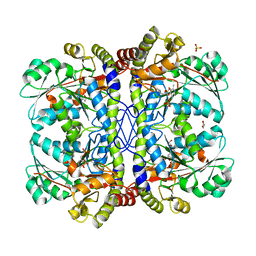
3QHX
 
 | |
3FW5
 | |
3FW4
 | |
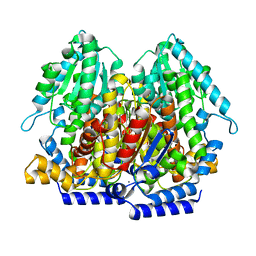
4EM6
 
 | |
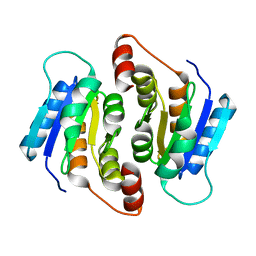
4EM8
 
 | |
4EX4
 
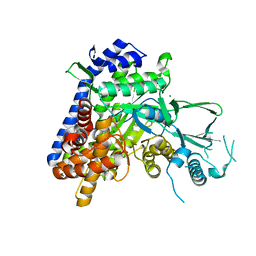 | |
4HR2
 
 | |
4JCP
 
 | |
4JPD
 
 | |
3S6M
 
 | |
3SDO
 
 | |
3S4K
 | |
3T1D
 
 | | The mutant structure of human Siderocalin W79A, R81A, Y106F bound to Enterobactin | | Descriptor: | 2,3-DIHYDROXY-BENZOIC ACID, 2-(2,3-DIHYDROXY-BENZOYLAMINO)-3-HYDROXY-PROPIONIC ACID, CHLORIDE ION, ... | | Authors: | Seattle Structural Genomics Center for Infectious Disease, Seattle Structural Genomics Center for Infectious Disease (SSGCID) | | Deposit date: | 2011-07-21 | | Release date: | 2011-08-17 | | Last modified: | 2023-09-13 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Parsing the functional specificity of Siderocalin/Lipocalin 2/NGAL for siderophores and related small-molecule ligands.
J Struct Biol X, 2, 2019
|
|
3TF6
 
 | |
3TK8
 
 | | Structure of a 2,3,4,5-tetrahydropyridine-2,6-dicarboxylate N-succinyltransferase from Burkholderia pseudomallei | | Descriptor: | 1,2-ETHANEDIOL, 2,3,4,5-tetrahydropyridine-2,6-dicarboxylate N-succinyltransferase, SULFATE ION, ... | | Authors: | Seattle Structural Genomics Center for Infectious Disease (SSGCID) | | Deposit date: | 2011-08-25 | | Release date: | 2011-10-12 | | Last modified: | 2024-02-28 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Structure of a 2,3,4,5-tetrahydropyridine-2,6-dicarboxylate N-succinyltransferase from Burkholderia pseudomallei
TO BE PUBLISHED
|
|
3TV2
 
 | |
3TZS
 
 | |
3UZU
 
 | |
3V7O
 
 | |
4W5K
 
 | |
4WBS
 
 | |
5DWN
 
 | |
4Y0V
 
 | |
4YLG
 
 | |
4ZIY
 
 | |
