1XL8
 
 | | Crystal structure of mouse carnitine octanoyltransferase in complex with octanoylcarnitine | | Descriptor: | (4S)-2-METHYL-2,4-PENTANEDIOL, CARNITINE, OCTANOYLCARNITINE, ... | | Authors: | Jogl, G, Hsiao, Y.S, Tong, L. | | Deposit date: | 2004-09-30 | | Release date: | 2004-10-19 | | Last modified: | 2023-11-15 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Crystal structure of mouse carnitine octanoyltransferase and molecular determinants of substrate selectivity.
J.Biol.Chem., 280, 2005
|
|
1XMC
 
 | | C323M mutant structure of mouse carnitine octanoyltransferase | | Descriptor: | (4S)-2-METHYL-2,4-PENTANEDIOL, 4-(2-HYDROXYETHYL)-1-PIPERAZINE ETHANESULFONIC ACID, Peroxisomal carnitine O-octanoyltransferase | | Authors: | Jogl, G, Hsiao, Y.S, Tong, L. | | Deposit date: | 2004-10-01 | | Release date: | 2004-10-19 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Crystal structure of mouse carnitine octanoyltransferase and molecular determinants of substrate selectivity.
J.Biol.Chem., 280, 2005
|
|
1XL7
 
 | | Crystal Structure of Mouse Carnitine Octanoyltransferase | | Descriptor: | (4S)-2-METHYL-2,4-PENTANEDIOL, 4-(2-HYDROXYETHYL)-1-PIPERAZINE ETHANESULFONIC ACID, Peroxisomal carnitine O-octanoyltransferase | | Authors: | Jogl, G, Hsiao, Y.S, Tong, L. | | Deposit date: | 2004-09-30 | | Release date: | 2004-10-19 | | Last modified: | 2017-10-11 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Crystal structure of mouse carnitine octanoyltransferase and molecular determinants of substrate selectivity.
J.Biol.Chem., 280, 2005
|
|
1XMD
 
 | | M335V mutant structure of mouse carnitine octanoyltransferase | | Descriptor: | (4S)-2-METHYL-2,4-PENTANEDIOL, 4-(2-HYDROXYETHYL)-1-PIPERAZINE ETHANESULFONIC ACID, Peroxisomal carnitine O-octanoyltransferase | | Authors: | Jogl, G, Hsiao, Y.S, Tong, L. | | Deposit date: | 2004-10-01 | | Release date: | 2004-10-19 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Crystal structure of mouse carnitine octanoyltransferase and molecular determinants of substrate selectivity.
J.Biol.Chem., 280, 2005
|
|
1MI1
 
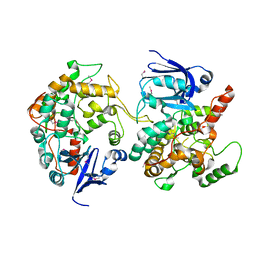 | | Crystal Structure of the PH-BEACH Domain of Human Neurobeachin | | Descriptor: | Neurobeachin | | Authors: | Jogl, G, Shen, Y, Gebauer, D, Li, J, Wiegmann, K, Kashkar, H, Kroenke, M, Tong, L, Northeast Structural Genomics Consortium (NESG) | | Deposit date: | 2002-08-21 | | Release date: | 2002-09-27 | | Last modified: | 2011-07-13 | | Method: | X-RAY DIFFRACTION (2.9 Å) | | Cite: | Crystal structure of the BEACH domain reveals an unusual fold and extensive association with a novel PH domain.
EMBO J., 21, 2002
|
|
1RY2
 
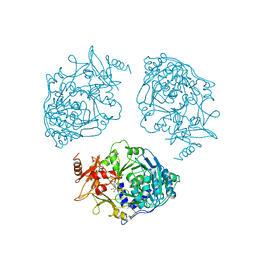 | |
1NF0
 
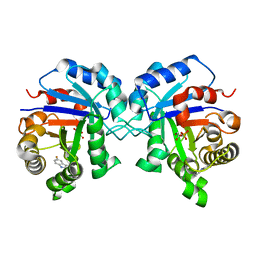 | | Triosephosphate Isomerase in Complex with DHAP | | Descriptor: | 1,3-DIHYDROXYACETONEPHOSPHATE, triosephosphate isomerase | | Authors: | Jogl, G, Rozovsky, S, McDermott, A.E, Tong, L. | | Deposit date: | 2002-12-12 | | Release date: | 2003-01-07 | | Last modified: | 2023-08-16 | | Method: | X-RAY DIFFRACTION (1.6 Å) | | Cite: | Optimal alignment for enzymatic proton transfer: Structure of the
Michaelis complex of triosephosphate isomerase at 1.2-A resolution
Proc.Natl.Acad.Sci.USA, 100, 2003
|
|
1NDI
 
