-Search query
-Search result
Showing 1 - 50 of 479 items for (author: tani & k)

EMDB-18664: 
Structure of the native microtubule lattice nucleated from the yeast spindle pole body
Method: subtomogram averaging / : Dendooven T, Yatskevich S, Burt A, Bellini D, Kilmartin J, Barford D

EMDB-18665: 
Structure of the native y-Tubulin Ring Complex (yTuRC) capping microtubule minus ends at the spindle pole body
Method: subtomogram averaging / : Dendooven T, Yatskevich S, Burt A, Bellini D, Kilmartin J, Barford D

EMDB-18666: 
Structure of the y-Tubulin Small Complex (yTuSC) as part of the native y-Tubulin Ring Complex (yTuRC) capping microtubule minus ends at the spindle pole body
Method: subtomogram averaging / : Dendooven T, Yatskevich S, Burt A, Bellini D, Kilmartin J, Barford D

PDB-8qv0: 
Structure of the native microtubule lattice nucleated from the yeast spindle pole body
Method: subtomogram averaging / : Dendooven T, Yatskevich S, Burt A, Bellini D, Kilmartin J, Barford D

PDB-8qv2: 
Structure of the native y-Tubulin Ring Complex (yTuRC) capping microtubule minus ends at the spindle pole body
Method: subtomogram averaging / : Dendooven T, Yatskevich S, Burt A, Bellini D, Kilmartin J, Barford D

PDB-8qv3: 
Structure of the y-Tubulin Small Complex (yTuSC) as part of the native y-Tubulin Ring Complex (yTuRC) capping microtubule minus ends at the spindle pole body
Method: subtomogram averaging / : Dendooven T, Yatskevich S, Burt A, Bellini D, Kilmartin J, Barford D

EMDB-39463: 
2.6A b-Galactosidase Structure Solved Using an Indirect Scintillator-Coupled CMOS Detector at 300 kV
Method: single particle / : Aramaki S, Yoshida Y, Tanihara T, Oyama K, Otsuki K, Terada Y, Matsunaga N, Ohdo S, Mayanagi K

EMDB-39481: 
2.08A Apoferritin Structure Solved Using an Indirect Scintillator-Coupled CMOS Detector at 300 kV
Method: single particle / : Aramaki S, Yoshida Y, Tanihara T, Oyama K, Otsuki K, Terada Y, Matsunaga N, Ohdo S, Mayanagi K

EMDB-17814: 
Structure of the human outer kinetochore KMN network complex
Method: single particle / : Yatskevich S, Barford D

PDB-8ppr: 
Structure of the human outer kinetochore KMN network complex
Method: single particle / : Yatskevich S, Barford D

EMDB-38291: 
Cryo-EM structure of human XKR8-basigin complex in lipid nanodisc
Method: single particle / : Sakuragi TS, Kanai RK, Kikkawa MK, Toyoshima CT, Nagata SN

PDB-8xej: 
Cryo-EM structure of human XKR8-basigin complex in lipid nanodisc
Method: single particle / : Sakuragi TS, Kanai RK, Kikkawa MK, Toyoshima CT, Nagata SN

EMDB-37465: 
Photosynthetic LH1-RC complex from the purple sulfur bacterium Allochromatium vinosum purified by sucrose density
Method: single particle / : Tani K, Kanno R, Harada A, Kobayashi A, Minamino A, Nakamura N, Ji XC, Purba ER, Hall M, Yu LJ, Madigan MT, Mizoguchi A, Iwasaki K, Humbel BM, Kimura Y, Wang-Otomo ZY

EMDB-37466: 
Photosynthetic LH1-RC complex from the purple sulfur bacterium Allochromatium vinosum purified by Ca2+-DEAE
Method: single particle / : Tani K, Kanno R, Harada A, Kobayashi A, Minamino A, Nakamura N, Ji XC, Purba ER, Hall M, Yu LJ, Madigan MT, Mizoguchi A, Iwasaki K, Humbel BM, Kimura Y, Wang-Otomo ZY

PDB-8wdu: 
Photosynthetic LH1-RC complex from the purple sulfur bacterium Allochromatium vinosum purified by sucrose density
Method: single particle / : Tani K, Kanno R, Harada A, Kobayashi A, Minamino A, Nakamura N, Ji XC, Purba ER, Hall M, Yu LJ, Madigan MT, Mizoguchi A, Iwasaki K, Humbel BM, Kimura Y, Wang-Otomo ZY

PDB-8wdv: 
Photosynthetic LH1-RC complex from the purple sulfur bacterium Allochromatium vinosum purified by Ca2+-DEAE
Method: single particle / : Tani K, Kanno R, Harada A, Kobayashi A, Minamino A, Nakamura N, Ji XC, Purba ER, Hall M, Yu LJ, Madigan MT, Mizoguchi A, Iwasaki K, Humbel BM, Kimura Y, Wang-Otomo ZY

EMDB-18387: 
FtsH1 protease from P.aeruginosa clone C in negative stain
Method: single particle / : Mawla GD, Mansour Kamal S, Cao LY, Purhonen P, Hebert H, Sauer RT, Baker TA, Romling U

