6HK5
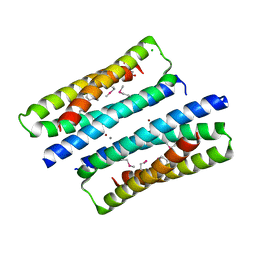 
 | | X-ray structure of a truncated mutant of the metallochaperone CooJ with a high-affinity nickel-binding site | | Descriptor: | 3,3',3''-phosphoryltripropanoic acid, CALCIUM ION, CHLORIDE ION, ... | | Authors: | Alfano, M, Perard, J, Basset, C, Carpentier, P, Zambelli, B, Timm, J, Crouzy, S, Ciurli, S, Cavazza, C. | | Deposit date: | 2018-09-05 | | Release date: | 2019-03-27 | | Last modified: | 2019-10-16 | | Method: | X-RAY DIFFRACTION (2.042 Å) | | Cite: | The carbon monoxide dehydrogenase accessory protein CooJ is a histidine-rich multidomain dimer containing an unexpected Ni(II)-binding site.
J.Biol.Chem., 294, 2019
|
|
2Q0M
 
 | | Tricarbonylmanganese(I)-lysozyme complex : a structurally characterized organometallic protein | | Descriptor: | CARBON MONOXIDE, CHLORIDE ION, Lysozyme C, ... | | Authors: | Razavet, M, Artero, V, Cavazza, C, Oudart, Y, Fontecilla-Camps, J.C, Fontecave, M. | | Deposit date: | 2007-05-22 | | Release date: | 2007-12-11 | | Last modified: | 2023-07-26 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | Tricarbonylmanganese(I)-lysozyme complex: a structurally characterized organometallic protein.
Chem.Commun.(Camb.), 2007
|
|
4DCX
 
 | | X-ray structure of NikA in complex with Fe(1R,2R)-N,N'-Bis(2-pyridylmethyl)-N,N'-dicarboxymethyl-1,2-cyclohexanediamine | | Descriptor: | ACETATE ION, GLYCEROL, Nickel-binding periplasmic protein, ... | | Authors: | Cherrier, M.V, Girgenti, E, Amara, P, Iannello, M, Marchi-Delapierre, C, Fontecilla-Camps, J.C, Menage, S, Cavazza, C. | | Deposit date: | 2012-01-18 | | Release date: | 2012-05-09 | | Last modified: | 2023-09-13 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | The structure of the periplasmic nickel-binding protein NikA provides insights for artificial metalloenzyme design.
J.Biol.Inorg.Chem., 17, 2012
|
|
4DCY
 
 | | X-ray structure of NikA in complex with Fe(1S,2S)-N,N-kappa-Bis(2-pyridylmethyl)-N-carboxymethyl-N-kappa-methyl-1,2-cyclohexanediamine | | Descriptor: | ACETATE ION, GLYCEROL, Nickel-binding periplasmic protein, ... | | Authors: | Cherrier, M.V, Girgenti, E, Amara, P, Iannello, M, Marchi-Delapierre, C, Fontecilla-Camps, J.C, Menage, S, Cavazza, C. | | Deposit date: | 2012-01-18 | | Release date: | 2012-05-09 | | Last modified: | 2023-09-13 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | The structure of the periplasmic nickel-binding protein NikA provides insights for artificial metalloenzyme design.
J.Biol.Inorg.Chem., 17, 2012
|
|
1YQ9
 
 | | Structure of the unready oxidized form of [NiFe] hydrogenase | | Descriptor: | CARBONMONOXIDE-(DICYANO) IRON, FE3-S4 CLUSTER, GLYCEROL, ... | | Authors: | Volbeda, A, Martin, L, Cavazza, C, Matho, M, Faber, B.W, Roseboom, W, Albracht, S.P, Garcin, E, Rousset, M, Fontecilla-Camps, J.C. | | Deposit date: | 2005-02-01 | | Release date: | 2005-04-19 | | Last modified: | 2023-08-23 | | Method: | X-RAY DIFFRACTION (2.35 Å) | | Cite: | Structural differences between the ready and unready oxidized states of [NiFe] hydrogenases.
J.Biol.Inorg.Chem., 10, 2005
|
|
5L8D
 
 | |
3DP8
 
 | | Structural characterization of a putative endogenous metal chelator in the periplasmic nickel transporter NikA (nickel butane-1,2,4-tricarboxylate form) | | Descriptor: | (2R)-butane-1,2,4-tricarboxylic acid, ACETATE ION, CHLORIDE ION, ... | | Authors: | Cherrier, M.V, Cavazza, C, Bochot, C, Lemaire, D, Fontecilla-Camps, J.C. | | Deposit date: | 2008-07-07 | | Release date: | 2008-09-16 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | Structural characterization of a putative endogenous metal chelator in the periplasmic nickel transporter NikA
Biochemistry, 47, 2008
|
|
3E3K
 
