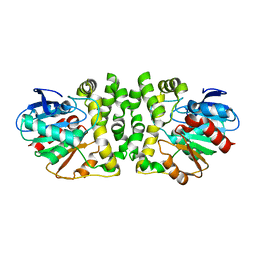
8HGU
 
 | |
1MFY
 
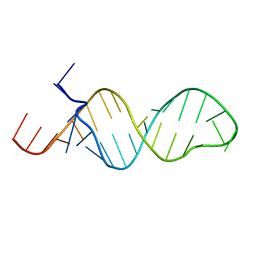 | | SOLUTION STRUCTURE OF INFLUENZA A VIRUS C4 PROMOTER | | Descriptor: | C4 promoter of influneza A virus | | Authors: | Lee, M.-K, Bae, S.-H, Park, C.-J, Cheong, H.-K, Cheong, C, Choi, B.-S. | | Deposit date: | 2002-08-14 | | Release date: | 2002-09-18 | | Last modified: | 2024-05-22 | | Method: | SOLUTION NMR | | Cite: | A single-nucleotide natural variation (U4 to C4) in an influenza A virus promoter exhibits a large structural change: implications for differential viral RNA synthesis by RNA-dependent RNA polymerase.
Nucleic Acids Res., 31, 2003
|
|
6WMZ
 
 | | Crystal structure of human SFPQ/NONO complex | | Descriptor: | Non-POU domain-containing octamer-binding protein, SULFATE ION, Splicing factor, ... | | Authors: | Lee, M, Bond, C.S. | | Deposit date: | 2020-04-22 | | Release date: | 2021-04-28 | | Last modified: | 2023-10-18 | | Method: | X-RAY DIFFRACTION (2.85 Å) | | Cite: | Structure analysis of human SFPQ/NONO complex
To Be Published
|
|
3TL1
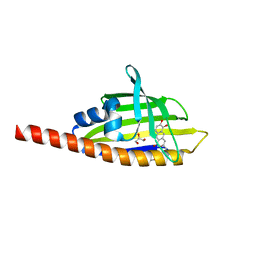 
 | | Crystal structure of the Streptomyces coelicolor WhiE ORFVI polyketide aromatase/cyclase | | Descriptor: | 6,7,9-trihydroxy-3-methyl-1H-benzo[g]isochromen-1-one, GLYCEROL, Polyketide cyclase | | Authors: | Lee, M.-Y, Ames, B.D, Zhang, W, Tang, Y, Tsai, S.-C. | | Deposit date: | 2011-08-29 | | Release date: | 2012-04-04 | | Last modified: | 2024-02-28 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Insight into the Molecular Basis of Aromatic Polyketide Cyclization: Crystal Structure and in Vitro Characterization of WhiE-ORFVI.
Biochemistry, 51, 2012
|
|
4WII
 
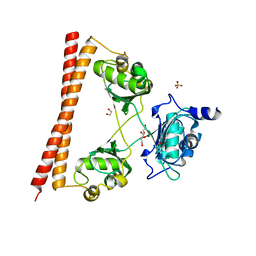 | | HUMAN SPLICING FACTOR, CONSTRUCT 3 | | Descriptor: | 1,2-ETHANEDIOL, SULFATE ION, Splicing factor, ... | | Authors: | lee, M, bond, c.s. | | Deposit date: | 2014-09-26 | | Release date: | 2015-04-01 | | Last modified: | 2023-12-27 | | Method: | X-RAY DIFFRACTION (2.05 Å) | | Cite: | The structure of human SFPQ reveals a coiled-coil mediated polymer essential for functional aggregation in gene regulation.
Nucleic Acids Res., 43, 2015
|
|
4WIK
 
