-Search query
-Search result
Showing 1 - 50 of 94 items for (author: leiman & p)
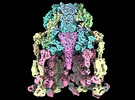
EMDB-29383: 
Structure of baseplate with receptor binding complex of Agrobacterium phage Milano
Method: single particle / : Sonani RR, Leiman PG, Wang F, Kreutzberger MAB, Sebastian A, Esteves NC, Kelly RJ, Scharf B, Egelman EH

PDB-8fqc: 
Structure of baseplate with receptor binding complex of Agrobacterium phage Milano
Method: single particle / : Sonani RR, Leiman PG, Wang F, Kreutzberger MAB, Sebastian A, Esteves NC, Kelly RJ, Scharf B, Egelman EH
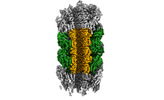
EMDB-29353: 
Structure of Agrobacterium tumefaciens bacteriophage Milano curved tail
Method: single particle / : Sonani RR, Leiman PG, Wang F, Kreutzberger MAB, Sebastian A, Esteves NC, Kelly RJ, Scharf B, Egelman EH

EMDB-29354: 
Structure of Agrobacterium tumefaciens bacteriophage Milano contracted tail-tube
Method: helical / : Sonani RR, Leiman PG, Wang F, Kreutzberger MAB, Sebastian A, Esteves NC, Kelly RJ, Scharf B, Egelman EH

EMDB-29355: 
Structure of Agrobacterium tumefaciens bacteriophage Milano contracted tail-sheath
Method: helical / : Sonani RR, Leiman PG, Wang F, Kreutzberger MAB, Sebastian A, Esteves NC, Kelly RJ, Scharf B, Egelman EH

PDB-8fop: 
Structure of Agrobacterium tumefaciens bacteriophage Milano curved tail
Method: single particle / : Sonani RR, Leiman PG, Wang F, Kreutzberger MAB, Sebastian A, Esteves NC, Kelly RJ, Scharf B, Egelman EH

PDB-8fou: 
Structure of Agrobacterium tumefaciens bacteriophage Milano contracted tail-tube
Method: helical / : Sonani RR, Leiman PG, Wang F, Kreutzberger MAB, Sebastian A, Esteves NC, Kelly RJ, Scharf B, Egelman EH

PDB-8foy: 
Structure of Agrobacterium tumefaciens bacteriophage Milano contracted tail-sheath
Method: helical / : Sonani RR, Leiman PG, Wang F, Kreutzberger MAB, Sebastian A, Esteves NC, Kelly RJ, Scharf B, Egelman EH
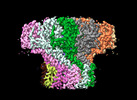
EMDB-29500: 
Portal assembly of Agrobacterium phage Milano
Method: single particle / : Sonani RR, Wang F, Esteves NC, Kelly RJ, Sebastian A, Kreutzberger MAB, Leiman PG, Scharf BE, Egelman EH
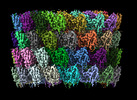
EMDB-29501: 
Collar sheath structure of Agrobacterium phage Milano
Method: single particle / : Sonani RR, Wang F, Esteves NC, Kelly RJ, Sebastian A, Kreutzberger MAB, Leiman PG, Scharf BE, Egelman EH

EMDB-29503: 
Neck structure of Agrobacterium phage Milano, C3 symmetry
Method: single particle / : Sonani RR, Wang F, Esteves NC, Kelly RJ, Sebastian A, Kreutzberger MAB, Leiman PG, Scharf BE, Egelman EH
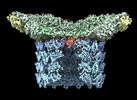
EMDB-29504: 
Structure of neck and portal vertex of Agrobacterium phage Milano, C5 symmetry
Method: single particle / : Sonani RR, Wang F, Esteves NC, Kelly RJ, Sebastian A, Kreutzberger MAB, Leiman PG, Scharf BE, Egelman EH

