-Search query
-Search result
Showing 1 - 50 of 379 items for (author: nguyen & f)

EMDB-40918: 
Human glutaminase C (Y466W) with L-Gln and Pi, filamentous form
Method: helical / : Feng S, Aplin C, Nguyen TTT, Milano SK, Cerione RA

EMDB-40920: 
Human liver-type glutaminase (Apo form)
Method: single particle / : Feng S, Aplin C, Nguyen TTT, Milano SK, Cerione RA
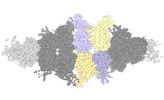
EMDB-40950: 
Human liver-type glutaminase (K253A) with L-Gln, filamentous form
Method: helical / : Feng S, Aplin C, Nguyen TTT, Milano SK, Cerione RA

EMDB-43533: 
Human liver-type glutaminase, bound with inhibitor Compound 968
Method: single particle / : Feng S, Aplin C, Nguyen TTT, Milano SK, Cerione RA

PDB-8szj: 
Human glutaminase C (Y466W) with L-Gln and Pi, filamentous form
Method: helical / : Feng S, Aplin C, Nguyen TTT, Milano SK, Cerione RA

PDB-8szl: 
Human liver-type glutaminase (Apo form)
Method: single particle / : Feng S, Aplin C, Nguyen TTT, Milano SK, Cerione RA

PDB-8t0z: 
Human liver-type glutaminase (K253A) with L-Gln, filamentous form
Method: helical / : Feng S, Aplin C, Nguyen TTT, Milano SK, Cerione RA

EMDB-29907: 
Structure of human NDS.1 Fab and 1G01 Fab in complex with influenza virus neuraminidase from A/Indiana/10/2011 (H3N2v); consensus map with only Fab 1G01 resolved
Method: single particle / : Tsybovsky Y, Lederhofer J, Kwong PD, Kanekiyo M

EMDB-29908: 
Structure of human NDS.1 Fab and 1G01 Fab in complex with influenza virus neuraminidase from A/Indiana/10/2011 (H3N2v), locally refined map
Method: single particle / : Tsybovsky Y, Lederhofer J, Kwong PD, Kanekiyo M

EMDB-29909: 
Structure of human NDS.3 Fab in complex with influenza virus neuraminidase from A/Darwin/09/2021 (H3N2)
Method: single particle / : Tsybovsky Y, Lederhofer J, Kwong PD, Kanekiyo M

PDB-8gat: 
Structure of human NDS.1 Fab and 1G01 Fab in complex with influenza virus neuraminidase from A/Indiana/10/2011 (H3N2v), based on consensus cryo-EM map with only Fab 1G01 resolved
Method: single particle / : Tsybovsky Y, Lederhofer J, Kwong PD, Kanekiyo M

PDB-8gau: 
Structure of human NDS.1 Fab and 1G01 Fab in complex with influenza virus neuraminidase from A/Indiana/10/2011 (H3N2v)
Method: single particle / : Tsybovsky Y, Lederhofer J, Kwong PD, Kanekiyo M

PDB-8gav: 
Structure of human NDS.3 Fab in complex with influenza virus neuraminidase from A/Darwin/09/2021 (H3N2)
Method: single particle / : Tsybovsky Y, Lederhofer J, Kwong PD, Kanekiyo M

EMDB-41133: 
Autographa californica multiple nucleopolyhedrovirus VP39
Method: helical / : Benning FMC, Chao LH

EMDB-41171: 
Cryo-EM structure of cardiac amyloid fibril from a variant ATTR I84S amyloidosis patient-3, wild-type morphology
Method: helical / : Nguyen BA, Singh V, Saelices L

EMDB-41172: 
Cryo-EM structure of cardiac amyloid fibril from a variant ATTR I84S amyloidosis patient-3, variant-type morphology
Method: helical / : Nguyen BA, Singh V, Saelices L
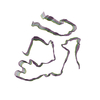
PDB-8tdn: 
Cryo-EM structure of cardiac amyloid fibril from a variant ATTR I84S amyloidosis patient-3, wild-type morphology
Method: helical / : Nguyen BA, Singh V, Saelices L
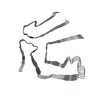
PDB-8tdo: 
Cryo-EM structure of cardiac amyloid fibril from a variant ATTR I84S amyloidosis patient-3, variant-type morphology
Method: helical / : Nguyen BA, Singh V, Saelices L