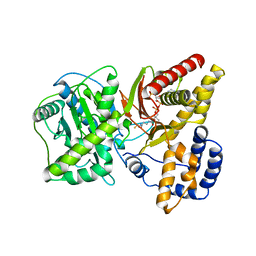 | | Carnitine Acetyltransferase in complex with CoA | | Descriptor: | COENZYME A, Carnitine Acetyltransferase | | Authors: | Jogl, G, Tong, L. | | Deposit date: | 2002-12-09 | | Release date: | 2003-01-28 | | Last modified: | 2023-08-16 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Crystal Structure of Carnitine Acetyltransferase and Implications for the Catalytic Mechanism and Fatty Acid Transport
Cell(Cambridge,Mass.), 112, 2003
|
|
1NDF
 
 | | Carnitine Acetyltransferase in Complex with Carnitine | | Descriptor: | CARNITINE, Carnitine Acetyltransferase | | Authors: | Jogl, G, Tong, L. | | Deposit date: | 2002-12-09 | | Release date: | 2003-01-28 | | Last modified: | 2023-08-16 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Crystal Structure of Carnitine Acetyltransferase and Implications for the Catalytic Mechanism and Fatty Acid Transport
Cell(Cambridge,Mass.), 112, 2003
|
|
1NEY
 
 | | Triosephosphate Isomerase in Complex with DHAP | | Descriptor: | 1,3-DIHYDROXYACETONEPHOSPHATE, triosephosphate isomerase | | Authors: | Jogl, G, Rozovsky, S, McDermott, A.E, Tong, L. | | Deposit date: | 2002-12-12 | | Release date: | 2003-01-07 | | Last modified: | 2023-08-16 | | Method: | X-RAY DIFFRACTION (1.2 Å) | | Cite: | Optimal alignment for enzymatic proton transfer: Structure of the
Michaelis complex of triosephosphate isomerase at 1.2-A resolution.
Proc.Natl.Acad.Sci.USA, 100, 2003
|
|
1NDB
 
 | | Crystal structure of Carnitine Acetyltransferase | | Descriptor: | Carnitine Acetyltransferase | | Authors: | Jogl, G, Tong, L. | | Deposit date: | 2002-12-09 | | Release date: | 2003-01-28 | | Last modified: | 2017-10-11 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Crystal Structure of Carnitine Acetyltransferase and Implications for the Catalytic Mechanism and Fatty Acid Transport
Cell(Cambridge,Mass.), 112, 2003
|
|
6O90
 
 | |
6W6P
 
 | |
6WUB
 
 | |
6WUA
 
 | |
6WU9
 
 | |
6O8W
 
 | |
6O8Y
 
 | |
6O8Z
 
 | |
6O8X
 
 | |
8F43
 
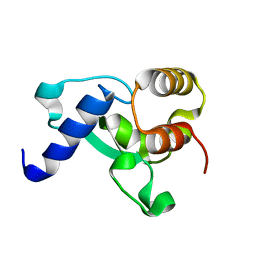 | | HNH Nuclease Domain from G. stearothermophilus Cas9, K597A mutant | | Descriptor: | CRISPR-associated endonuclease Cas9 | | Authors: | D'Ordine, A.M, Belato, H.B, Lisi, G.P, Jogl, G. | | Deposit date: | 2022-11-10 | | Release date: | 2022-12-21 | | Last modified: | 2024-05-01 | | Method: | X-RAY DIFFRACTION (1.37 Å) | | Cite: | Disruption of electrostatic contacts in the HNH nuclease from a thermophilic Cas9 rewires allosteric motions and enhances high-temperature DNA cleavage.
J.Chem.Phys., 157, 2022
|
|
2RDN
 
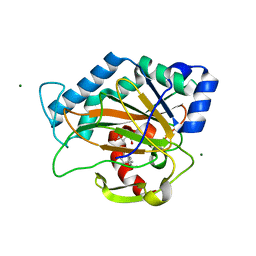 | | Crystal Structure of PtlH with AKG and ent-1PL bound | | Descriptor: | (1S,3aS,5aR,8aS)-1,7,7-trimethyl-1,2,3,3a,5a,6,7,8-octahydrocyclopenta[c]pentalene-4-carboxylic acid, 1-deoxypentalenic acid 11-beta hydroxylase; Fe(II)/alpha-ketoglutarate dependent hydroxylase, 2-OXOGLUTARIC ACID, ... | | Authors: | You, Z, Omura, S, Ikeda, H, Cane, D.E, Jogl, G. | | Deposit date: | 2007-09-24 | | Release date: | 2007-10-16 | | Last modified: | 2024-02-21 | | Method: | X-RAY DIFFRACTION (1.35 Å) | | Cite: | Crystal Structure of the Non-heme Iron Dioxygenase PtlH in Pentalenolactone Biosynthesis.
J.Biol.Chem., 282, 2007
|
|
7MPZ
 
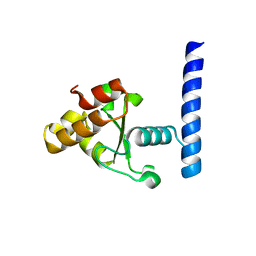 | |
8SCA
 
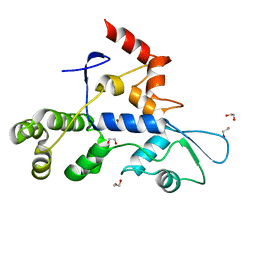 | | Rec3 Domain from S. pyogenes Cas9 | | Descriptor: | 1,2-ETHANEDIOL, CRISPR-associated endonuclease Cas9/Csn1 | | Authors: | D'Ordine, A.M, Skeens, E, Lisi, G.P, Jogl, G. | | Deposit date: | 2023-04-05 | | Release date: | 2024-03-13 | | Method: | X-RAY DIFFRACTION (1.67 Å) | | Cite: | High-fidelity, hyper-accurate, and evolved mutants rewire atomic-level communication in CRISPR-Cas9.
Sci Adv, 10, 2024
|
|
8SP7
 | |
