3KIA
 
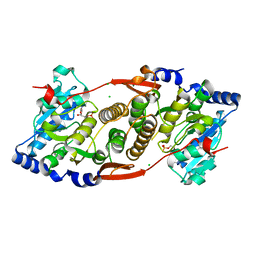 | | Crystal structure of mannosyl-3-phosphoglycerate synthase from Rubrobacter xylanophilus | | Descriptor: | CHLORIDE ION, GUANOSINE-5'-MONOPHOSPHATE, MAGNESIUM ION, ... | | Authors: | Macedo-Ribeiro, S, Pereira, P.J.B, Empadinhas, N, da Costa, M.S. | | Deposit date: | 2009-11-01 | | Release date: | 2010-11-03 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (2.8 Å) | | Cite: | Functional and structural characterization of a novel mannosyl-3-phosphoglycerate synthase from Rubrobacter xylanophilus reveals its dual substrate specificity
Mol.Microbiol., 79, 2011
|
|
3O3P
 
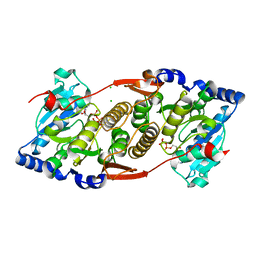 | | Crystal structure of R. xylanophilus MpgS in complex with GDP-Mannose | | Descriptor: | CHLORIDE ION, GUANOSINE-5'-DIPHOSPHATE, GUANOSINE-5'-DIPHOSPHATE-ALPHA-D-MANNOSE, ... | | Authors: | Macedo-Ribeiro, S, Pereira, P.J.B, Empadinhas, N, da Costa, M.S. | | Deposit date: | 2010-07-25 | | Release date: | 2010-11-03 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (2.529 Å) | | Cite: | Functional and structural characterization of a novel mannosyl-3-phosphoglycerate synthase from Rubrobacter xylanophilus reveals its dual substrate specificity
Mol.Microbiol., 79, 2011
|
|
3F1Y
 
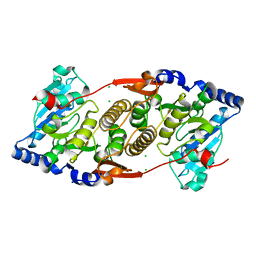 | | Mannosyl-3-phosphoglycerate synthase from Rubrobacter xylanophilus | | Descriptor: | CHLORIDE ION, MAGNESIUM ION, Mannosyl-3-phosphoglycerate synthase | | Authors: | Macedo-Ribeiro, S, Pereira, P.J.B, Empadinhas, N, da Costa, M.S. | | Deposit date: | 2008-10-28 | | Release date: | 2009-11-03 | | Last modified: | 2023-12-27 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Functional and structural characterization of a novel mannosyl-3-phosphoglycerate synthase from Rubrobacter xylanophilus reveals its dual substrate specificity
Mol.Microbiol., 79, 2011
|
|
4WZY
 
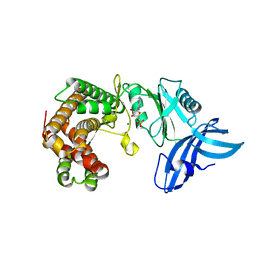 | | Structure of mycobacterial maltokinase, the missing link in the essential GlgE-pathway (ATP complex) | | Descriptor: | ADENOSINE-5'-TRIPHOSPHATE, MAGNESIUM ION, Maltokinase | | Authors: | Fraga, J, Empadinhas, N, Pereira, P.J.B, Macedo-Ribeiro, S. | | Deposit date: | 2014-11-20 | | Release date: | 2015-02-11 | | Last modified: | 2024-01-10 | | Method: | X-RAY DIFFRACTION (1.71 Å) | | Cite: | Structure of mycobacterial maltokinase, the missing link in the essential GlgE-pathway.
Sci Rep, 5, 2015
|
|
4U98
 