EMDB-29980: 
Cryo-EM structure of serine 87 O-GlcNAc-modified alpha-synuclein fibrils
Method: helical / : Balana JA, Nguyen AB, Saelices L, Pratt RM

PDB-8gf7: 
Cryo-EM structure of serine 87 O-GlcNAc-modified alpha-synuclein fibrils
Method: helical / : Balana JA, Nguyen AB, Saelices L, Pratt RM

EMDB-29783: 
Cryo-EM structure of T/F100 SOSIP.664 HIV-1 Env trimer with LMHS mutations in complex with 8ANC195 and 10-1074
Method: single particle / : Chen Y, Zhou F, Huang R, Tolbert W, Pazgier M

EMDB-41613: 
Cryo-EM structure of BG505 SOSIP.664 HIV-1 Env trimer in complex with temsavir, 8ANC195, and 10-1074
Method: single particle / : Tolbert WD, Pozharski E, Pazgier M

PDB-8g6u: 
Cryo-EM structure of T/F100 SOSIP.664 HIV-1 Env trimer with LMHS mutations in complex with 8ANC195 and 10-1074
Method: single particle / : Chen Y, Zhou F, Huang R, Tolbert W, Pazgier M

PDB-8ttw: 
Cryo-EM structure of BG505 SOSIP.664 HIV-1 Env trimer in complex with temsavir, 8ANC195, and 10-1074
Method: single particle / : Tolbert WD, Pozharski E, Pazgier M

EMDB-27031: 
Accurate computational design of genetically encoded 3D protein crystals
Method: single particle / : Li Z, Borst AJ, Baker D

EMDB-40926: 
CryoEM Structure of Computationally Designed Nanocage O32-ZL4
Method: single particle / : Weidle C, Borst A

PDB-8cwy: 
Accurate computational design of genetically encoded 3D protein crystals
Method: single particle / : Li Z, Borst AJ, Baker D

PDB-8szz: 
CryoEM Structure of Computationally Designed Nanocage O32-ZL4
Method: single particle / : Weidle C, Borst A

EMDB-41994: 
KCTD5/Cullin3/Gbeta1gamma2 Complex: Local Refinement of KCTD5(CTD)/Gbeta1gamma2
Method: single particle / : Nguyen DM, Narayanan N, Kuntz DA, Prive GG

EMDB-41995: 
KCTD5/Cullin3/Gbeta1gamma2 Complex: Local Refinement of KCTD5(BTB)/Cullin3(NTD)
Method: single particle / : Nguyen DM, Narayanan N, Kuntz DA, Prive GG

EMDB-41996: 
KCTD5/Cullin3/Gbeta1gamma2 Complex State A: Composite Map from RELION Multi-body Refinement
Method: single particle / : Nguyen DM, Narayanan N, Kuntz DA, Prive GG

EMDB-41997: 
KCTD5/Cullin3/Gbeta1gamma2 Complex State A: Reference Map for Composite Map EMD-41996
Method: single particle / : Nguyen DM, Narayanan N, Kuntz DA, Prive GG

EMDB-41998: 
KCTD5/Cullin3/Gbeta1gamma2 Complex State A Body 1: RELION Multi-body Refinement of KCTD5(CTD)/Gbeta1gamma2
Method: single particle / : Nguyen DM, Narayanan N, Kuntz DA, Prive GG

EMDB-41999: 
KCTD5/Cullin3/Gbeta1gamma2 Complex State A Body 2: RELION Multi-body Refinemnent of KCTD5(BTB)/Cullin3(NTD)
Method: single particle / : Nguyen DM, Narayanan N, Kuntz DA, Prive GG