EMDB-18388: 
FtsH2 protease from P.aeruginosa clone C in negative stain
Method: single particle / : Mawla GD, Mansour Kamal S, Cao LY, Purhonen P, Hebert H, Sauer RT, Baker TA, Romling U

EMDB-18389: 
P.aeruginosa clone C construct PaFtsH2-H1-link32 in negative stain
Method: single particle / : Mawla GD, Mansour Kamal S, Cao LY, Purhonen P, Hebert H, Sauer RT, Baker TA, Romling U

EMDB-36905: 
SARS-CoV-2 BA.1 RBD with UT28-RD
Method: single particle / : Chen L, Kita S, Anraku Y, Maenaka K

EMDB-36906: 
SARS-CoV-2 BA.1 spike with UT28-RD
Method: single particle / : Chen L, Kita S, Anraku Y, Maenaka K

EMDB-16532: 
Structure of Tau filaments Type I from Subacute Sclerosing Panencephalitis
Method: helical / : Qi C, Hasegawa M, Takao M, Sakai M, Akagi M, Iwasaki Y, Yoshida M, Scheres SHW, Goedert M

EMDB-16535: 
Structure of Tau filaments Type II from Subacute Sclerosing Panencephalitis
Method: helical / : Qi C, Hasegawa M, Takao M, Sakai M, Akagi M, Iwasaki Y, Yoshida M, Scheres SHW, Goedert M
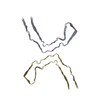
PDB-8caq: 
Structure of Tau filaments Type I from Subacute Sclerosing Panencephalitis
Method: helical / : Qi C, Hasegawa M, Takao M, Sakai M, Akagi M, Iwasaki Y, Yoshida M, Scheres SHW, Goedert M

PDB-8cax: 
Structure of Tau filaments Type II from Subacute Sclerosing Panencephalitis
Method: helical / : Qi C, Hasegawa M, Takao M, Sakai M, Akagi M, Iwasaki Y, Yoshida M, Scheres SHW, Goedert M

EMDB-26583: 
Cryo-EM structure of Antibody 12-16 in complex with prefusion SARS-CoV-2 Spike glycoprotein
Method: single particle / : Casner RG, Shapiro L

PDB-7ukl: 
Cryo-EM structure of Antibody 12-16 in complex with prefusion SARS-CoV-2 Spike glycoprotein
Method: single particle / : Casner RG, Shapiro L

EMDB-35912: 
Cryo-EM structure of the AsCas12f-sgRNA-target DNA ternary complex
Method: single particle / : Hino T, Omura NS, Nakagawa R, Togashi T, Takeda NS, Hiramoto T, Tasaka S, Hirano H, Tokuyama T, Uosaki H, Ishiguro H, Yamano H, Ozaki Y, Motooka D, Mori H, Kirita Y, Kise Y, Itoh Y, Matoba S, Aburatani H, Yachie N, Siksnys V, Ohmori T, Hoshino A, Nureki O

EMDB-35926: 
Cryo-EM structure of the AsCas12f-YHAM-sgRNAS3-5v7-target DNA
Method: single particle / : Hino T, Omura NS, Nakagawa R, Togashi T, Takeda NS, Hiramoto T, Tasaka S, Hirano H, Tokuyama T, Uosaki H, Ishiguro H, Yamano H, Ozaki Y, Motooka D, Mori H, Kirita Y, Kise Y, Itoh Y, Matoba S, Aburatani H, Yachie N, Siksnys V, Ohmori T, Hoshino A, Nureki O

EMDB-35965: 
Cryo-EM structure of the AsCas12f-HKRA-sgRNAS3-5v7-target DNA
Method: single particle / : Hino T, Omura NS, Nakagawa R, Togashi T, Takeda NS, Hiramoto T, Tasaka S, Hirano H, Tokuyama T, Uosaki H, Ishiguro H, Yamano H, Ozaki Y, Motooka D, Mori H, Kirita Y, Kise Y, Itoh Y, Matoba S, Aburatani H, Yachie N, Siksnys V, Ohmori T, Hoshino A, Nureki O

PDB-8j12: 
Cryo-EM structure of the AsCas12f-sgRNA-target DNA ternary complex
Method: single particle / : Hino T, Omura NS, Nakagawa R, Togashi T, Takeda NS, Hiramoto T, Tasaka S, Hirano H, Tokuyama T, Uosaki H, Ishiguro H, Yamano H, Ozaki Y, Motooka D, Mori H, Kirita Y, Kise Y, Itoh Y, Matoba S, Aburatani H, Yachie N, Siksnys V, Ohmori T, Hoshino A, Nureki O

PDB-8j1j: 
Cryo-EM structure of the AsCas12f-YHAM-sgRNAS3-5v7-target DNA
Method: single particle / : Hino T, Omura NS, Nakagawa R, Togashi T, Takeda NS, Hiramoto T, Tasaka S, Hirano H, Tokuyama T, Uosaki H, Ishiguro H, Yamano H, Ozaki Y, Motooka D, Mori H, Kirita Y, Kise Y, Itoh Y, Matoba S, Aburatani H, Yachie N, Siksnys V, Ohmori T, Hoshino A, Nureki O

