-検索条件
-検索結果
検索 (著者・登録者: carter & ap)の結果119件中、1から50件目までを表示しています

EMDB-17835: 
Consensus cryo-EM structure of Dynein-Dynactin-JIP3(1-185)-LIS1

EMDB-17836: 
Consensus cryo-EM structure of Dynein-dynactin-JIP3(1-560)-LIS1

EMDB-17825: 
Cytoplasmic dynein-1 motor domain in post-powerstroke state

EMDB-17826: 
Cytoplasmic dynein-1 motor domain bound to dynactin-p150glued and LIS1

EMDB-17828: 
Cytoplasmic dynein-1 motor domain bound to LIS1

EMDB-17829: 
Cytoplasmic dynein-1 A1/A2 motor domains bound to LIS1

EMDB-17830: 
Cytoplasmic dynein-A heavy chain bound to dynactin-p150glued and IC-LC tower

EMDB-17831: 
Cytoplasmic dynein-B heavy chain bound to IC-LC tower

EMDB-17832: 
Cytoplasmic dynein-1 heavy chain bound to JIP3-LZI

EMDB-17833: 
Cytoplasmic dynein-1 heavy chain bound to JIP3-RH1

EMDB-17834: 
Dynactin pointed end bound to JIP3

EMDB-17873: 
Composite structure of Dynein-Dynactin-JIP3-LIS1

EMDB-18121: 
A cryo-ET study of ciliary rootlet organization - Example cell-like tomogram

EMDB-18122: 
A cryo-ET study of ciliary rootlet organization - purified rootlet example 1

EMDB-18123: 
A cryo-ET study of ciliary rootlet organization - purified rootlet example 2

EMDB-18124: 
A cryo-ET study of ciliary rootlet organization - purified rootlet example 3

EMDB-18125: 
A cryo-ET study of ciliary rootlet organization - purified rootlet example 4

EMDB-18475: 
Cryo-electron tomogram of an induced S2 cell protrusion. The cell was treated with 2uM thapsigargin (5h) and with 2.5uM Cytochalasin D (2h).

EMDB-16877: 
Subtomogram averaging structure of cofilactin filament inside microtubule lumen of Drosophila S2 cell protrusion.

EMDB-16685: 
Tomogram of an induced protrusion of a Drosophila S2 cell

EMDB-16693: 
Tomogram of an induced protrusion of a Drosophila S2 cell

EMDB-16695: 
Tomogram of an induced protrusion of a Drosophila S2 alpha-tubulin acetyltransferase knock-out (dTAT KO) cell with a filament inside the microtubule lumen

EMDB-16720: 
Tomogram of an induced protrusion of a Drosophila S2 cell with filaments inside the microtubule lumen.

EMDB-16800: 
Tomogram of an induced protrusion of a cofilin knock-down Drosophila S2 cell with filaments inside the microtubule lumen

EMDB-16811: 
Tomogram of an induced protrusion of a cofilin knock-down Drosophila S2 cell with a filament inside the microtubule lumen.

EMDB-33748: 
Spike_GSAS_6P protomer RBD domain bound with R1-32 Fab and ACE2 with 3:3:3 ratio
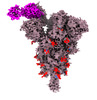
EMDB-33760: 
Spike_GSAS_6P and R1-32 Fab with 3to1 ratio
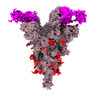
EMDB-33764: 
SARS-COV-2 Spike_GSAS_6P bound with two R1-32 Fabs
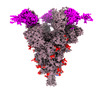
EMDB-33766: 
SARS-CoV-2 Spike (6P) in complex with 3 R1-32 Fabs

EMDB-33772: 
SARS-CoV-2 Spike (6P) in complex with 3 R1-32 Fabs and 3 ACE2

EMDB-14552: 
The barbed end complex of dynactin bound to BICDR1 and the cytoplasmic dynein tails (A2, B1, B2)

EMDB-14549: 
Composite structure of dynein-dynactin-BICDR on microtubules

EMDB-14550: 
Cytoplasmic dynein-1 motor domain

EMDB-14551: 
Cytoplasmic dynein-1 motor domain AAA1, AAA2, and AAA3 subunits

EMDB-14553: 
Cytoplasmic dynein (A2) bound to BICDR1

EMDB-14555: 
Cytoplasmic dynein (A1) bound to BICDR1

EMDB-14556: 
Cytoplasmic dynein light intermediate chain (B1) bound to the motor domain (A2).

EMDB-14559: 
The pointed end complex of dynactin bound to BICDR1

EMDB-15396: 
Consensus map of dynein-dynactin-BICDR on microtubules

EMDB-33453: 
Structure of SARS-CoV-2 Spike Protein with Engineered x3 Disulfide (x3(D427C, V987C) and single Arg S1/S2 cleavage site), Locked-1 Conformation

EMDB-33454: 
Structure of SARS-CoV-2 Spike Protein with Engineered x3 Disulfide (x3(D427C, V987C) and single Arg S1/S2 cleavage site), Locked-211 Conformation

EMDB-33455: 
Structure of SARS-CoV-2 Spike Protein with Engineered x3 Disulfide (x3(D427C, V987C) and single Arg S1/S2 cleavage site), Locked-122 Conformation

EMDB-33456: 
Structure of SARS-CoV-2 Spike Protein with Engineered x3 Disulfide (x3(D427C, V987C) and single Arg S1/S2 cleavage site), Locked-2 Conformation

EMDB-33457: 
Structure of SARS-CoV-2 Spike Protein with Engineered x3 Disulfide (x3(D427C, V987C) and single Arg S1/S2 cleavage site), Closed Conformation

EMDB-33458: 
Structure of SARS-CoV-2 D614G Spike Protein with Engineered x3 Disulfide (x3(D427C, V987C) and single Arg S1/S2 cleavage site), Locked-2 Conformation

EMDB-33459: 
Structure of SARS-CoV-2 D614G Spike Protein with Engineered x3 Disulfide (x3(D427C, V987C) and single Arg S1/S2 cleavage site), Closed Conformation

EMDB-33460: 
Structure of SARS-CoV-2 Spike Protein with Engineered x3 Disulfide (x3(D427C, V987C) and single Arg S1/S2 cleavage site), incubated in Low pH after 40-Day Storage in PBS, Locked-2 Conformation

EMDB-33461: 
Structure of SARS-CoV-2 Spike Protein with Engineered x3 Disulfide (x3(D427C, V987C) and single Arg S1/S2 cleavage site) after 40-Day Storage in PBS, Closed Conformation

EMDB-33462: 
Structure of SARS-CoV-2 Spike Protein with Engineered x3 Disulfide (x3(D427C, V987C) and single Arg S1/S2 cleavage site), incubated in Low pH after 40-Day Storage in PBS, Closed Conformation

EMDB-33463: 
Structure of SARS-CoV-2 D614G spike protein (single Arg S1/S2 cleavage site), Closed Conformation
ページ:
 ムービー
ムービー コントローラー
コントローラー 構造ビューア
構造ビューア EMN検索について
EMN検索について



 wwPDBはEMDBデータモデルのバージョン3へ移行します
wwPDBはEMDBデータモデルのバージョン3へ移行します