EMDB-29512: 
Structure of tail-neck junction of Agrobacterium phage Milano
Method: single particle / : Sonani RR, Wang F, Esteves NC, Kelly RJ, Sebastian A, Kreutzberger MAB, Leiman PG, Scharf BE, Egelman EH

EMDB-29540: 
Structure of capsid of Agrobacterium phage Milano
Method: single particle / : Sonani RR, Wang F, Esteves NC, Kelly RJ, Sebastian A, Kreutzberger MAB, Leiman PG, Scharf BE, Egelman EH
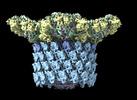
EMDB-29541: 
Structure of neck with portal vertex of capsid of Agrobacterium phage Milano
Method: single particle / : Sonani RR, Wang F, Esteves NC, Kelly RJ, Sebastian A, Kreutzberger MAB, Leiman PG, Scharf BE, Egelman EH

PDB-8fwb: 
Portal assembly of Agrobacterium phage Milano
Method: single particle / : Sonani RR, Wang F, Esteves NC, Kelly RJ, Sebastian A, Kreutzberger MAB, Leiman PG, Scharf BE, Egelman EH

PDB-8fwc: 
Collar sheath structure of Agrobacterium phage Milano
Method: single particle / : Sonani RR, Wang F, Esteves NC, Kelly RJ, Sebastian A, Kreutzberger MAB, Leiman PG, Scharf BE, Egelman EH

PDB-8fwe: 
Neck structure of Agrobacterium phage Milano, C3 symmetry
Method: single particle / : Sonani RR, Wang F, Esteves NC, Kelly RJ, Sebastian A, Kreutzberger MAB, Leiman PG, Scharf BE, Egelman EH

PDB-8fwg: 
Structure of neck and portal vertex of Agrobacterium phage Milano, C5 symmetry
Method: single particle / : Sonani RR, Wang F, Esteves NC, Kelly RJ, Sebastian A, Kreutzberger MAB, Leiman PG, Scharf BE, Egelman EH

PDB-8fwm: 
Structure of tail-neck junction of Agrobacterium phage Milano
Method: single particle / : Sonani RR, Wang F, Esteves NC, Kelly RJ, Sebastian A, Kreutzberger MAB, Leiman PG, Scharf BE, Egelman EH

PDB-8fxp: 
Structure of capsid of Agrobacterium phage Milano
Method: single particle / : Sonani RR, Wang F, Esteves NC, Kelly RJ, Sebastian A, Kreutzberger MAB, Leiman PG, Scharf BE, Egelman EH

PDB-8fxr: 
Structure of neck with portal vertex of capsid of Agrobacterium phage Milano
Method: single particle / : Sonani RR, Wang F, Esteves NC, Kelly RJ, Sebastian A, Kreutzberger MAB, Leiman PG, Scharf BE, Egelman EH

EMDB-25443: 
CryoEM map of the manually-picked stalled contraction intermediate state of bacteriophage A511.
Method: single particle / : Fraser A, Leiman PG

EMDB-25444: 
CryoEM map of the CNN-picked stalled contraction intermediate state of bacteriophage A511.
Method: single particle / : Fraser A, Leiman PG

EMDB-24763: 
Cryo-EM map of the phage AR9 non-virion RNA polymerase holoenzyme in complex with DNA containing the AR9 P077 promoter
Method: single particle / : Fraser A, Leiman PG, Sokolova ML
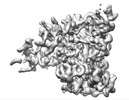
EMDB-24765: 
Cryo-EM map of the phage AR9 non-virion RNA polymerase holoenzyme
Method: single particle / : Fraser A, Leiman PG, Sokolova ML

PDB-7um0: 
Structure of the phage AR9 non-virion RNA polymerase holoenzyme in complex with two DNA oligonucleotides containing the AR9 P077 promoter as determined by cryo-EM
Method: single particle / : Leiman PG, Fraser A, Sokolova ML

