-Search query
-Search result
Showing 1 - 50 of 360 items for (author: li & jw)

EMDB-43161: 
Human transthyretin covalently modified with A2-derived stilbene in the canonical conformation
Method: single particle / : Basanta B, Nugroho K, Yan N, Kline GM, Tsai FJ, Wu M, Kelly JW, Lander GC

EMDB-43162: 
Unliganded human transthyretin in the canonical conformation
Method: single particle / : Basanta B, Nugroho K, Yan N, Kline GM, Tsai FJ, Wu M, Kelly JW, Lander GC

EMDB-43163: 
Unliganded human transthyretin in the compressed conformation
Method: single particle / : Basanta B, Nugroho K, Yan N, Kline GM, Tsai FJ, Wu M, Kelly JW, Lander GC

EMDB-43164: 
Unliganded human transthyretin in the frayed conformation
Method: single particle / : Basanta B, Nugroho K, Yan N, Kline GM, Tsai FJ, Wu M, Kelly JW, Lander GC

EMDB-43165: 
(Biarylamine-FT2-WT)1(C10A)3-human transthyretin in the compressed conformation
Method: single particle / : Basanta B, Nugroho K, Yan N, Kline GM, Tsai FJ, Wu M, Kelly JW, Lander GC

EMDB-43166: 
(Biarylamine-FT2-WT)1(C10A)3-human transthyretin in the frayed conformation
Method: single particle / : Basanta B, Nugroho K, Yan N, Kline GM, Tsai FJ, Wu M, Kelly JW, Lander GC

EMDB-18889: 
Mouse teneurin-3 compact dimer - A1B1 isoform
Method: single particle / : Gogou C, Meijer DH

EMDB-18890: 
Mouse teneurin-3 non-compact subunit - A1B1 isoform
Method: single particle / : Gogou C, Meijer DH

EMDB-18891: 
Mouse teneurin-3 non-compact subunit - A0B0 isoform
Method: single particle / : Gogou C, Meijer DH

EMDB-19409: 
Mouse teneurin-3 compact dimer - A1B0 isoform
Method: single particle / : Gogou C, Meijer DH

EMDB-40861: 
Cryo-EM of the GDP-bound human dynamin polymer assembled on the membrane in the super constricted state
Method: helical / : Jimah JR, Canagarajah BJ, Hinshaw JE
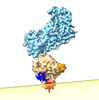
EMDB-40901: 
Cryo-EM of the GDP-bound human dynamin polymer assembled on the membrane in the super constricted state showing the PH domain
Method: helical / : Jimah JR, Canagarajah BJ, Hinshaw JE
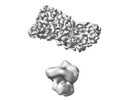
EMDB-40902: 
Cryo-EM of the GDP-bound human dynamin polymer assembled on the membrane in the super constricted state showing the second PH domain
Method: helical / : Jimah JR, Canagarajah BJ, Hinshaw JE

EMDB-40903: 
Cryo-EM of the GDP-bound human dynamin polymer assembled on the membrane in the super constricted state (full helix)
Method: helical / : Jimah JR, Canagarajah BJ, Hinshaw JE
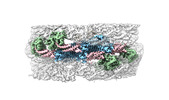

EMDB-40942: 
Cryo-EM of the GDP-bound human dynamin (full-length) polymer assembled on the membrane in the super constricted state
Method: helical / : Jimah JR, Canagarajah BJ, Hinshaw JE

EMDB-40946: 
Cryo-EM of the GDP-bound human dynamin (full-length) polymer assembled on the membrane in the super constricted state (full helix)
Method: helical / : Jimah JR, Canagarajah BJ, Hinshaw JE

EMDB-19042: 
MAP7 MTBD (microtubule binding domain) decorated microtubule protofilament
Method: single particle / : Bangera M, Moores CA

EMDB-19043: 
MAP7 MTBD (microtubule binding domain) decorated Taxol stabilized microtubule (13pf) protofilament
Method: single particle / : Bangera M, Moores CA

EMDB-19044: 
MAP7 MTBD (microtubule binding domain) decorated Taxol stabilized microtubule (14pf) protofilament
Method: single particle / : Bangera M, Moores CA

EMDB-19194: 
In situ cryo-electron tomogram of an autophagosome in the projection of an iPSC-derived neuron #2
Method: electron tomography / : Hoyer MJ, Capitanio C, Smith IR, Paoli JC, Bieber A, Jiang Y, Paulo JA, Gonzalez-Lozano MA, Baumeister W, Wilfling F, Schulman BA, Harper WJ
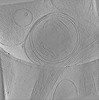

EMDB-19346: 
In situ cryo-electron tomogram of an autophagosome in the projection of an iPSC-derived neuron #1
Method: electron tomography / : Hoyer MJ, Capitanio C, Smith IR, Paoli JC, Bieber A, Jiang Y, Paulo JA, Gonzalez-Lozano MA, Baumeister W, Wilfling F, Schulman BA, Harper WJ

EMDB-43160: 
Human transthyretin covalently modified with A2-derived stilbene in the compressed conformation
Method: single particle / : Basanta B, Nugroho K, Yan N, Kline GM, Tsai FJ, Wu M, Kelly JW, Lander GC

EMDB-40783: 
MBP-tagged fusion fatty acid synthase from S. cerevisiae
Method: single particle / : Mazhab-Jafari MT, Khalili Samani E

EMDB-40784: 
MBP tagged and Mixed chain fatty acid synthase, FAS2_deltaACP
Method: single particle / : Mazhab-Jafari MT, Khalili Samani E

