGANIC) CLUSTER, Methylxanthine N1-demethylase NdmA, ... | | Authors: | Kim, J.H, Kim, B.H, Kang, S.Y, Song, H.K. | | Deposit date: | 2018-09-06 | | Release date: | 2019-09-04 | | Last modified: | 2024-03-27 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Structural and Mechanistic Insights into Caffeine Degradation by the Bacterial N-Demethylase Complex.
J.Mol.Biol., 431, 2019
|
|
6AED
 
 | |
6SJ2
 
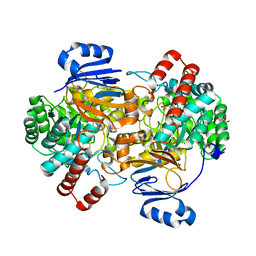 | | Amidohydrolase, AHS with 3-HAA | | Descriptor: | 3-HYDROXYANTHRANILIC ACID, Amidohydrolase, GLYCEROL, ... | | Authors: | Naismith, J.H, Song, H. | | Deposit date: | 2019-08-12 | | Release date: | 2020-01-15 | | Last modified: | 2024-05-15 | | Method: | X-RAY DIFFRACTION (1.25 Å) | | Cite: | The Biosynthesis of the Benzoxazole in Nataxazole Proceeds via an Unstable Ester and has Synthetic Utility.
Angew.Chem.Int.Ed.Engl., 59, 2020
|
|
6SIW
 
 | | PaaK family AMP-ligase with AMP | | Descriptor: | 1,2-ETHANEDIOL, 2-(2-(2-(2-(2-(2-ETHOXYETHOXY)ETHOXY)ETHOXY)ETHOXY)ETHOXY)ETHANOL, ADENOSINE MONOPHOSPHATE, ... | | Authors: | Naismith, J.H, Song, H. | | Deposit date: | 2019-08-12 | | Release date: | 2020-01-15 | | Last modified: | 2024-05-15 | | Method: | X-RAY DIFFRACTION (1.96 Å) | | Cite: | The Biosynthesis of the Benzoxazole in Nataxazole Proceeds via an Unstable Ester and has Synthetic Utility.
Angew.Chem.Int.Ed.Engl., 59, 2020
|
|
6SJ3
 
 | | Amidohydrolase, AHS with 3-HBA | | Descriptor: | 1,2-ETHANEDIOL, 3-HYDROXYBENZOIC ACID, Amidohydrolase, ... | | Authors: | Naismith, J.H, Song, H. | | Deposit date: | 2019-08-12 | | Release date: | 2020-01-15 | | Last modified: | 2024-01-24 | | Method: | X-RAY DIFFRACTION (1.17 Å) | | Cite: | The Biosynthesis of the Benzoxazole in Nataxazole Proceeds via an Unstable Ester and has Synthetic Utility.
Angew.Chem.Int.Ed.Engl., 59, 2020
|
|
6SIZ
 
 | | PaaK family AMP-ligase with ANP and substrate | | Descriptor: | 1,2-ETHANEDIOL, 3-HYDROXYANTHRANILIC ACID, AMP-dependent synthetase and ligase, ... | | Authors: | Naismith, J.H, Song, H. | | Deposit date: | 2019-08-12 | | Release date: | 2020-01-15 | | Last modified: | 2024-01-24 | | Method: | X-RAY DIFFRACTION (1.77 Å) | | Cite: | The Biosynthesis of the Benzoxazole in Nataxazole Proceeds via an Unstable Ester and has Synthetic Utility.
Angew.Chem.Int.Ed.Engl., 59, 2020
|
|
6SIY
 
 | | PaaK family AMP-ligase with AMP and substrate | | Descriptor: | 1,2-ETHANEDIOL, 3-HYDROXYANTHRANILIC ACID, ADENOSINE MONOPHOSPHATE, ... | | Authors: | Naismith, J.H, Song, H. | | Deposit date: | 2019-08-12 | | Release date: | 2020-01-15 | | Last modified: | 2024-01-24 | | Method: | X-RAY DIFFRACTION (1.95 Å) | | Cite: | The Biosynthesis of the Benzoxazole in Nataxazole Proceeds via an Unstable Ester and has Synthetic Utility.
Angew.Chem.Int.Ed.Engl., 59, 2020
|
|
6SIX
 
 | | PaaK family AMP-ligase with ANP | | Descriptor: | 1,2-ETHANEDIOL, AMP-dependent synthetase and ligase, CITRATE ANION, ... | | Authors: | Naismith, J.H, Song, H. | | Deposit date: | 2019-08-12 | | Release date: | 2020-01-15 | | Last modified: | 2024-05-15 | | Method: | X-RAY DIFFRACTION (1.88 Å) | | Cite: | The Biosynthesis of the Benzoxazole in Nataxazole Proceeds via an Unstable Ester and has Synthetic Utility.
Angew.Chem.Int.Ed.Engl., 59, 2020
|
|
6TM4
 
 | | NatL2 in complex with two molecules of salicylic acid | | Descriptor: | 2-HYDROXYBENZOIC ACID, ADENOSINE MONOPHOSPHATE, MAGNESIUM ION, ... | | Authors: | Naismith, J.H, Song, H. | | Deposit date: | 2019-12-03 | | Release date: | 2020-01-15 | | Last modified: | 2024-01-24 | | Method: | X-RAY DIFFRACTION (1.89 Å) | | Cite: | The Biosynthesis of the Benzoxazole in Nataxazole Proceeds via an Unstable Ester and has Synthetic Utility.
Angew.Chem.Int.Ed.Engl., 59, 2020
|
|
1QZY
 
 | | Human Methionine Aminopeptidase in complex with bengamide inhibitor LAF153 and cobalt | | Descriptor: | (E)-(2R,3R,4S,5R)-3,4,5-TRIHYDROXY-2-METHOXY-8,8-DIMETHYL-NON-6-ENOIC ACID ((3S,6R)-6-HYDROXY-2-OXO-AZEPAN-3-YL)-AMIDE, COBALT (II) ION, Methionine aminopeptidase 2, ... | | Authors: | Eck, M.J, Song, H.K, Morollo, A. | | Deposit date: | 2003-09-18 | | Release date: | 2003-11-25 | | Last modified: | 2011-07-13 | | Method: | X-RAY DIFFRACTION (1.6 Å) | | Cite: | Proteomics-based target identification: bengamides as a new class of methionine aminopeptidase inhibitors.
J.Biol.Chem., 278, 2003
|
|
3KTJ
 
 | |
3KTI
 | | Structure of ClpP in complex with ADEP1 | | Descriptor: | 2-[N-CYCLOHEXYLAMINO]ETHANE SULFONIC ACID, ATP-dependent Clp protease proteolytic subunit, Acyldepsipeptide 1, ... | | Authors: | Lee, B.-G, Brotz-Oesterhelt, H, Song, H.K. | | Deposit date: | 2009-11-25 | | Release date: | 2010-03-23 | | Last modified: | 2013-02-27 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Structures of ClpP in complex with acyldepsipeptide antibiotics reveal its activation mechanism
Nat.Struct.Mol.Biol., 17, 2010
|
|
3KTK
 | |
3KTG
 | |
3KTH
 
 | |