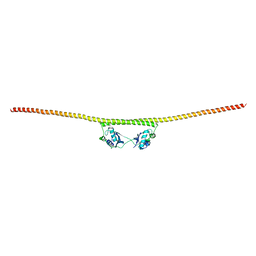 | | HUMAN SPLICING FACTOR, CONSTRUCT 2 | | Descriptor: | Splicing factor, proline- and glutamine-rich | | Authors: | lee, M, bond, c.s. | | Deposit date: | 2014-09-26 | | Release date: | 2015-04-01 | | Last modified: | 2023-12-27 | | Method: | X-RAY DIFFRACTION (3 Å) | | Cite: | The structure of human SFPQ reveals a coiled-coil mediated polymer essential for functional aggregation in gene regulation.
Nucleic Acids Res., 43, 2015
|
|
4WIJ
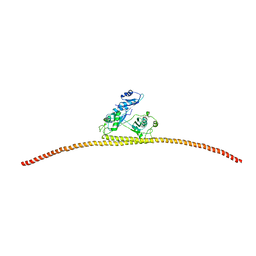 
 | | HUMAN SPLICING FACTOR, CONSTRUCT 1 | | Descriptor: | Splicing factor, proline- and glutamine-rich | | Authors: | lee, M, bond, c.s. | | Deposit date: | 2014-09-26 | | Release date: | 2015-04-01 | | Last modified: | 2023-12-27 | | Method: | X-RAY DIFFRACTION (3.49 Å) | | Cite: | The structure of human SFPQ reveals a coiled-coil mediated polymer essential for functional aggregation in gene regulation.
Nucleic Acids Res., 43, 2015
|
|
1MKD
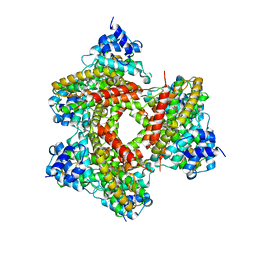 
 | | crystal structure of PDE4D catalytic domain and zardaverine complex | | Descriptor: | 6-(4-DIFLUOROMETHOXY-3-METHOXY-PHENYL)-2H-PYRIDAZIN-3-ONE, MAGNESIUM ION, Phosphodiesterase 4D, ... | | Authors: | Lee, M.E, Markowitz, J, Lee, J.-O, Lee, H. | | Deposit date: | 2002-08-29 | | Release date: | 2003-03-01 | | Last modified: | 2024-03-13 | | Method: | X-RAY DIFFRACTION (2.9 Å) | | Cite: | Crystal structure of phosphodiesterase 4D and inhibitor complex
FEBS LETT., 530, 2002
|
|
7SP0
 
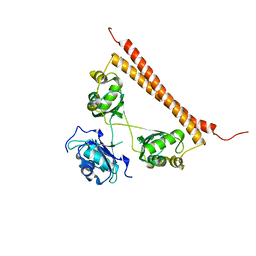 | | Crystal structure of human SFPQ L534I mutant in complex with zinc | | Descriptor: | Splicing factor, proline- and glutamine-rich, ZINC ION | | Authors: | Lee, M. | | Deposit date: | 2021-11-01 | | Release date: | 2022-10-12 | | Last modified: | 2023-10-18 | | Method: | X-RAY DIFFRACTION (1.83 Å) | | Cite: | Familial ALS-associated SFPQ variants promote the formation of SFPQ cytoplasmic aggregates in primary neurons.
Open Biology, 12, 2022
|
|
6VVI
 
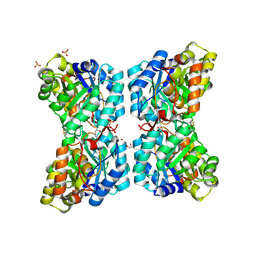 | |
6VVH
 
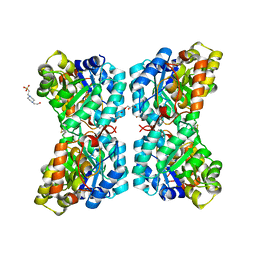 | |
6K2K
 
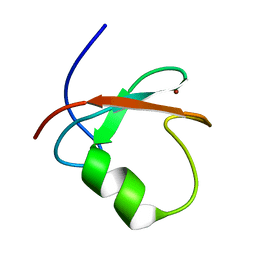 | | Solution structure of MUL1-RING domain | | Descriptor: | Mitochondrial ubiquitin ligase activator of NFKB 1, ZINC ION | | Authors: | Lee, M.S, Lee, M.K, Ryu, K.S, Chi, S.W. | | Deposit date: | 2019-05-14 | | Release date: | 2019-07-10 | | Last modified: | 2024-05-15 | | Method: | SOLUTION NMR | | Cite: | Solution structure of MUL1-RING domain and its interaction with p53 transactivation domain.
Biochem.Biophys.Res.Commun., 516, 2019
|
|
5K3X
 
