3K0O
 
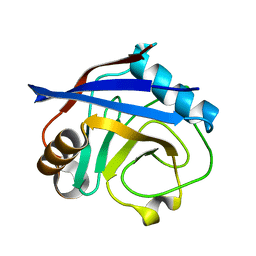 | |
2O7A
 
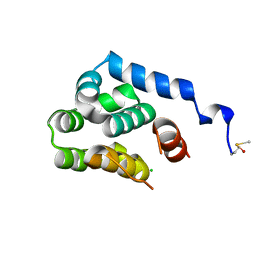 | | T4 lysozyme C-terminal fragment | | Descriptor: | ACETATE ION, CHLORIDE ION, Lysozyme | | Authors: | Echols, N, Kwon, E, Marqusee, S.M, Alber, T. | | Deposit date: | 2006-12-10 | | Release date: | 2007-04-10 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (0.84 Å) | | Cite: | Exploring subdomain cooperativity in T4 lysozyme I: Structural and energetic studies of a circular permutant and protein fragment.
Protein Sci., 16, 2007
|
|
3N6Q
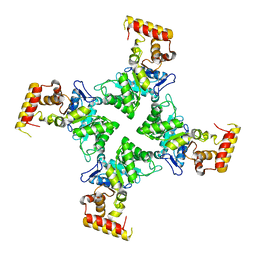 
 | | Crystal structure of YghZ from E. coli | | Descriptor: | MAGNESIUM ION, YghZ aldo-keto reductase | | Authors: | Zubieta, C, Totir, M, Echols, N, May, A, Alber, T. | | Deposit date: | 2010-05-26 | | Release date: | 2011-06-15 | | Last modified: | 2023-09-06 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Macro-to-Micro Structural Proteomics: Native Source Proteins for High-Throughput Crystallization.
Plos One, 7, 2012
|
|
3OUN
 
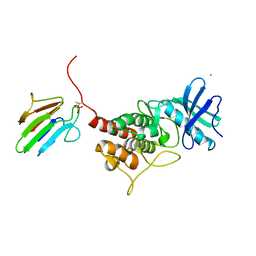 | |
3ORI
 
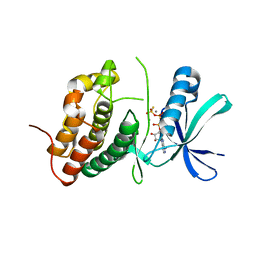 | | Mycobacterium tuberculosis PknB kinase domain L33D mutant (crystal form 1) | | Descriptor: | MANGANESE (II) ION, PHOSPHOTHIOPHOSPHORIC ACID-ADENYLATE ESTER, Serine/threonine protein kinase | | Authors: | Lombana, T.N, Echols, N, Good, M.C, Thomsen, N.D, Ng, H.-L, Alber, T, TB Structural Genomics Consortium (TBSGC) | | Deposit date: | 2010-09-07 | | Release date: | 2010-12-15 | | Last modified: | 2023-09-06 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Allosteric activation mechanism of the Mycobacterium tuberculosis receptor Ser/Thr protein kinase, PknB.
Structure, 18, 2010
|
|
3OUK
 
 | |
3ORL
 
 | | Mycobacterium tuberculosis PknB kinase domain L33D mutant (crystal form 3) | | Descriptor: | MANGANESE (II) ION, PHOSPHOTHIOPHOSPHORIC ACID-ADENYLATE ESTER, Serine/threonine protein kinase | | Authors: | Echols, N, Lombana, T.N, Thomsen, N.D, Ng, H.-L, Alber, T, TB Structural Genomics Consortium (TBSGC) | | Deposit date: | 2010-09-07 | | Release date: | 2010-12-15 | | Last modified: | 2023-09-06 | | Method: | X-RAY DIFFRACTION (2.9 Å) | | Cite: | Allosteric activation mechanism of the Mycobacterium tuberculosis receptor Ser/Thr kinase, PknB
Structure, 18, 2010
|
|
3ORP
 
 | | Mycobacterium tuberculosis PknB kinase domain L33D mutant (crystal form 5) | | Descriptor: | PHOSPHOTHIOPHOSPHORIC ACID-ADENYLATE ESTER, Serine/threonine protein kinase | | Authors: | Good, M.C, Echols, N, Lombana, T.N, Alber, T, TB Structural Genomics Consortium (TBSGC) | | Deposit date: | 2010-09-07 | | Release date: | 2010-12-15 | | Last modified: | 2023-09-06 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Allosteric activation mechanism of the Mycobacterium tuberculosis receptor Ser/Thr kinase, PknB
Structure, 18, 2010
|
|
3OTV
 
