3UJP
 
 | | Structure of MntC protein at 2.7A | | Descriptor: | CACODYLATE ION, MANGANESE (II) ION, Mn transporter subunit, ... | | Authors: | Kanteev, M, Adir, N. | | Deposit date: | 2011-11-08 | | Release date: | 2012-11-14 | | Last modified: | 2013-06-19 | | Method: | X-RAY DIFFRACTION (2.7 Å) | | Cite: | Arginine 116 stabilizes the entrance to the metal ion-binding site of the MntC protein.
Acta Crystallogr.,Sect.F, 69, 2013
|
|
4J6V
 
 | | Crystal Structure of Tyrosinase from Bacillus megaterium N205D mutant | | Descriptor: | COPPER (II) ION, Tyrosinase | | Authors: | Kanteev, M, Goldfeder, M, Adir, N, Fishman, A. | | Deposit date: | 2013-02-12 | | Release date: | 2013-12-25 | | Last modified: | 2024-02-28 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | The mechanism of copper uptake by tyrosinase from Bacillus megaterium.
J.Biol.Inorg.Chem., 18, 2013
|
|
4J6T
 
 | | Crystal Structure of Tyrosinase from Bacillus megaterium F197A mutant | | Descriptor: | COPPER (II) ION, Tyrosinase | | Authors: | Kanteev, M, Goldfeder, M, Adir, N, Fishman, A. | | Deposit date: | 2013-02-12 | | Release date: | 2013-12-25 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (2.43 Å) | | Cite: | The mechanism of copper uptake by tyrosinase from Bacillus megaterium.
J.Biol.Inorg.Chem., 18, 2013
|
|
4J6U
 
 | |
5I38
 
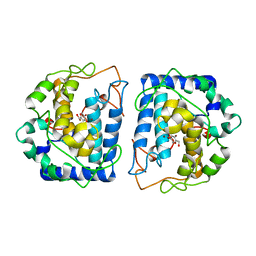 | | Crystal Structure of tyrosinase from Bacillus megaterium with inhibitor kojic acid in the active site | | Descriptor: | 5-HYDROXY-2-(HYDROXYMETHYL)-4H-PYRAN-4-ONE, COPPER (II) ION, Tyrosinase | | Authors: | Kanteev, M, Goldfeder, M, Deri, B, Adir, N, Fishman, A. | | Deposit date: | 2016-02-10 | | Release date: | 2016-10-12 | | Last modified: | 2024-01-10 | | Method: | X-RAY DIFFRACTION (2.6 Å) | | Cite: | The unravelling of the complex pattern of tyrosinase inhibition.
Sci Rep, 6, 2016
|
|
4X71
 
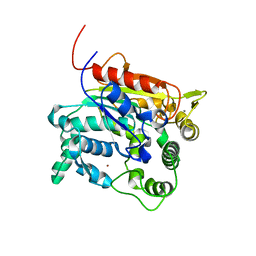 | | Crystal Structure of lipase from Geobacillus stearothermophilus T6 methanol stable variant A269T | | Descriptor: | CALCIUM ION, Lipase, ZINC ION | | Authors: | Kanteev, M, Dror, A, Gihaz, S, Shahar, A, Fishman, A. | | Deposit date: | 2014-12-09 | | Release date: | 2015-06-10 | | Last modified: | 2024-05-08 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Structural insights into methanol-stable variants of lipase T6 from Geobacillus stearothermophilus.
Appl.Microbiol.Biotechnol., 99, 2015
|
|
4X6U
 
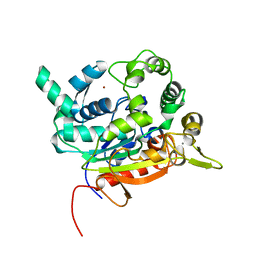 | | Crystal Structure of lipase from Geobacillus stearothermophilus T6 | | Descriptor: | CALCIUM ION, Lipase, ZINC ION | | Authors: | Kanteev, M, Dror, A, Gihaz, S, Fishman, A. | | Deposit date: | 2014-12-09 | | Release date: | 2015-06-10 | | Last modified: | 2024-05-08 | | Method: | X-RAY DIFFRACTION (2.201 Å) | | Cite: | Structural insights into methanol-stable variants of lipase T6 from Geobacillus stearothermophilus.
Appl.Microbiol.Biotechnol., 99, 2015
|
|
4X7B
 