EMDB-40785: 
MBP tagged mixed chain FAS, fusFAS_deltaACP
Method: single particle / : Mazhab-Jafari MT, Khalili Samani E

EMDB-37203: 
Cryo-EM structure of HSV-1 gB with D48 Fab complex
Method: single particle / : Yang J, Sun C, Fang X, Zeng M, Liu Z

EMDB-18900: 
Mouse teneurin-3 non-compact subunit - A0B1 isoform
Method: single particle / : Gogou C, Meijer DH

EMDB-18902: 
Mouse teneurin-3 non-compact subunit - A1B0 isoform
Method: single particle / : Gogou C, Meijer DH

EMDB-35775: 
The rice Na+/H+ antiporter SOS1 in an auto-inhibited state
Method: single particle / : Zhang XY, Tang LH, Zhang CR, Nie JW

EMDB-35950: 
The truncated rice Na+/H+ antiporter SOS1 (1-976) in a constitutively active state
Method: single particle / : Zhang XY, Tang LH, Zhang CR, Nie JW

EMDB-29292: 
HIV-1 envelope trimer bound to one CD4 molecule on membranes
Method: subtomogram averaging / : Li W, Qin Z, Nand E, Grunst MW, Grover JR, Uchil PD, Mothes W

EMDB-29293: 
HIV-1 envelope trimer bound to two CD4 molecules on membranes
Method: subtomogram averaging / : Li W, Qin Z, Nand E, Grunst MW, Grover JR, Uchil PD, Mothes W

EMDB-29294: 
HIV-1 envelope trimer bound to three CD4 molecules on membranes
Method: subtomogram averaging / : Li W, Qin Z, Nand E, Grunst MW, Grover JR, Uchil PD, Mothes W

EMDB-29295: 
Consensus structure of HIV-1 envelope trimer bound to CD4 receptors on membranes
Method: subtomogram averaging / : Li W, Qin Z, Nand E, Grunst MW, Grover JR, Uchil PD, Mothes W

EMDB-41045: 
Unliganded HIV-1 Env on virion
Method: subtomogram averaging / : Li W, Qin Z, Nand E, Grunst MW, Grover JR, Uchil PD, Mothes W

EMDB-28156: 
Cryo-EM structure of human DNMT3B homo-tetramer (form I)
Method: single particle / : Lu JW, Song JK

EMDB-28157: 
Cryo-EM structure of human DNMT3B homo-tetramer (form II)
Method: single particle / : Lu JW, Song JK

EMDB-28158: 
Cryo-EM structure of human DNMT3B homo-trimer
Method: single particle / : Lu JW, Song JK

EMDB-28159: 
Cryo-EM structure of human DNMT3B homo-hexamer
Method: single particle / : Lu JW, Song JK

EMDB-35452: 
Cryo-EM structure of ochratoxin A-detoxifying amidohydrolase ADH3
Method: single particle / : Dai LH, Niu D, Huang JW, Li X, Shen PP, Li H, Hu YM, Yang Y, Chen CC, Guo RT

EMDB-35453: 
Cryo-EM structure of ochratoxin A-detoxifying amidohydrolase ADH3 in complex with Phe
Method: single particle / : Dai LH, Niu D, Huang JW, Li X, Shen PP, Li H, Hu YM, Yang Y, Chen CC, Guo RT

EMDB-35454: 
Cryo-EM structure of ochratoxin A-detoxifying amidohydrolase ADH3 in complex with ochratoxin A
Method: single particle / : Dai LH, Niu D, Huang JW, Li X, Shen PP, Li H, Hu YM, Yang Y, Chen CC, Guo RT

EMDB-36060: 
Cryo-EM structure of ochratoxin A-detoxifying amidohydrolase ADH3 mutant S88E in complex with ochratoxin A
Method: single particle / : Dai LH, Niu D, Huang JW, Li X, Shen PP, Li H, Hu YM, Yang Y, Chen CC, Guo RT

EMDB-15522: 
RCII/PSI complex, class 3
Method: single particle / : Zhao Z, Vercellino I, Knoppova J, Sobotka R, Murray JW, Nixon PJ, Sazanov LA, Komenda J

EMDB-15618: 
RCII/PSI complex, class 2
Method: single particle / : Zhao Z, Vercellino I, Knoppova J, Sobotka R, Murray JW, Nixon PJ, Sazanov LA, Komenda J

EMDB-15621: 
RCII/PSI complex, focused refinement of PSI
Method: single particle / : Zhao Z, Vercellino I, Knoppova J, Sobotka R, Murray JW, Nixon PJ, Sazanov LA, Komenda J

EMDB-33734: 
Cryo-EM structure of SARS-CoV-2 spike in complex with K202.B bispecific antibody
Method: single particle / : Yoo Y, Cho HS

EMDB-33448: 
Cryo-EM structure of Listeria monocytogenes man-PTS complexed with pediocin PA-1
Method: single particle / : Zeng JW, Wang JW, Zhu LY

EMDB-40422: 
Cryo-EM Structure of RyR1
Method: single particle / : Cholak S, Saville JW, Zhu X, Berezuk AM, Tuttle KS, Haji-Ghassemi O, Van Petegem F, Subramaniam S

EMDB-40423: 
Cryo-EM Structure of RyR1 + ATP-gamma-S
Method: single particle / : Cholak S, Saville JW, Zhu X, Berezuk AM, Tuttle KS, Haji-Ghassemi O, Van Petegem F, Subramaniam S
Pages:
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search



 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