 | |
3ORO
 
 | | Mycobacterium tuberculosis PknB kinase domain L33D mutant (crystal form 4) | | Descriptor: | 2-[N-CYCLOHEXYLAMINO]ETHANE SULFONIC ACID, PHOSPHOTHIOPHOSPHORIC ACID-ADENYLATE ESTER, Serine/threonine protein kinase | | Authors: | Good, M.C, Echols, N, Lombana, T.N, Alber, T, TB Structural Genomics Consortium (TBSGC) | | Deposit date: | 2010-09-07 | | Release date: | 2010-12-15 | | Last modified: | 2023-09-06 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Allosteric activation mechanism of the Mycobacterium tuberculosis receptor Ser/Thr protein kinase, PknB.
Structure, 18, 2010
|
|
3OXH
 
 | | Mycobacterium tuberculosis kinase inhibitor homolog RV0577 | | Descriptor: | CHLORIDE ION, PARA-MERCURY-BENZENESULFONIC ACID, RV0577 PROTEIN, ... | | Authors: | Echols, N, Flynn, E.M, Stephenson, S, Ng, H.-L, Alber, T, TB Structural Genomics Consortium (TBSGC) | | Deposit date: | 2010-09-21 | | Release date: | 2011-10-19 | | Last modified: | 2024-02-21 | | Method: | X-RAY DIFFRACTION (1.75 Å) | | Cite: | Structural and Biophysical Characterization of the Mycobacterium tuberculosis Protein Rv0577, a Protein Associated with Neutral Red Staining of Virulent Tuberculosis Strains and Homologue of the Streptomyces coelicolor Protein KbpA.
Biochemistry, 2017
|
|
3ORK
 
 | | Mycobacterium tuberculosis PknB kinase domain L33D mutant (crystal form 2) | | Descriptor: | 2-AMINO-2-HYDROXYMETHYL-PROPANE-1,3-DIOL, MANGANESE (II) ION, PHOSPHOTHIOPHOSPHORIC ACID-ADENYLATE ESTER, ... | | Authors: | Echols, N, Lombana, T.N, Thomsen, N.D, Ng, H.-L, Alber, T, TB Structural Genomics Consortium (TBSGC) | | Deposit date: | 2010-09-07 | | Release date: | 2010-12-15 | | Last modified: | 2023-09-06 | | Method: | X-RAY DIFFRACTION (1.6 Å) | | Cite: | Allosteric activation mechanism of the Mycobacterium tuberculosis receptor Ser/Thr kinase, PknB
Structure, 18, 2010
|
|
3ORM
 
 | | Mycobacterium tuberculosis PknB kinase domain D76A mutant | | Descriptor: | MANGANESE (II) ION, PHOSPHOTHIOPHOSPHORIC ACID-ADENYLATE ESTER, Serine/threonine protein kinase | | Authors: | Echols, N, Lombana, T.N, Alber, T, TB Structural Genomics Consortium (TBSGC) | | Deposit date: | 2010-09-07 | | Release date: | 2010-12-15 | | Last modified: | 2023-09-06 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | Allosteric activation mechanism of the Mycobacterium tuberculosis receptor Ser/Thr protein kinase, PknB.
Structure, 18, 2010
|
|
3ORT
 
 | | Mycobacterium tuberculosis PknB kinase domain L33D mutant (crystal form 6) | | Descriptor: | 2-(N-MORPHOLINO)-ETHANESULFONIC ACID, PHOSPHOTHIOPHOSPHORIC ACID-ADENYLATE ESTER, Serine/threonine protein kinase | | Authors: | Good, M.C, Echols, N, Lombana, T.N, Alber, T, TB Structural Genomics Consortium (TBSGC) | | Deposit date: | 2010-09-07 | | Release date: | 2010-12-15 | | Last modified: | 2023-09-06 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Allosteric activation mechanism of the Mycobacterium tuberculosis receptor Ser/Thr kinase, PknB
Structure, 18, 2010
|
|
3OUV
 
 | |
1L10
 
 | |
1I1J
 
 | | STRUCTURE OF MELANOMA INHIBITORY ACTIVITY PROTEIN: A MEMBER OF A NEW FAMILY OF SECRETED PROTEINS | | Descriptor: | MELANOMA DERIVED GROWTH REGULATORY PROTEIN | | Authors: | Lougheed, J.C, Holton, J.M, Alber, T, Bazan, J.F, Handel, T.M. | | Deposit date: | 2001-02-02 | | Release date: | 2001-05-16 | | Last modified: | 2011-07-13 | | Method: | X-RAY DIFFRACTION (1.39 Å) | | Cite: | Structure of melanoma inhibitory activity protein, a member of a recently identified family of secreted proteins.
Proc.Natl.Acad.Sci.USA, 98, 2001
|
|
1L01
 
 | |
1L30
 
 | |
1L28
 
 | |
1L26
 
 | |
1L32
 
 | |
1L25
 
 | |
1L31
 
 | |
1L29
 
 | |