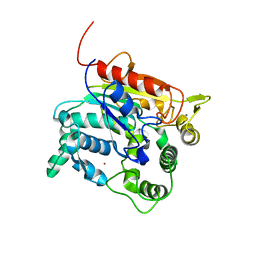 | | Crystal Structure of lipase from Geobacillus stearothermophilus T6 methanol stable variant H86Y/A269T | | Descriptor: | CALCIUM ION, Lipase, ZINC ION | | Authors: | Kanteev, M, Dror, A, Gihaz, S, Fishman, A. | | Deposit date: | 2014-12-09 | | Release date: | 2015-06-10 | | Last modified: | 2024-05-08 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | Structural insights into methanol-stable variants of lipase T6 from Geobacillus stearothermophilus.
Appl.Microbiol.Biotechnol., 99, 2015
|
|
4X85
 
 | | Crystal Structure of lipase from Geobacillus stearothermophilus T6 methanol stable variant H86Y/A269T/R374W | | Descriptor: | CALCIUM ION, Lipase, ZINC ION | | Authors: | Kanteev, M, Dror, A, Gihaz, S, Fishman, A. | | Deposit date: | 2014-12-10 | | Release date: | 2015-06-10 | | Last modified: | 2024-05-08 | | Method: | X-RAY DIFFRACTION (2.192 Å) | | Cite: | Structural insights into methanol-stable variants of lipase T6 from Geobacillus stearothermophilus.
Appl.Microbiol.Biotechnol., 99, 2015
|
|
5I3A
 
 | | Crystal Structure of tyrosinase from Bacillus megaterium with configuration A of hydroquinone inhibitor in the active site | | Descriptor: | Tyrosinase, ZINC ION, benzene-1,4-diol | | Authors: | Kanteev, M, Deri, B, Adir, N, Fishman, A. | | Deposit date: | 2016-02-10 | | Release date: | 2016-10-12 | | Last modified: | 2024-01-10 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | The unravelling of the complex pattern of tyrosinase inhibition.
Sci Rep, 6, 2016
|
|
5I3B
 
 | | Crystal Structure of tyrosinase from Bacillus megaterium with configuration B of hydroquinone inhibitor in the active site | | Descriptor: | Tyrosinase, ZINC ION, benzene-1,4-diol | | Authors: | Kanteev, M, Deri, B, Adir, N, Fishman, A. | | Deposit date: | 2016-02-10 | | Release date: | 2016-10-12 | | Last modified: | 2024-01-10 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | The unravelling of the complex pattern of tyrosinase inhibition.
Sci Rep, 6, 2016
|
|
4Z2U
 
 | |
4Z2R
 
 | |
4Z2T
 
 | |
5BRT
 
 | | Crystal Structure of 2-hydroxybiphenyl 3-monooxygenase from Pseudomonas azelaica with 2-hydroxybiphenyl in the active site | | Descriptor: | 2-HYDROXYBIPHENYL, 2-hydroxybiphenyl-3-monooxygenase, FLAVIN-ADENINE DINUCLEOTIDE | | Authors: | Kanteev, M, Bregman-Cohen, A, Deri, B, Adir, N, Fishman, A. | | Deposit date: | 2015-06-01 | | Release date: | 2015-08-19 | | Last modified: | 2024-01-10 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | A crystal structure of 2-hydroxybiphenyl 3-monooxygenase with bound substrate provides insights into the enzymatic mechanism.
Biochim.Biophys.Acta, 1854, 2015
|
|
4IRM
 
 | |
4HD7
 
 | | Crystal Structure of Tyrosinase from Bacillus megaterium V218G mutant soaked in CuSO4 | | Descriptor: | COPPER (II) ION, Tyrosinase | | Authors: | Goldfeder, M, Kanteev, M, Adir, N, Fishman, A. | | Deposit date: | 2012-10-02 | | Release date: | 2013-01-23 | | Last modified: | 2024-02-28 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Influencing the monophenolase/diphenolase activity ratio in tyrosinase.
Biochim.Biophys.Acta, 1834, 2013
|
|
4HD6
 
