1IAV
 
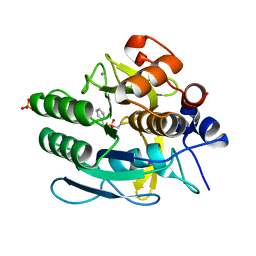 | |
1A8D
 
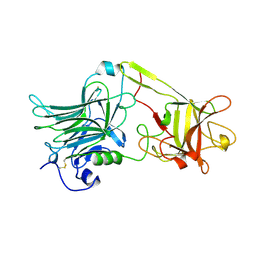 | | TETANUS TOXIN C FRAGMENT | | Descriptor: | GOLD ION, TETANUS NEUROTOXIN | | Authors: | Knapp, M, Rupp, B. | | Deposit date: | 1998-03-23 | | Release date: | 1998-10-14 | | Last modified: | 2011-07-13 | | Method: | X-RAY DIFFRACTION (1.57 Å) | | Cite: | The 1.61 Angstrom Structure of the Tetanus Toxin Ganglioside Binding Region: Solved by MAD and Mir Phase Combination
Am.Cryst.Assoc.,Abstr.Papers (Annual Meeting), 25, 1998
|
|
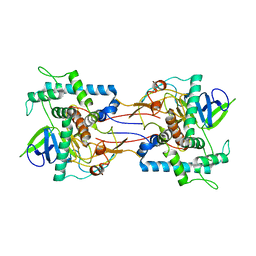
1E1H
 
 | |
1Q5P
 
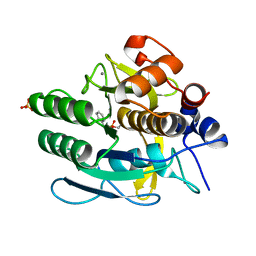 | | S156E/S166D variant of Bacillus lentus subtilisin | | Descriptor: | CALCIUM ION, SULFATE ION, Serine protease | | Authors: | Bott, R.R, Chan, G, Domingo, B, Ganshaw, G, Hsia, C.Y, Knapp, M, Murray, C.J. | | Deposit date: | 2003-08-08 | | Release date: | 2003-11-11 | | Last modified: | 2018-04-04 | | Method: | X-RAY DIFFRACTION (1.6 Å) | | Cite: | Do enzymes change the nature of transition states? Mapping the transition state for general acid-base catalysis of a serine protease
Biochemistry, 42, 2003
|
|
6UD7
 
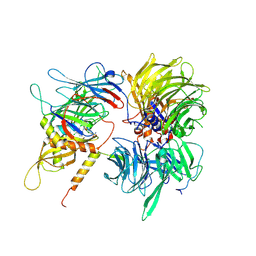 | | Crystal structure of full-length human DCAF15-DDB1(deltaBPB)-DDA1-RBM39 in complex with indisulam | | Descriptor: | DDB1- and CUL4-associated factor 15, DET1- and DDB1-associated protein 1, DNA damage-binding protein 1,DNA damage-binding protein 1, ... | | Authors: | Bussiere, D.E, Shu, W, Xie, L, Knapp, M. | | Deposit date: | 2019-09-18 | | Release date: | 2019-12-18 | | Last modified: | 2020-01-22 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Structural basis of indisulam-mediated RBM39 recruitment to DCAF15 E3 ligase complex.
Nat.Chem.Biol., 16, 2020
|
|
3FLN
 
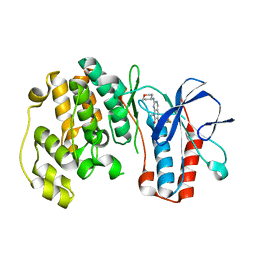 | | P38 kinase crystal structure in complex with R1487 | | Descriptor: | 6-(2,4-difluorophenoxy)-8-methyl-2-(tetrahydro-2H-pyran-4-ylamino)pyrido[2,3-d]pyrimidin-7(8H)-one, Mitogen-activated protein kinase 14 | | Authors: | Kuglstatter, A, Knapp, M. | | Deposit date: | 2008-12-19 | | Release date: | 2009-12-22 | | Last modified: | 2024-02-21 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | The Discovery of Pamapimod, R1503 and R1487 as Orally Bioavailable and Highly Selective Inhibitors of p38 Map Kinase
To be Published
|
|
3FLQ
 
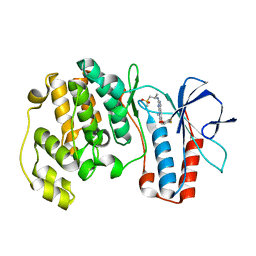 | |
3FI4
 
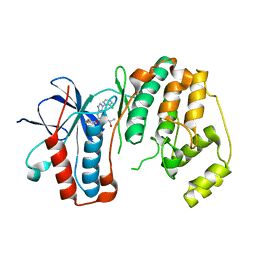 | | P38 kinase crystal structure in complex with RO4499 | | Descriptor: | (2S)-1-{[3-(2-chlorophenyl)-6-(2,4-difluorophenoxy)-1H-pyrazolo[3,4-d]pyrimidin-4-yl]amino}propan-2-ol, Mitogen-activated protein kinase 14 | | Authors: | Kuglstatter, A, Knapp, M, Dunten, P. | | Deposit date: | 2008-12-10 | | Release date: | 2009-12-22 | | Last modified: | 2023-09-06 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Mapping Binding Pocket Volume: Potential Applications towards Ligand Design and Selectivity
To be Published
|
|
3FKO
 
 | | P38 kinase crystal structure in complex with RO3668 | | Descriptor: | 3-(2-chlorophenyl)-6-(2-fluorophenoxy)-2H-indazole, Mitogen-activated protein kinase 14 | | Authors: | Kuglstatter, A, Knapp, M. | | Deposit date: | 2008-12-17 | | Release date: | 2009-12-22 | | Last modified: | 2023-09-06 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Mapping Binding Pocket Volume: Potential Applications towards Ligand Design and Selectivity
To be Published
|
|
3FL4
 
 | | P38 kinase crystal structure in complex with RO5634 | | Descriptor: | 6-(2,4-difluorophenoxy)-3-(2-methylphenyl)-1H-pyrazolo[3,4-d]pyrimidine, Mitogen-activated protein kinase 14 | | Authors: | Kuglstatter, A, Knapp, M. | | Deposit date: | 2008-12-18 | | Release date: | 2009-12-22 | | Last modified: | 2024-02-21 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Mapping Binding Pocket Volume: Potential Applications towards Ligand Design and Selectivity
To be Published
|
|