 | | Structural characterization of a putative endogenous metal chelator in the periplasmic nickel transporter NikA (butane-1,2,4-tricarboxylate without nickel form) | | Descriptor: | (2R)-butane-1,2,4-tricarboxylic acid, ACETATE ION, CHLORIDE ION, ... | | Authors: | Cherrier, M.V, Cavazza, C, Bochot, C, Lemaire, D, Fontecilla-Camps, J.C. | | Deposit date: | 2008-08-07 | | Release date: | 2008-09-16 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (2.8 Å) | | Cite: | Structural characterization of a putative endogenous metal chelator in the periplasmic nickel transporter NikA
Biochemistry, 47, 2008
|
|
4I9D
 
 | | X-ray structure of NikA in complex with Fe-N,N'-Bis(2-pyridylmethyl)-N-carboxymethyl-N'-methyl | | Descriptor: | ACETATE ION, GLYCEROL, Nickel-binding periplasmic protein, ... | | Authors: | Cherrier, M.V, Amara, P, Iannello, M, Cavazza, C. | | Deposit date: | 2012-12-05 | | Release date: | 2013-04-17 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | An artificial oxygenase built from scratch: substrate binding site identified using a docking approach.
Angew.Chem.Int.Ed.Engl., 52, 2013
|
|
4XKN
 
 | |
4XKP
 
 | |
4XKR
 
 | | Crystal structure of NikA from Staphylococcus aureus in complex with Ni-(L-His)(2-methyl-thiazolidine dicarboxylate) (co-crystallization with Ni(II) and CDdeltaHis medium supernatant) | | Descriptor: | (2R,4R)-2-methyl-1,3-thiazolidine-2,4-dicarboxylic acid, 4-(2-HYDROXYETHYL)-1-PIPERAZINE ETHANESULFONIC ACID, GLYCEROL, ... | | Authors: | Lebrette, H, Cavazza, C. | | Deposit date: | 2015-01-12 | | Release date: | 2015-02-11 | | Last modified: | 2024-01-10 | | Method: | X-RAY DIFFRACTION (1.75 Å) | | Cite: | Novel insights into nickel import in Staphylococcus aureus: the positive role of free histidine and structural characterization of a new thiazolidine-type nickel chelator.
Metallomics, 7, 2015
|
|
4XKQ
 
 | |
1YRQ
 
 | | Structure of the ready oxidized form of [NiFe]-hydrogenase | | Descriptor: | CARBONMONOXIDE-(DICYANO) IRON, FE3-S4 CLUSTER, IRON/SULFUR CLUSTER, ... | | Authors: | Volbeda, A, Martin, L, Cavazza, C, Matho, M, Faber, B.W, Roseboom, W, Albracht, S.P, Garcin, E, Rousset, M, Fontecilla-Camps, J.C. | | Deposit date: | 2005-02-04 | | Release date: | 2005-04-19 | | Last modified: | 2023-12-06 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Structural differences between the ready and unready oxidized states of [NiFe] hydrogenases.
J.Biol.Inorg.Chem., 10, 2005
|
|
1ZLQ
 
 | | Crystallographic and spectroscopic evidence for high affinity binding of Fe EDTA (H2O)- to the periplasmic nickel transporter NikA | | Descriptor: | 2,3-DIHYDROXY-1,4-DITHIOBUTANE, ACETATE ION, CHLORIDE ION, ... | | Authors: | Cherrier, M.V, Martin, L, Cavazza, C, Jacquamet, L, Lemaire, D, Gaillard, J, Fontecilla Camps, J.C. | | Deposit date: | 2005-05-09 | | Release date: | 2005-08-02 | | Last modified: | 2023-08-23 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Crystallographic and Spectroscopic Evidence for High Affinity Binding of FeEDTA(H(2)O)(-) to the Periplasmic Nickel Transporter NikA
J.Am.Chem.Soc., 127, 2005
|
|
4OEU
 
 | | Crystal structure of NikZ from Campylobacter jejuni in complex with Ni(L-His) | | Descriptor: | DI(HYDROXYETHYL)ETHER, GLYCEROL, HISTIDINE, ... | | Authors: | Lebrette, H, Cavazza, C. | | Deposit date: | 2014-01-13 | | Release date: | 2014-10-01 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Promiscuous nickel import in human pathogens: structure, thermodynamics, and evolution of extracytoplasmic nickel-binding proteins.
Structure, 22, 2014
|
|
4OET
 