PDB-8j3r: 
Cryo-EM structure of the AsCas12f-HKRA-sgRNAS3-5v7-target DNA
Method: single particle / : Hino T, Omura NS, Nakagawa R, Togashi T, Takeda NS, Hiramoto T, Tasaka S, Hirano H, Tokuyama T, Uosaki H, Ishiguro H, Yamano H, Ozaki Y, Motooka D, Mori H, Kirita Y, Kise Y, Itoh Y, Matoba S, Aburatani H, Yachie N, Siksnys V, Ohmori T, Hoshino A, Nureki O

EMDB-17600: 
DNA-origami turbine with left-handed chirality
Method: single particle / : Shi X, Pumm AK, Maffeo C, Kohler F, Feigl E, Zhao W, Verschueren D, Golestanian R, Aksimentiev A, Dietz H, Dekker C

EMDB-17606: 
DNA-origami turbine with right-handed chirality
Method: single particle / : Shi X, Pumm AK, Maffeo C, Kohler F, Feigl E, Zhao W, Verschueren D, Golestanian R, Aksimentiev A, Dietz H, Dekker C
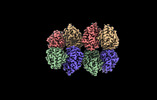
EMDB-28013: 
Cryo-EM structure of the Glutaminase C core filament (fGAC)
Method: single particle / : Ambrosio AL, Dias SM, Quesnay JE, Portugal RV, Cassago A, van Heel MG, Islam Z, Rodrigues CT

PDB-8ec6: 
Cryo-EM structure of the Glutaminase C core filament (fGAC)
Method: single particle / : Ambrosio AL, Dias SM, Quesnay JE, Portugal RV, Cassago A, van Heel MG, Islam Z, Rodrigues CT

EMDB-16677: 
Structure of TDP-43 amyloid filament from type A FTLD-TDP (variant 1)
Method: helical / : Arseni D, Ryskeldi-Falcon B

EMDB-16681: 
Structure of TDP-43 amyloid filaments from type A FTLD-TDP (individual 2, variant 2)
Method: helical / : Arseni D, Ryskeldi-Falcon B

EMDB-16682: 
Structure of TDP-43 amyloid filaments from type A FTLD-TDP (individual 3, variant 1)
Method: helical / : Arseni D, Ryskeldi-Falcon B

EMDB-17224: 
Cryo-EM structure of CBF1-CCAN bound topologically to centromeric DNA
Method: single particle / : Dendooven TD, Zhang Z, Yang J, McLaughlin S, Schwabb J, Scheres S, Yatskevich S, Barford D

EMDB-17225: 
Cryo-EM structure of yeast CENP-OPQU+ bound to the CENP-A N-terminus
Method: single particle / : Dendooven TD, Zhang Z, Yang J, McLaughlin S, Schwabb J, Scheres S, Yatskevich S

EMDB-17226: 
Cryo-EM structure of CBF1-CCAN bound topologically to a centromeric CENP-A nucleosome
Method: single particle / : Dendooven TD, Zhang Z, Yang J, McLaughlin S, Schwabb J, Scheres S, Yatskevich S, Barford D

EMDB-17227: 
Cryo-EM structure of the yeast Inner kinetochore bound to a CENP-A nucleosome.
Method: single particle / : Dendooven TD, Zhang Z, Yang J, McLaughlin S, Schwabb J, Scheres S, Yatskevich S, Barford D

EMDB-17368: 
Multibody map of the CENP-A nucleosome as part of the inner kinetochore
Method: single particle / : Dendooven TD, Zhang Z, Yang J, McLaughlin S, Schwabb J, Scheres S, Yatskevich S, Barford D

EMDB-17371: 
Multibody map of CCAN(Topo) bound to CON3 DNA as part of the Inner kinetochore.
Method: single particle / : Dendooven TD, Zhang Z, Yang J, McLaughlin S, Schwabb J, Scheres S, Yatskevich S, Barford D

EMDB-17372: 
Multibody map of CCAN(Non-Topo) bound to C0N3 DNA as part of the inner kinetochore.
Method: single particle / : Dendooven TD, Zhang Z, Yang J, McLaughlin S, Schwabb J, Scheres S, Yatskevich S, Barford D

EMDB-17374: 
Multibody map of CBF3:CENP-HIK as part of the inner kinetochore
Method: single particle / : Dendooven TD, Zhang Z, Yang J, McLaughlin S, Schwabb J, Scheres S, Yatskevich S, Barford D

EMDB-17376: 
Consensus map of the yeast inner kinetochore
Method: single particle / : Dendooven TD, Zhang Z, Yang J, McLaughlin S, Schwabb J, Scheres S, Yatskevich S, Barford D

PDB-8ovw: 
Cryo-EM structure of CBF1-CCAN bound topologically to centromeric DNA
Method: single particle / : Dendooven TD, Zhang Z, Yang J, McLaughlin S, Schwabb J, Scheres S, Yatskevich S, Barford D
Pages:
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search



 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