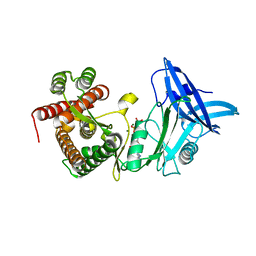 | | Structure of mycobacterial maltokinase, the missing link in the essential GlgE-pathway (AppCp complex) | | Descriptor: | MAGNESIUM ION, Maltokinase, PHOSPHOMETHYLPHOSPHONIC ACID ADENYLATE ESTER | | Authors: | Fraga, J, Empadinhas, N, Pereira, P.J.B, Macedo-Ribeiro, S. | | Deposit date: | 2014-08-05 | | Release date: | 2015-02-11 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (1.15 Å) | | Cite: | Structure of mycobacterial maltokinase, the missing link in the essential GlgE-pathway.
Sci Rep, 5, 2015
|
|
4U94
 
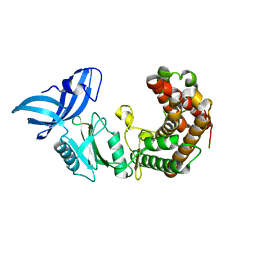 | | Structure of mycobacterial maltokinase, the missing link in the essential GlgE-pathway | | Descriptor: | MAGNESIUM ION, Maltokinase | | Authors: | Fraga, J, Empadinhas, N, Pereira, P.J.B, Macedo-Ribeiro, S. | | Deposit date: | 2014-08-05 | | Release date: | 2015-02-11 | | Last modified: | 2024-05-08 | | Method: | X-RAY DIFFRACTION (1.473 Å) | | Cite: | Structure of mycobacterial maltokinase, the missing link in the essential GlgE-pathway.
Sci Rep, 5, 2015
|
|
5CJ5
 
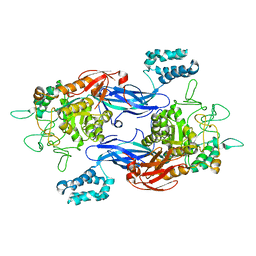 | | Structure of Mycobacterium thermoresistibile GlgE APO form at 3.13A resolution | | Descriptor: | Alpha-1,4-glucan:maltose-1-phosphate maltosyltransferase | | Authors: | Mendes, V, Blaszczyk, M, Maranha, A, Empadinhas, N, Blundell, T.L. | | Deposit date: | 2015-07-13 | | Release date: | 2015-12-09 | | Last modified: | 2024-01-10 | | Method: | X-RAY DIFFRACTION (3.13 Å) | | Cite: | Structure of Mycobacterium thermoresistibile GlgE defines novel conformational states that contribute to the catalytic mechanism.
Sci Rep, 5, 2015
|
|
5CGM
 
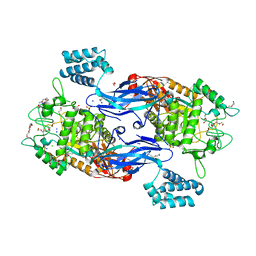 | | Structure of Mycobacterium thermoresistibile GlgE in complex with maltose at 1.95A resolution | | Descriptor: | 1,2-ETHANEDIOL, Alpha-1,4-glucan:maltose-1-phosphate maltosyltransferase, CHLORIDE ION, ... | | Authors: | Mendes, V, Blaszczyk, M, Maranha, A, Empadinhas, N, Blundell, T.L. | | Deposit date: | 2015-07-09 | | Release date: | 2015-12-09 | | Last modified: | 2024-01-10 | | Method: | X-RAY DIFFRACTION (1.95 Å) | | Cite: | Structure of Mycobacterium thermoresistibile GlgE defines novel conformational states that contribute to the catalytic mechanism.
Sci Rep, 5, 2015
|
|
5CIM
 