 | | Crystal structure of NikZ from Campylobacter jejuni, unliganded form | | Descriptor: | GLYCEROL, Putative peptide ABC-transport system periplasmic peptide-binding protein | | Authors: | Lebrette, H, Cavazza, C. | | Deposit date: | 2014-01-13 | | Release date: | 2014-10-01 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | Promiscuous nickel import in human pathogens: structure, thermodynamics, and evolution of extracytoplasmic nickel-binding proteins.
Structure, 22, 2014
|
|
4OFL
 
 | |
4OES
 
 | | Crystal structure of NikA from Brucella suis in complex with Fe(III)-EDTA | | Descriptor: | FE (III) ION, GLYCEROL, NikA protein, ... | | Authors: | Lebrette, H, Cavazza, C. | | Deposit date: | 2014-01-13 | | Release date: | 2014-10-01 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (1.951 Å) | | Cite: | Promiscuous nickel import in human pathogens: structure, thermodynamics, and evolution of extracytoplasmic nickel-binding proteins.
Structure, 22, 2014
|
|
4OER
 
 | | Crystal structure of NikA from Brucella suis, unliganded form | | Descriptor: | GLYCEROL, NikA protein, SULFATE ION | | Authors: | Lebrette, H, Cavazza, C. | | Deposit date: | 2014-01-13 | | Release date: | 2014-10-01 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (1.85 Å) | | Cite: | Promiscuous nickel import in human pathogens: structure, thermodynamics, and evolution of extracytoplasmic nickel-binding proteins.
Structure, 22, 2014
|
|
4OFO
 
 | | Crystal structure of YntA from Yersinia pestis, unliganded form | | Descriptor: | Extracytoplasmic Nickel-Binding Protein YpYntA, NICKEL (II) ION | | Authors: | Lebrette, H, Cavazza, C. | | Deposit date: | 2014-01-15 | | Release date: | 2014-10-01 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (3 Å) | | Cite: | Promiscuous nickel import in human pathogens: structure, thermodynamics, and evolution of extracytoplasmic nickel-binding proteins.
Structure, 22, 2014
|
|
4OEV
 
 | | Crystal structure of NikZ from Campylobacter jejuni in complex with Ni(II) ion | | Descriptor: | GLYCEROL, NICKEL (II) ION, OXALATE ION, ... | | Authors: | Lebrette, H, Cavazza, C. | | Deposit date: | 2014-01-13 | | Release date: | 2014-10-01 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Promiscuous nickel import in human pathogens: structure, thermodynamics, and evolution of extracytoplasmic nickel-binding proteins.
Structure, 22, 2014
|
|
4I8C
 
 | | X-ray structure of NikA in complex with Ni-(L-His)2 | | Descriptor: | ACETATE ION, CHLORIDE ION, GLYCEROL, ... | | Authors: | Lebrette, H, Iannello, M, Fontecilla-Camps, J.C, Cavazza, C. | | Deposit date: | 2012-12-03 | | Release date: | 2013-01-30 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (2.503 Å) | | Cite: | The binding mode of Ni-((L)-His)(2) in NikA revealed by X-ray crystallography.
J.Inorg.Biochem., 121C, 2012
|
|
4KO3
 
 | | Low X-ray dose structure of anaerobically purified Dm. baculatum [NiFeSe]-hydrogenase after crystallization under air | | Descriptor: | CALCIUM ION, CARBONMONOXIDE-(DICYANO) IRON, CHLORIDE ION, ... | | Authors: | Volbeda, A, Cavazza, C, Fontecilla-Camps, J.C. | | Deposit date: | 2013-05-11 | | Release date: | 2013-07-10 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | Structural foundations for the O2 resistance of Desulfomicrobium baculatum [NiFeSe]-hydrogenase.
Chem.Commun.(Camb.), 49, 2013
|
|
4KO2
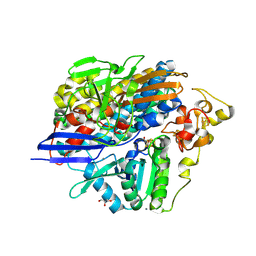 
 | | Low X-ray dose structure of H2-activated anaerobically purified Dm. baculatum [NiFeSe]-hydrogenase after crystallization under air | | Descriptor: | CALCIUM ION, CARBONMONOXIDE-(DICYANO) IRON, GLYCEROL, ... | | Authors: | Volbeda, A, Cavazza, C, Fontecilla-Camps, J.C. | | Deposit date: | 2013-05-11 | | Release date: | 2013-07-10 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (1.6 Å) | | Cite: | Structural foundations for the O2 resistance of Desulfomicrobium baculatum [NiFeSe]-hydrogenase.
Chem. Commun. (Camb.), 49, 2013
|
|
