5TK7
 
 | |
5TKA
 
 | |
5TK6
 
 | |
5UL3
 
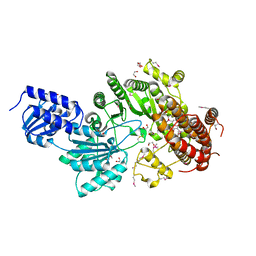
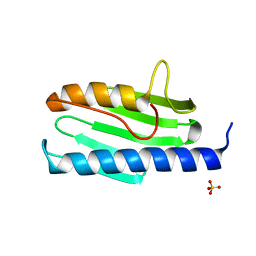
 | | Structure of Cobalamin-dependent S-adenosylmethionine radical enzyme OxsB with aqua-cobalamin bound | | Descriptor: | 1,2-ETHANEDIOL, 2,3-DIHYDROXY-1,4-DITHIOBUTANE, 2-(N-MORPHOLINO)-ETHANESULFONIC ACID, ... | | Authors: | Bridwell-Rabb, J, Drennan, C.L. | | Deposit date: | 2017-01-24 | | Release date: | 2017-04-19 | | Last modified: | 2024-04-03 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | A B12-dependent radical SAM enzyme involved in oxetanocin A biosynthesis.
Nature, 544, 2017
|
|
5UL2
 
 | |
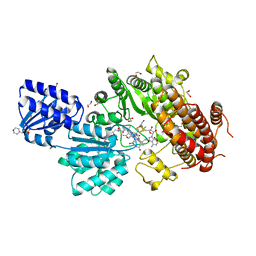
5UL4
 
 | |
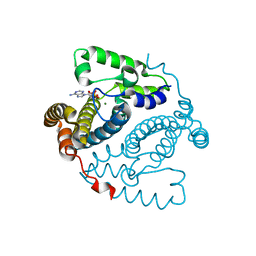
5TK8
 
 | |
5TK9
 
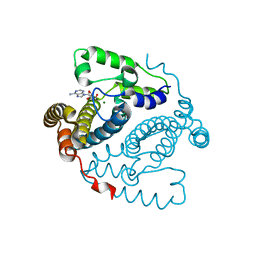 | | Structure of the HD-domain phosphohydrolase OxsA with Oxetanocin-A bound | | Descriptor: | MAGNESIUM ION, OxsA protein, [(2S,3R,4R)-4-(6-amino-9H-purin-9-yl)oxetane-2,3-diyl]dimethanol | | Authors: | Bridwell-Rabb, J, Drennan, C.L. | | Deposit date: | 2016-10-06 | | Release date: | 2016-11-16 | | Last modified: | 2024-03-06 | | Method: | X-RAY DIFFRACTION (1.843 Å) | | Cite: | An HD domain phosphohydrolase active site tailored for oxetanocin-A biosynthesis.
Proc. Natl. Acad. Sci. U.S.A., 113, 2016
|
|
3T3J
 
 | |
3T3L
 
 | |
3T3K
 
 | |
3T3X
 
 | |
3T3T
 
 | |
7SZE
 
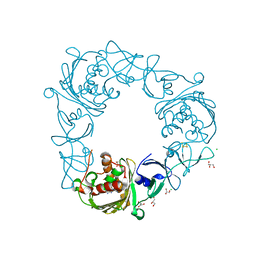 | |
7SZG
 
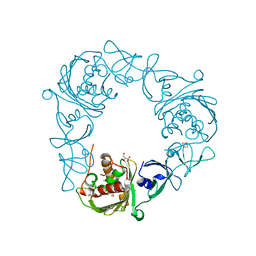 | | Structure of the Rieske Non-heme Iron Oxygenase GxtA Pressurized with Xenon | | Descriptor: | 2,3-DIHYDROXY-1,4-DITHIOBUTANE, CHLORIDE ION, FE (III) ION, ... | | Authors: | Bridwell-Rabb, J, Liu, J. | | Deposit date: | 2021-11-27 | | Release date: | 2022-01-19 | | Last modified: | 2023-10-18 | | Method: | X-RAY DIFFRACTION (2.69 Å) | | Cite: | Design principles for site-selective hydroxylation by a Rieske oxygenase.
Nat Commun, 13, 2022
|
|
7SZF
 