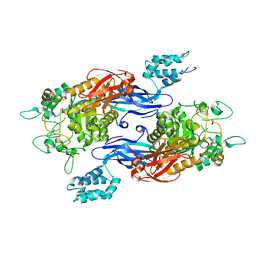 | | Structure of Mycobacterium thermoresistibile GlgE in complex with maltose (cocrystallisation with maltose-1-phosphate) at 3.32A resolution | | Descriptor: | Alpha-1,4-glucan:maltose-1-phosphate maltosyltransferase, alpha-D-glucopyranose-(1-4)-alpha-D-glucopyranose | | Authors: | Mendes, V, Blaszczyk, M, Maranha, A, Empadinhas, N, Blundell, T.L. | | Deposit date: | 2015-07-13 | | Release date: | 2015-12-09 | | Last modified: | 2024-01-10 | | Method: | X-RAY DIFFRACTION (3.32 Å) | | Cite: | Structure of Mycobacterium thermoresistibile GlgE defines novel conformational states that contribute to the catalytic mechanism.
Sci Rep, 5, 2015
|
|
3E26
 
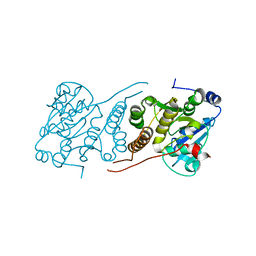 | |
3E25
 
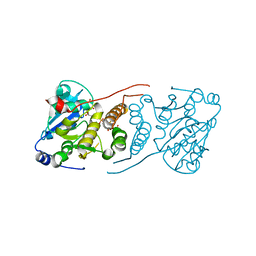 | | Crystal structure of M. tuberculosis glucosyl-3-phosphoglycerate synthase | | Descriptor: | 3-PHOSPHOGLYCERIC ACID, MAGNESIUM ION, Putative uncharacterized protein, ... | | Authors: | Pereira, P.J.B, Empadinhas, N, Costa, M.S, Macedo-Ribeiro, S. | | Deposit date: | 2008-08-05 | | Release date: | 2008-12-02 | | Last modified: | 2024-04-03 | | Method: | X-RAY DIFFRACTION (2.7 Å) | | Cite: | Mycobacterium tuberculosis glucosyl-3-phosphoglycerate synthase: structure of a key enzyme in methylglucose lipopolysaccharide biosynthesis
Plos One, 3, 2008
|
|
6Q5T
 
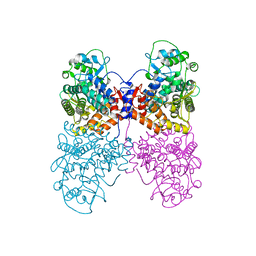 | |
7P5L
 
 | |
7P8G
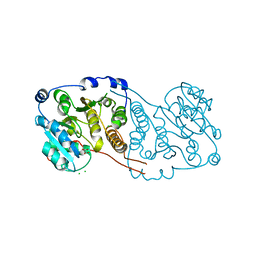 
 | | Crystal structure of Mycobacterium hassiacum glucosyl-3-phosphoglycerate synthase at pH 5.5 - apo form | | Descriptor: | CHLORIDE ION, Glucosyl-3-phosphoglycerate synthase, MALONATE ION, ... | | Authors: | Silva, A, Nunes-Costa, D, Barbosa Pereira, P.J, Macedo-Ribeiro, S. | | Deposit date: | 2021-07-21 | | Release date: | 2023-01-25 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (2.13 Å) | | Cite: | Crystal structure of Mycobacterium hassiacum glucosyl-3-phosphoglycerate synthase at pH 5.5 - apo form
To Be Published
|
|
6S2Q
 
