2F36
 
 | |
2F34
 
 | |
2F35
 
 | |
1IIT
 
 | | GLUR0 LIGAND BINDING CORE COMPLEX WITH L-SERINE | | Descriptor: | SERINE, Slr1257 protein | | Authors: | Mayer, M.L, Olson, R, Gouaux, E. | | Deposit date: | 2001-04-24 | | Release date: | 2001-09-19 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Mechanisms for ligand binding to GluR0 ion channels: crystal structures of the glutamate and serine complexes and a closed apo state.
J.Mol.Biol., 311, 2001
|
|
1TXF
 
 | |
1TT1
 
 | |
1II5
 
 | |
1S7Y
 
 | |
1S50
 
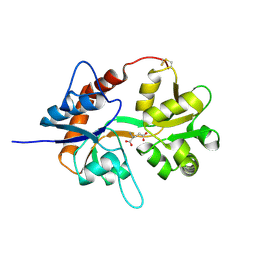 | |
1SD3
 
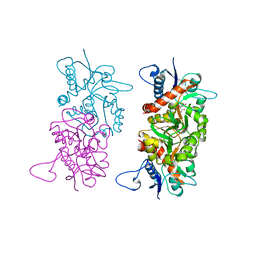 | |
1S9T
 
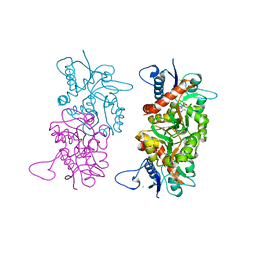 | |
1IIW
 
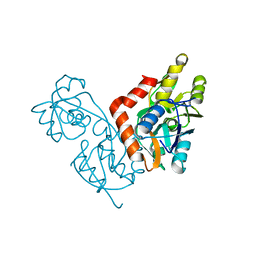 | |
3C32
 
 | |
3C35
 
 | |
3C31
 
 | |
3C33
 
 | |
3C36
 
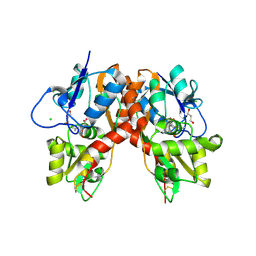 | |
3C34
 
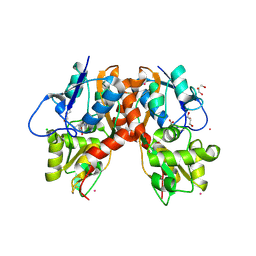 | |
2I0C
 
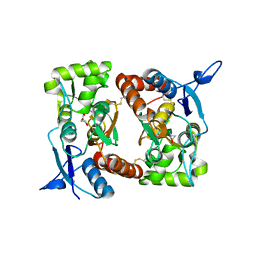 | |
2I0B
 
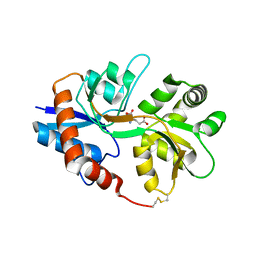 | |
2GFE
 
 | |
5CMB
 
 | |
5CMC
 
 | |
2OJT
 
 | | Structure and mechanism of kainate receptor modulation by anions | | Descriptor: | (S)-1-(2-AMINO-2-CARBOXYETHYL)-3(2-CARBOXYTHIOPHENE-3-YL-METHYL)-5-METHYLPYRIMIDINE-2,4-DIONE, BROMIDE ION, Glutamate receptor, ... | | Authors: | Mayer, M.L. | | Deposit date: | 2007-01-14 | | Release date: | 2007-04-03 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (1.95 Å) | | Cite: | Structure and mechanism of kainate receptor modulation by anions.
Neuron, 53, 2007
|
|
4UQK
 
 | | Electron density map of GluA2em in complex with quisqualate and LY451646 | | Descriptor: | (S)-2-AMINO-3-(3,5-DIOXO-[1,2,4]OXADIAZOLIDIN-2-YL)-PROPIONIC ACID, GLUTAMATE RECEPTOR 2 | | Authors: | Meyerson, J.R, Kumar, J, Chittori, S, Rao, P, Pierson, J, Bartesaghi, A, Mayer, M.L, Subramaniam, S. | | Deposit date: | 2014-06-24 | | Release date: | 2014-08-13 | | Last modified: | 2017-08-02 | | Method: | ELECTRON MICROSCOPY (16.4 Å) | | Cite: | Structural Mechanism of Glutamate Receptor Activation and Desensitization
Nature, 514, 2014
|
|
