2ZQ0
 
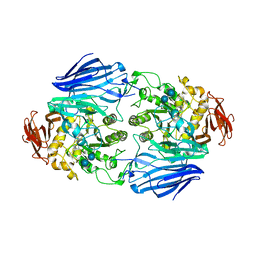 | | Crystal structure of SusB complexed with acarbose | | Descriptor: | 4,6-dideoxy-4-{[(1S,4R,5S,6S)-4,5,6-trihydroxy-3-(hydroxymethyl)cyclohex-2-en-1-yl]amino}-alpha-D-glucopyranose-(1-4)-alpha-D-glucopyranose-(1-4)-alpha-D-glucopyranose, Alpha-glucosidase (Alpha-glucosidase SusB), CALCIUM ION | | Authors: | Yao, M, Tanaka, I, Kitamura, M. | | Deposit date: | 2008-07-31 | | Release date: | 2008-10-28 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (1.6 Å) | | Cite: | Structural and functional analysis of a glycoside hydrolase family 97 enzyme from Bacteroides thetaiotaomicron.
J.Biol.Chem., 283, 2008
|
|
5ZIC
 
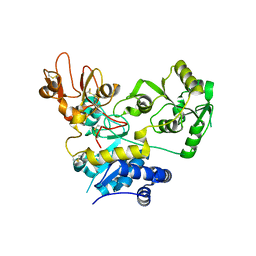 | | Crystal structure of human GnT-V luminal domain in complex with acceptor sugar | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-2)-6-thio-alpha-D-mannopyranose-(1-6)-beta-D-mannopyranose, Alpha-1,6-mannosylglycoprotein 6-beta-N-acetylglucosaminyltransferase A | | Authors: | Nagae, M, Yamaguchi, Y. | | Deposit date: | 2018-03-14 | | Release date: | 2018-08-01 | | Last modified: | 2023-11-22 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Structure and mechanism of cancer-associated N-acetylglucosaminyltransferase-V.
Nat Commun, 9, 2018
|
|
5ZIB
 
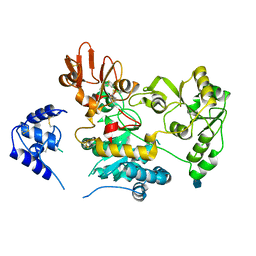 | | Crystal structure of human GnT-V luminal domain in apo form | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, Alpha-1,6-mannosylglycoprotein 6-beta-N-acetylglucosaminyltransferase A | | Authors: | Nagae, M, Yamaguchi, Y. | | Deposit date: | 2018-03-14 | | Release date: | 2018-08-01 | | Last modified: | 2020-07-29 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Structure and mechanism of cancer-associated N-acetylglucosaminyltransferase-V.
Nat Commun, 9, 2018
|
|
5XCX
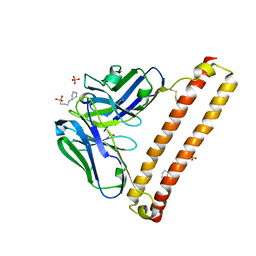 
 | | Crystal structure of TS2/16 Fv-clasp fragment | | Descriptor: | 3-CYCLOHEXYL-1-PROPYLSULFONIC ACID, GLYCEROL, PHOSPHATE ION, ... | | Authors: | Arimori, T, Takagi, J. | | Deposit date: | 2017-03-23 | | Release date: | 2017-10-04 | | Last modified: | 2017-10-18 | | Method: | X-RAY DIFFRACTION (2.04 Å) | | Cite: | Fv-clasp: An Artificially Designed Small Antibody Fragment with Improved Production Compatibility, Stability, and Crystallizability
Structure, 25, 2017
|
|
5XCS
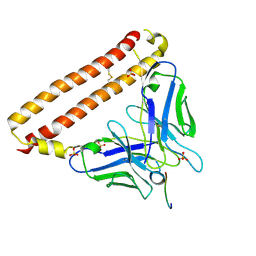 
 | | Crystal structure of 12CA5 Fv-clasp fragment with its antigen peptide | | Descriptor: | HA peptide, SULFATE ION, VH-SARAH(Y35C) chimera, ... | | Authors: | Arimori, T, Takagi, J. | | Deposit date: | 2017-03-23 | | Release date: | 2017-10-04 | | Last modified: | 2023-11-22 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Fv-clasp: An Artificially Designed Small Antibody Fragment with Improved Production Compatibility, Stability, and Crystallizability
Structure, 25, 2017
|
|
5XCQ
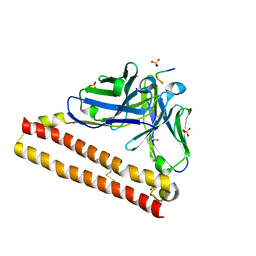 
 | | Crystal structure of P20.1 Fv-clasp fragment with its antigen peptide | | Descriptor: | C8 peptide, SULFATE ION, VH-SARAH(Y35C)chimera, ... | | Authors: | Arimori, T, Takagi, J. | | Deposit date: | 2017-03-23 | | Release date: | 2017-10-04 | | Last modified: | 2023-11-22 | | Method: | X-RAY DIFFRACTION (1.313 Å) | | Cite: | Fv-clasp: An Artificially Designed Small Antibody Fragment with Improved Production Compatibility, Stability, and Crystallizability
Structure, 25, 2017
|
|
5XCU
 