 | |
6S2R
 
 | |
7PVL
 
 | |
7QCP
 
 | |
7QOQ
 
 | | Crystal structure of Mycobacterium hassiacum glucosyl-3-phosphoglycerate synthase at pH 8.5 in complex with UMP and Magnesium | | Descriptor: | CHLORIDE ION, Glucosyl-3-phosphoglycerate synthase, MAGNESIUM ION, ... | | Authors: | Silva, A, Barbosa Pereira, P.J, Macedo-Ribeiro, S, Costa, D. | | Deposit date: | 2021-12-27 | | Release date: | 2023-01-18 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (1.93 Å) | | Cite: | Crystal structure of Mycobacterium hassiacum glucosyl-3-phosphoglycerate synthase at pH 8.5 in complex with UMP and Magnesium
To Be Published
|
|
6G80
 
 | |
6G3N
 
 | |
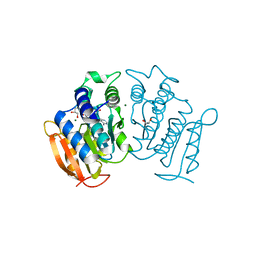
6G7D
 
 | |
5OO2
 
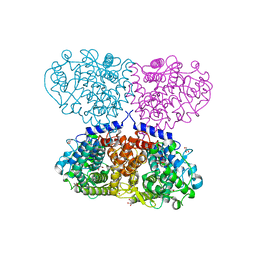 | | Crystal structure of Mycolicibacterium hassiacum glucosylglycerate hydrolase (MhGgH) E419A variant in complex with glucosylglycolate | | Descriptor: | (alpha-D-glucopyranosyloxy)acetic acid, GLYCEROL, SERINE, ... | | Authors: | Cereija, T.B, Macedo-Ribeiro, S, Pereira, P.J.B. | | Deposit date: | 2017-08-05 | | Release date: | 2018-08-29 | | Last modified: | 2024-01-17 | | Method: | X-RAY DIFFRACTION (2.06 Å) | | Cite: | The structural characterization of a glucosylglycerate hydrolase provides insights into the molecular mechanism of mycobacterial recovery from nitrogen starvation.
Iucrj, 6, 2019
|
|
5ONZ
 
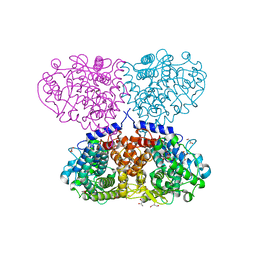 | | Crystal structure of Mycolicibacterium hassiacum glucosylglycerate hydrolase (MhGgH) D182A variant in complex with glucosylglycolate | | Descriptor: | (alpha-D-glucopyranosyloxy)acetic acid, GLYCEROL, Hydrolase, ... | | Authors: | Cereija, T.B, Macedo-Ribeiro, S, Pereira, P.J.B. | | Deposit date: | 2017-08-04 | | Release date: | 2018-08-29 | | Last modified: | 2024-01-17 | | Method: | X-RAY DIFFRACTION (1.93 Å) | | Cite: | The structural characterization of a glucosylglycerate hydrolase provides insights into the molecular mechanism of mycobacterial recovery from nitrogen starvation.
Iucrj, 6, 2019
|
|
5ONT
 
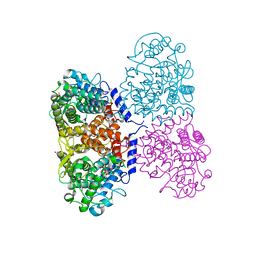 | | Crystal structure of Mycolicibacterium hassiacum glucosylglycerate hydrolase(MhGgH) E419A variant in complex with glucosylglycerol | | Descriptor: | 1,3-dihydroxypropan-2-yl alpha-D-glucopyranoside, GLYCEROL, Uncharacterized protein | | Authors: | Cereija, T.B, Macedo-Ribeiro, S, Pereira, P.J.B. | | Deposit date: | 2017-08-04 | | Release date: | 2018-08-29 | | Last modified: | 2024-01-17 | | Method: | X-RAY DIFFRACTION (2.05 Å) | | Cite: | The structural characterization of a glucosylglycerate hydrolase provides insights into the molecular mechanism of mycobacterial recovery from nitrogen starvation.
Iucrj, 6, 2019
|
|