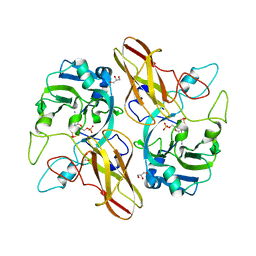 | | Crystal Structure of the sulfite dehydrogenase, SorT R78K mutant from Sinorhizobium meliloti | | Descriptor: | (MOLYBDOPTERIN-S,S)-OXO-MOLYBDENUM, GLYCEROL, Putative sulfite oxidase | | Authors: | Lee, M, McGrath, A, Maher, M. | | Deposit date: | 2016-05-20 | | Release date: | 2017-05-24 | | Last modified: | 2023-09-27 | | Method: | X-RAY DIFFRACTION (1.6 Å) | | Cite: | The central active site arginine in sulfite oxidizing enzymes alters kinetic properties by controlling electron transfer and redox interactions.
Biochim. Biophys. Acta, 1859, 2017
|
|
6WQA
 
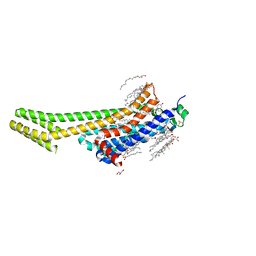 | | 2.0A angstrom A2a adenosine receptor structure using XFEL data collected in helium atmosphere. | | Descriptor: | (2R)-2,3-dihydroxypropyl (9Z)-octadec-9-enoate, (2S)-2,3-dihydroxypropyl (9Z)-octadec-9-enoate, 4-{2-[(7-amino-2-furan-2-yl[1,2,4]triazolo[1,5-a][1,3,5]triazin-5-yl)amino]ethyl}phenol, ... | | Authors: | Lee, M.-Y, Geiger, J, Ishchenko, A, Han, G.W, Barty, A, White, T.A, Gati, C, Batyuk, A, Hunter, M.S, Aquila, A, Boutet, S, Weierstall, U, Cherezov, V, Liu, W. | | Deposit date: | 2020-04-28 | | Release date: | 2020-11-18 | | Last modified: | 2023-10-18 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Harnessing the power of an X-ray laser for serial crystallography of membrane proteins crystallized in lipidic cubic phase
Iucrj, 7, 2020
|
|
1QH2
 
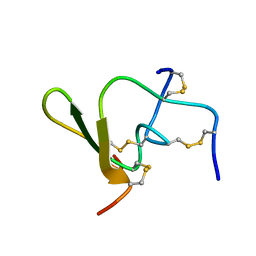 | |
1ZNF
 
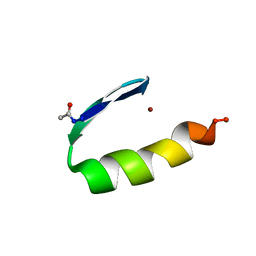 | | THREE-DIMENSIONAL SOLUTION STRUCTURE OF A SINGLE ZINC FINGER DNA-BINDING DOMAIN | | Descriptor: | 31ST ZINC FINGER FROM XFIN, ZINC ION | | Authors: | Lee, M.S, Gippert, G.P, Soman, K.V, Case, D.A, Wright, P.E. | | Deposit date: | 1989-09-25 | | Release date: | 1991-07-15 | | Last modified: | 2017-11-29 | | Method: | SOLUTION NMR | | Cite: | Three-dimensional solution structure of a single zinc finger DNA-binding domain.
Science, 245, 1989
|
|
6OWJ
 
 | | Zn-mediated polymerization of human SFPQ | | Descriptor: | Splicing factor, proline- and glutamine-rich, ZINC ION | | Authors: | Lee, M. | | Deposit date: | 2019-05-10 | | Release date: | 2020-02-05 | | Last modified: | 2023-10-11 | | Method: | X-RAY DIFFRACTION (1.94 Å) | | Cite: | Structural basis of the zinc-induced cytoplasmic aggregation of the RNA-binding protein SFPQ.
Nucleic Acids Res., 48, 2020
|
|
7F3V
 
 | | Crystal structure of YfiH with C107A mutation in complex with endogenous UDP-MurNAc | | Descriptor: | (2R)-2-{[(2R,3R,4R,5S,6R)-3-(acetylamino)-2-{[(S)-{[(R)-{[(2R,3S,4R,5R)-5-(2,4-dioxo-3,4-dihydropyrimidin-1(2H)-yl)-3,4-dihydroxytetrahydrofuran-2-yl]methoxy}(hydroxy)phosphoryl]oxy}(hydroxy)phosphoryl]oxy}-5-hydroxy-6-(hydroxymethyl)tetrahydro-2H-pyran-4-yl]oxy}propanoic acid, 2-AMINO-2-HYDROXYMETHYL-PROPANE-1,3-DIOL, PHOSPHATE ION, ... | | Authors: | Lee, M.S, Hsieh, K.Y, Chang, C.I. | | Deposit date: | 2021-06-17 | | Release date: | 2021-12-29 | | Last modified: | 2023-11-29 | | Method: | X-RAY DIFFRACTION (1.47 Å) | | Cite: | Structural Basis for the Peptidoglycan-Editing Activity of YfiH.
Mbio, 13, 2021
|
|
3HQ2
 