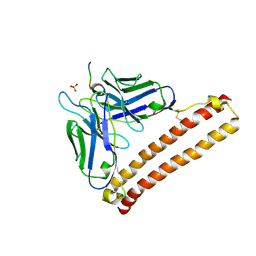 | | Crystal structure of 12CA5 Fv-clasp fragment with its antigen peptide | | Descriptor: | HA peptide, SULFATE ION, VH(S112C)-SARAH chimera,VH(S112C)-SARAH chimera, ... | | Authors: | Arimori, T, Takagi, J. | | Deposit date: | 2017-03-23 | | Release date: | 2017-10-04 | | Last modified: | 2023-11-22 | | Method: | X-RAY DIFFRACTION (2.454 Å) | | Cite: | Fv-clasp: An Artificially Designed Small Antibody Fragment with Improved Production Compatibility, Stability, and Crystallizability
Structure, 25, 2017
|
|
5XCV
 
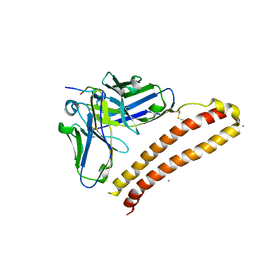 | | Crystal structure of NZ-1 Fv-clasp fragment with its antigen peptide | | Descriptor: | CALCIUM ION, PA14 peptide, VH(S112C)-SARAH chimera,VH(S112C)-SARAH chimera, ... | | Authors: | Arimori, T, Takagi, J. | | Deposit date: | 2017-03-23 | | Release date: | 2017-10-11 | | Last modified: | 2023-11-22 | | Method: | X-RAY DIFFRACTION (2.143 Å) | | Cite: | Fv-clasp: An Artificially Designed Small Antibody Fragment with Improved Production Compatibility, Stability, and Crystallizability
Structure, 25, 2017
|
|
5XCT
 
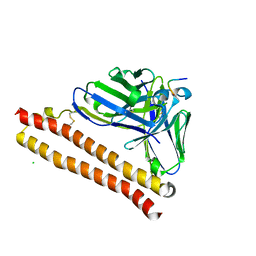 | | Crystal structure of P20.1 Fv-clasp fragment with its antigen peptide | | Descriptor: | C8 peptide, CHLORIDE ION, VH(S112C)-SARAH chimera, ... | | Authors: | Arimori, T, Takagi, J. | | Deposit date: | 2017-03-23 | | Release date: | 2017-10-04 | | Last modified: | 2023-11-22 | | Method: | X-RAY DIFFRACTION (1.17 Å) | | Cite: | Fv-clasp: An Artificially Designed Small Antibody Fragment with Improved Production Compatibility, Stability, and Crystallizability
Structure, 25, 2017
|
|
5XCR
 
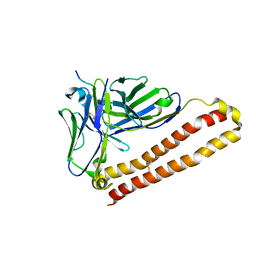 | |
2D73
 
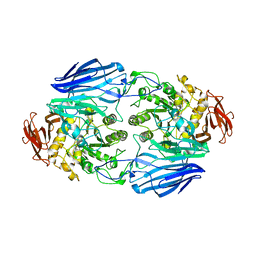 | | Crystal Structure Analysis of SusB | | Descriptor: | CALCIUM ION, alpha-glucosidase SusB | | Authors: | Kitamura, M, Yao, M. | | Deposit date: | 2005-11-15 | | Release date: | 2007-02-27 | | Last modified: | 2024-03-13 | | Method: | X-RAY DIFFRACTION (1.6 Å) | | Cite: | Structural and functional analysis of a glycoside hydrolase family 97 enzyme from Bacteroides thetaiotaomicron.
J.Biol.Chem., 283, 2008
|
|
2D5A
 
 | |
2D59
 
 | |