 | | Crystal Structure of Tyrosinase from Bacillus megaterium V218F mutant soaked in CuSO4 | | Descriptor: | COPPER (II) ION, Tyrosinase | | Authors: | Goldfeder, M, Kanteev, M, Adir, N, Fishman, A. | | Deposit date: | 2012-10-02 | | Release date: | 2013-01-23 | | Last modified: | 2024-02-28 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Influencing the monophenolase/diphenolase activity ratio in tyrosinase.
Biochim.Biophys.Acta, 1834, 2013
|
|
4HD4
 
 | | Crystal Structure of Tyrosinase from Bacillus megaterium V218F mutant | | Descriptor: | COPPER (II) ION, Tyrosinase | | Authors: | Goldfeder, M, Kanteev, M, Adir, N, Fishman, A. | | Deposit date: | 2012-10-02 | | Release date: | 2013-01-23 | | Last modified: | 2024-02-28 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Influencing the monophenolase/diphenolase activity ratio in tyrosinase.
Biochim.Biophys.Acta, 1834, 2013
|
|
4P6S
 
 | | Crystal Structure of tyrosinase from Bacillus megaterium with L-DOPA in the active site | | Descriptor: | 3,4-DIHYDROXYPHENYLALANINE, Tyrosinase, ZINC ION | | Authors: | Goldfeder, M, Kanteev, M, Adir, N, Fishman, A. | | Deposit date: | 2014-03-25 | | Release date: | 2014-07-30 | | Last modified: | 2023-12-27 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Determination of tyrosinase substrate-binding modes reveals mechanistic differences between type-3 copper proteins.
Nat Commun, 5, 2014
|
|
3NQ1
 
 | | Crystal Structure of Tyrosinase from Bacillus megaterium in complex with inhibitor kojic acid | | Descriptor: | 5-HYDROXY-2-(HYDROXYMETHYL)-4H-PYRAN-4-ONE, COPPER (II) ION, Tyrosinase, ... | | Authors: | Sendovski, M, Kanteev, M, Adir, N, Fishman, A. | | Deposit date: | 2010-06-29 | | Release date: | 2010-11-17 | | Last modified: | 2023-12-27 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | First structures of an active bacterial tyrosinase reveal copper plasticity.
J.Mol.Biol., 405, 2011
|
|
3NQ5
 
 | | Crystal Structure of Tyrosinase from Bacillus megaterium R209H mutant | | Descriptor: | COPPER (II) ION, Tyrosinase, ZINC ION | | Authors: | Sendovski, M, Kanteev, M, Adir, N, Fishman, A. | | Deposit date: | 2010-06-29 | | Release date: | 2010-11-17 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | First structures of an active bacterial tyrosinase reveal copper plasticity.
J.Mol.Biol., 405, 2011
|
|
3NPY
 
 | | Crystal Structure of Tyrosinase from Bacillus megaterium soaked in CuSO4 | | Descriptor: | CHLORIDE ION, COPPER (II) ION, Tyrosinase, ... | | Authors: | Sendovski, M, Kanteev, M, Adir, N, Fishman, A. | | Deposit date: | 2010-06-29 | | Release date: | 2010-11-17 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (2.192 Å) | | Cite: | First structures of an active bacterial tyrosinase reveal copper plasticity.
J.Mol.Biol., 405, 2011
|
|
3NM8
 
 | | Crystal structure of Tyrosinase from Bacillus megaterium | | Descriptor: | CHLORIDE ION, COPPER (II) ION, Tyrosinase, ... | | Authors: | Sendovski, M, Kanteev, M, Adir, N, Fishman, A. | | Deposit date: | 2010-06-22 | | Release date: | 2010-11-17 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | First structures of an active bacterial tyrosinase reveal copper plasticity
J.Mol.Biol., 405, 2011
|
|
4P6T
 
 | | Crystal Structure of tyrosinase from Bacillus megaterium with p-tyrosol in the active site | | Descriptor: | 4-(2-hydroxyethyl)phenol, COPPER (II) ION, Tyrosinase | | Authors: | Goldfeder, M, Kanteev, M, Adir, N, Fishman, A. | | Deposit date: | 2014-03-25 | | Release date: | 2014-07-30 | | Last modified: | 2023-12-27 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | Determination of tyrosinase substrate-binding modes reveals mechanistic differences between type-3 copper proteins.
Nat Commun, 5, 2014
|
|
