-検索条件
-検索結果
検索 (著者・登録者: wong & bh)の結果67件中、1から50件目までを表示しています
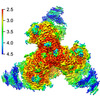
EMDB-28617: 
Cryo-EM structure of HIV-1 BG505 DS-SOSIP ENV trimer bound to VRC34.01 FAB

EMDB-28618: 
Cryo-EM structure of HIV-1 BG505 DS-SOSIP ENV trimer bound to VRC34.01-COMBO1 FAB

EMDB-28619: 
Cryo-EM structure of HIV-1 BG505 DS-SOSIP ENV trimer bound to VRC34.01-MM28 FAB

PDB-8euu: 
Cryo-EM structure of HIV-1 BG505 DS-SOSIP ENV trimer bound to VRC34.01 FAB

PDB-8euv: 
Cryo-EM structure of HIV-1 BG505 DS-SOSIP ENV trimer bound to VRC34.01-COMBO1 FAB

PDB-8euw: 
Cryo-EM structure of HIV-1 BG505 DS-SOSIP ENV trimer bound to VRC34.01-MM28 FAB

EMDB-29725: 
Vaccine-elicited human antibody 2C06 in complex with HIV-1 envelope trimer BG505 DS-SOSIP

EMDB-29731: 
Vaccine-elicited human antibody 2C09 in complex with HIV-1 envelope trimer BG505 DS-SOSIP

PDB-8g4m: 
Vaccine-elicited human antibody 2C06 in complex with HIV-1 envelope trimer BG505 DS-SOSIP

PDB-8g4t: 
Vaccine-elicited human antibody 2C09 in complex with HIV-1 envelope trimer BG505 DS-SOSIP
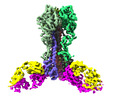
EMDB-27139: 
Cryo-EM structure of the VRC321 clinical trial, vaccine-elicited, human antibody 1B06 in complex with a stabilized NC99 HA trimer

PDB-8d21: 
Cryo-EM structure of the VRC321 clinical trial, vaccine-elicited, human antibody 1B06 in complex with a stabilized NC99 HA trimer
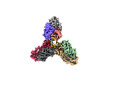
EMDB-28192: 
Cryo-EM structure of a potent anti-malarial antibody L9 in complex with Plasmodium falciparum circumsporozoite protein (PfCSP)(dominant class)

EMDB-28196: 
Cryo-EM structure of a potent anti-malarial antibody L9 in complex with Plasmodium falciparum circumsporozoite protein (PfCSP)(class 2)

PDB-8ek1: 
Cryo-EM structure of a potent anti-malarial antibody L9 in complex with Plasmodium falciparum circumsporozoite protein (PfCSP)(dominant class)

PDB-8eka: 
Cryo-EM structure of a potent anti-malarial antibody L9 in complex with Plasmodium falciparum circumsporozoite protein (PfCSP)(class 2)

EMDB-26044: 
H10ssF: ferritin-based nanoparticle displaying H10 hemagglutinin stabilized stem epitopes

EMDB-27194: 
CIS43 Variant 10 bound to pfCSP Class 4

EMDB-27195: 
CIS43 Variant 10 bound to pfCSP Class 2
EMDB-27202: 
CIS43 Variant 10 bound to pfCSP Class 1

EMDB-27222: 
CIS43 Variant 10 bound to pfCSP Class 3

EMDB-24075: 
Cryo-EM structure of NTD-directed neutralizing antibody LP5 Fab in complex with SARS-CoV-2 S2P spike

PDB-7mxp: 
Cryo-EM structure of NTD-directed neutralizing antibody LP5 Fab in complex with SARS-CoV-2 S2P spike

EMDB-22302: 
Cryo-EM Structure of Vaccine-Elicited Rhesus Antibody 789-203-3C12 in Complex with Stabilized SI06 (A/Solomon Islands/3/06) Influenza Hemagglutinin Trimer

PDB-6xsk: 
Cryo-EM Structure of Vaccine-Elicited Rhesus Antibody 789-203-3C12 in Complex with Stabilized SI06 (A/Solomon Islands/3/06) Influenza Hemagglutinin Trimer