EMDB-42000: 
KCTD5/Cullin3/Gbeta1gamma2 Complex State B: Composite Map from RELION Multi-body Refinement
Method: single particle / : Nguyen DM, Narayanan N, Kuntz DA, Prive GG

EMDB-42001: 
KCTD5/Cullin3/Gbeta1gamma2 Complex State B: Reference Map for Composite Map EMD-42000
Method: single particle / : Nguyen DM, Narayanan N, Kuntz DA, Prive GG

EMDB-42002: 
KCTD5/Cullin3/Gbeta1gamma2 Complex State B Body 1: RELION Multi-body Refinement of KCTD5(CTD)/Gbeta1gamma2
Method: single particle / : Nguyen DM, Narayanan N, Kuntz DA, Prive GG

EMDB-42003: 
KCTD5/Cullin3/Gbeta1gamma2 Complex State B Body 2: RELION Multi-body Refinement of KCTD5(BTB)/Cullin3(NTD)
Method: single particle / : Nguyen DM, Narayanan N, Kuntz DA, Prive GG

EMDB-42004: 
KCTD5/Cullin3/Gbeta1gamma2 Complex State C: Composite Map from RELION Multi-body Refinement
Method: single particle / : Nguyen DM, Narayanan N, Kuntz DA, Prive GG

EMDB-42005: 
KCTD5/Cullin3/Gbeta1gamma2 Complex State C: Reference Map for Composite Map EMD-42004
Method: single particle / : Nguyen DM, Narayanan N, Kuntz DA, Prive GG

EMDB-42006: 
KCTD5/Cullin3/Gbeta1gamma2 Complex State C Body 1: RELION Multi-body Refinement of KCTD5(CTD)/Gbeta1gamma2
Method: single particle / : Nguyen DM, Narayanan N, Kuntz DA, Prive GG

EMDB-42007: 
KCTD5/Cullin3/Gbeta1gamma2 Complex State C Body 2: RELION Multi-body Refinement of KCTD5(BTB)/Cullin3(NTD)
Method: single particle / : Nguyen DM, Narayanan N, Kuntz DA, Prive GG

EMDB-42008: 
KCTD5/Cullin3/Gbeta1gamma2 Complex State D: Composite Map from RELION Multi-body Refinement
Method: single particle / : Nguyen DM, Narayanan N, Kuntz DA, Prive GG

EMDB-42009: 
KCTD5/Cullin3/Gbeta1gamma2 Complex State D: Reference Map for Composite Map EMD-42008
Method: single particle / : Nguyen DM, Narayanan N, Kuntz DA, Prive GG

EMDB-42010: 
KCTD5/Cullin3/Gbeta1gamma2 Complex State D Body 1: RELION Multi-body Refinement of KCTD5(CTD)/Gbeta1gamma2
Method: single particle / : Nguyen DM, Narayanan N, Kuntz DA, Prive GG

EMDB-42011: 
KCTD5/Cullin3/Gbeta1gamma2 Complex State D Body 2: RELION Multi-body Refinemnent of KCTD5(BTB)/Cullin3(NTD)
Method: single particle / : Nguyen DM, Narayanan N, Kuntz DA, Prive GG
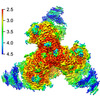
EMDB-28617: 
Cryo-EM structure of HIV-1 BG505 DS-SOSIP ENV trimer bound to VRC34.01 FAB
Method: single particle / : Pletnev S, Kwong P

EMDB-28618: 
Cryo-EM structure of HIV-1 BG505 DS-SOSIP ENV trimer bound to VRC34.01-COMBO1 FAB
Method: single particle / : Pletnev S, Kwong P

EMDB-28619: 
Cryo-EM structure of HIV-1 BG505 DS-SOSIP ENV trimer bound to VRC34.01-MM28 FAB
Method: single particle / : Pletnev S, Kwong P

PDB-8euu: 
Cryo-EM structure of HIV-1 BG505 DS-SOSIP ENV trimer bound to VRC34.01 FAB
Method: single particle / : Pletnev S, Kwong P
Pages:
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search



 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