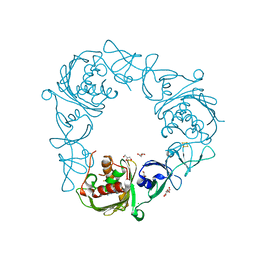 | |
7SZH
 
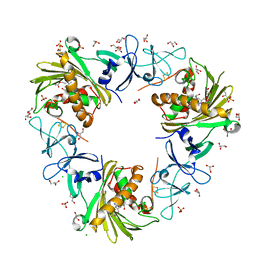 | |
7TXO
 
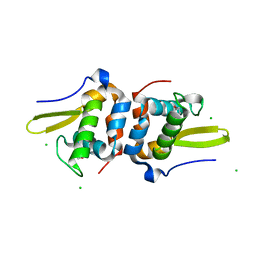 | | Selenomethionine Labeled Structure of RexT | | Descriptor: | CHLORIDE ION, Transcriptional regulator | | Authors: | Bridwell-Rabb, J, Li, B. | | Deposit date: | 2022-02-09 | | Release date: | 2022-03-23 | | Last modified: | 2022-04-13 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | Structural and mechanistic basis for redox sensing by the cyanobacterial transcription regulator RexT.
Commun Biol, 5, 2022
|
|
7TXM
 
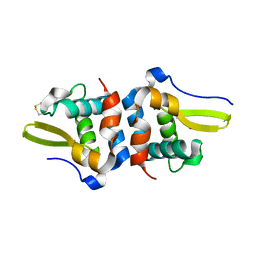 | | Oxidized Structure of RexT | | Descriptor: | HYDROGEN PEROXIDE, Transcriptional regulator | | Authors: | Bridwell-Rabb, J, Li, B. | | Deposit date: | 2022-02-09 | | Release date: | 2022-03-23 | | Last modified: | 2023-10-18 | | Method: | X-RAY DIFFRACTION (2.16 Å) | | Cite: | Structural and mechanistic basis for redox sensing by the cyanobacterial transcription regulator RexT.
Commun Biol, 5, 2022
|
|
7TXN
 
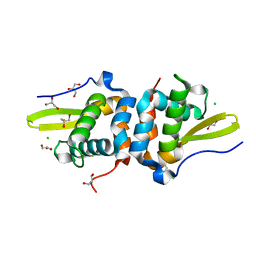 | | Reduced Structure of RexT | | Descriptor: | CHLORIDE ION, GLYCEROL, Transcriptional regulator | | Authors: | Bridwell-Rabb, J, Li, B. | | Deposit date: | 2022-02-09 | | Release date: | 2022-03-23 | | Last modified: | 2023-10-18 | | Method: | X-RAY DIFFRACTION (1.95 Å) | | Cite: | Structural and mechanistic basis for redox sensing by the cyanobacterial transcription regulator RexT.
Commun Biol, 5, 2022
|
|
8EYZ
 
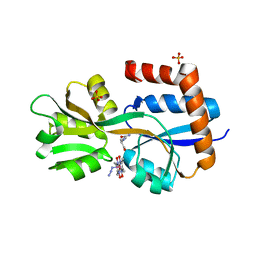 | | Engineered glutamine binding protein bound to GLN and a cobaloxime ligand | | Descriptor: | AZIDOBIS (DIMETHYLGLYOXIMATO) PYRIDINECOBALT, Amino acid ABC transporter substrate-binding protein, GLUTAMINE, ... | | Authors: | Bridwell-Rabb, J, Boggs, D.G, Olshansky, L, Fatima, S, Thompson, P. | | Deposit date: | 2022-10-29 | | Release date: | 2022-11-23 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (2.99 Å) | | Cite: | Engineering a Conformationally Switchable Artificial Metalloprotein.
J.Am.Chem.Soc., 144, 2022
|
|
6WN3
 
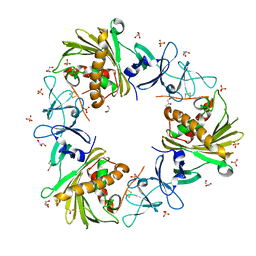 | |
6WNB
 
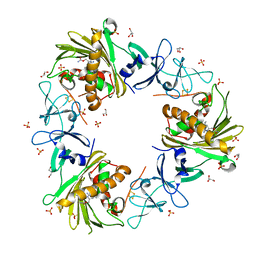 | |
6WNC
 
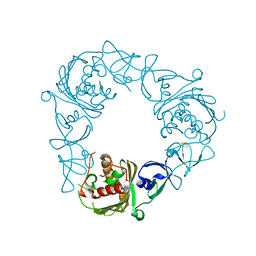 | |
6WND
 
 | |