 | | BsuCP Crystal Structure | | Descriptor: | Bacillus subtilis M32 carboxypeptidase, CHLORIDE ION, FLUORIDE ION, ... | | Authors: | Lee, M.M, Isaza, C.E, White, J.D, Chen, R.P.-Y, Liang, G.F.-C, He, H.T.-F, Chan, S.I, Chan, M.K. | | Deposit date: | 2009-06-05 | | Release date: | 2009-06-30 | | Last modified: | 2024-02-21 | | Method: | X-RAY DIFFRACTION (2.9 Å) | | Cite: | Insight into the substrate length restriction of M32 carboxypeptidases: Characterization of two distinct subfamilies.
Proteins, 77, 2009
|
|
3HOA
 
 | |
1OV8
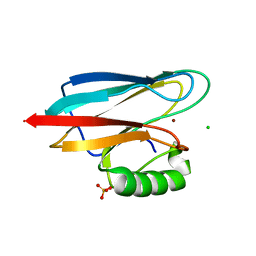 
 | | Auracyanin B structure in space group, P65 | | Descriptor: | Auracyanin B, CHLORIDE ION, COPPER (II) ION, ... | | Authors: | Lee, M, Maher, M.J, Freeman, H.C, Guss, J.M. | | Deposit date: | 2003-03-25 | | Release date: | 2003-09-09 | | Last modified: | 2023-08-16 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Auracyanin B structure in space group P6(5).
Acta Crystallogr.,Sect.D, 59, 2003
|
|
1XGE
 
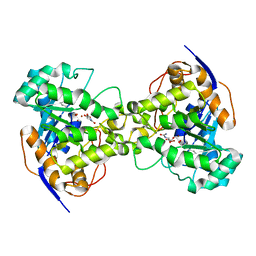 | | Dihydroorotase from Escherichia coli: Loop Movement and Cooperativity between subunits | | Descriptor: | (4S)-2,6-DIOXOHEXAHYDROPYRIMIDINE-4-CARBOXYLIC ACID, Dihydroorotase, N-CARBAMOYL-L-ASPARTATE, ... | | Authors: | Lee, M, Chan, C.W, Guss, J.M, Christopherson, R.I, Maher, M.J. | | Deposit date: | 2004-09-17 | | Release date: | 2005-04-26 | | Last modified: | 2023-11-15 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Dihydroorotase from Escherichia coli: Loop Movement and Cooperativity between Subunits
J.Mol.Biol., 348, 2005
|
|
2E25
 
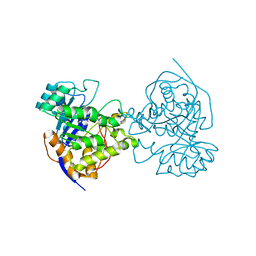 | | The Crystal Structure of the T109S mutant of E. coli Dihydroorotase complexed with an inhibitor 5-fluoroorotate | | Descriptor: | 5-FLUORO-2,6-DIOXO-1,2,3,6-TETRAHYDROPYRIMIDINE-4-CARBOXYLIC ACID, Dihydroorotase, ZINC ION | | Authors: | Lee, M, Maher, M.J, Guss, J.M. | | Deposit date: | 2006-11-08 | | Release date: | 2007-03-13 | | Last modified: | 2023-11-15 | | Method: | X-RAY DIFFRACTION (2.7 Å) | | Cite: | Structure of the T109S mutant of Escherichia coli dihydroorotase complexed with the inhibitor 5-fluoroorotate: catalytic activity is reflected by the crystal form
Acta Crystallogr.,Sect.F, 63, 2007
|
|
3DSQ
 
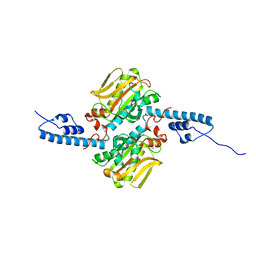 | |
4MF9
 
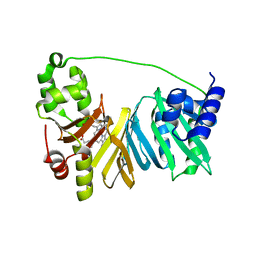 | |