EMDB-23883: 
Single-Particle Cryo-EM Structure of Major Facilitator Superfamily Domain containing 2A in complex with LPC-18:3

EMDB-23521: 
Prefusion RSV F glycoprotein bound by neutralizing site V-directed antibody ADI-14442

PDB-7lue: 
Prefusion RSV F glycoprotein bound by neutralizing site V-directed antibody ADI-14442

EMDB-23520: 
Cryo-EM structure of RSV preF bound by Fabs 32.4K and 01.4B

PDB-7luc: 
Cryo-EM structure of RSV preF bound by Fabs 32.4K and 01.4B

PDB-7l6o: 
Cryo-EM structure of HIV-1 Env CH848.3.D0949.10.17chim.6R.DS.SOSIP.664

EMDB-23518: 
Cryo-EM map of DH851.3 bound to HIV-1 CH505 Env

PDB-7lu9: 
Cryo-EM structure of DH851.3 bound to HIV-1 CH505 Env

EMDB-23519: 
Cryo-EM map of DH898.1 Fab-dimer bound near the CD4 binding site of HIV-1 Env CH848 SOSIP trimer

PDB-7lua: 
Cryo-EM structure of DH898.1 Fab-dimer bound near the CD4 binding site of HIV-1 Env CH848 SOSIP trimer

EMDB-23152: 
Cryo-electron microscopy reconstruction of antibody DH898.1 Fab-dimer bound to glycans 332, 392, and 396 of HIV Env CH848 10.17 SOSIP trimer

EMDB-23153: 
Cryo-electron microscopy local refinement of antibody DH898.1 Fab-dimer bound to glycans 332, 392, and 396 of HIV Env CH848 10.17 SOSIP trimer

EMDB-23016: 
Cryo-EM structure of prefusion SARS-CoV-2 spike glycoprotein in complex with 910-30 Fab

EMDB-23039: 
Cryo-EM map of SARS-CoV-2 spike glycoprotein in complex with 910-30 Fab (disrupted form)

PDB-7ks9: 
Cryo-EM structure of prefusion SARS-CoV-2 spike glycoprotein in complex with 910-30 Fab

EMDB-23124: 
Cryo-electron microcospy reconstruction of CH848.3.D0949.10.17chim.6R.DS.SOSIP.664 HIV Env

EMDB-23145: 
Cryo-electron microscopy reconstruction of locally refined antibody DH898.1 Fab-dimer

PDB-7l6m: 
Cryo-EM structure of DH898.1 Fab-dimer from local refinement of the Fab-dimer bound near the CD4 binding site of HIV-1 Env CH848 SOSIP trimer

EMDB-23094: 
Cryo-EM structure of SARS-CoV-2 2P S ectodomain bound to one copy of domain-swapped antibody 2G12

EMDB-23095: 
Cryo-EM structure of SARS-CoV-2 2P S ectodomain bound to two copies of domain-swapped antibody 2G12

EMDB-23097: 
Cryo-EM structure of SARS-CoV-2 2P S ectodomain bound domain-swapped antibody 2G12 from masked 3D refinement

PDB-7l02: 
Cryo-EM structure of SARS-CoV-2 2P S ectodomain bound to one copy of domain-swapped antibody 2G12

PDB-7l06: 
Cryo-EM structure of SARS-CoV-2 2P S ectodomain bound to two copies of domain-swapped antibody 2G12

PDB-7l09: 
Cryo-EM structure of SARS-CoV-2 2P S ectodomain bound domain-swapped antibody 2G12 from masked 3D refinement

EMDB-22295: 
Cryo-EM structure of SHIV-elicited RHA1.V2.01 in complex with HIV-1 Env BG505 DS-SOSIP.664
ページ:
 ムービー
ムービー コントローラー
コントローラー 構造ビューア
構造ビューア EMN検索について
EMN検索について



 wwPDBはEMDBデータモデルのバージョン3へ移行します
wwPDBはEMDBデータモデルのバージョン3へ移行します