PDB-7um1: 
Structure of bacteriophage AR9 non-virion RNAP polymerase holoenzyme determined by cryo-EM
Method: single particle / : Leiman PG, Fraser A, Sokolova ML

EMDB-11659: 
The contracted tail of myophage vb_EcoM_CBA120 (CBA120)
Method: helical / : Nazarov S, Leiman P

EMDB-4906: 
Cryo-EM reconstruction of TssA protein from T6SS of Escherichia coli.
Method: single particle / : Nazarov S, Shneider M, Basler M, Leiman P

PDB-6rju: 
C-terminal domain of TssA protein from T6SS of Escherichia coli.
Method: single particle / : Nazarov S, Shneider M, Basler M, Leiman P

EMDB-20526: 
CryoEM Structure of Pyocin R2 - precontracted - trunk
Method: helical / : Ge P, Avaylon J

EMDB-20643: 
CryoEM Structure of Pyocin R2 - precontracted - baseplate
Method: single particle / : Ge P, Avaylon J

EMDB-20644: 
CryoEM Structure of Pyocin R2 - precontracted - collar
Method: single particle / : Ge P, Avaylon J

EMDB-20646: 
CryoEM Structure of Pyocin R2 - precontracted - hub
Method: single particle / : Ge P, Avaylon J

EMDB-20647: 
CryoEM Structure of Pyocin R2 - postcontracted - collar
Method: single particle / : Ge P, Avaylon J

EMDB-20648: 
CryoEM Structure of Pyocin R2 - postcontracted - baseplate
Method: single particle / : Ge P, Avaylon J

PDB-6pyt: 
CryoEM Structure of Pyocin R2 - precontracted - trunk
Method: helical / : Ge P, Avaylon J, Scholl D, Shneider MM, Browning C, Buth SA, Plattner M, Ding K, Leiman PG, Miller JF, Zhou ZH

PDB-6u5b: 
CryoEM Structure of Pyocin R2 - precontracted - baseplate
Method: single particle / : Ge P, Avaylon J, Scholl D, Shneider MM, Browning C, Buth SA, Plattner M, Ding K, Leiman PG, Miller JF, Zhou ZH

PDB-6u5f: 
CryoEM Structure of Pyocin R2 - precontracted - collar
Method: single particle / : Ge P, Avaylon J, Scholl D, Shneider MM, Browning C, Buth SA, Plattner M, Ding K, Leiman PG, Miller JF, Zhou ZH

PDB-6u5h: 
CryoEM Structure of Pyocin R2 - precontracted - hub
Method: single particle / : Ge P, Avaylon J, Scholl D, Shneider MM, Browning C, Buth SA, Plattner M, Ding K, Leiman PG, Miller JF, Zhou ZH

PDB-6u5j: 
CryoEM Structure of Pyocin R2 - postcontracted - collar
Method: single particle / : Ge P, Avaylon J, Scholl D, Shneider MM, Browning C, Buth SA, Plattner M, Ding K, Leiman PG, Miller JF, Zhou ZH

PDB-6u5k: 
CryoEM Structure of Pyocin R2 - postcontracted - baseplate
Method: single particle / : Ge P, Avaylon J, Scholl D, Shneider MM, Browning C, Buth SA, Plattner M, Ding K, Leiman PG, Miller JF, Zhou ZH

EMDB-7559: 
Bacteriophage A511 baseplate in post-host attachment state (urea-induced)
Method: single particle / : Guerrero-Ferreira R

EMDB-7560: 
Bacteriophage A511 baseplate in pre-host attachment state
Method: single particle / : Guerrero-Ferreira R

EMDB-7561: 
Bacteriophage A511 baseplate in post-host attachment state
Method: single particle / : Guerrero-Ferreira R

PDB-5w5e: 
Re-refinement of the pyocin tube structure
Method: helical / : Wang F, Zheng W, Taylor NM, Guerrero-Ferreira RC, Leiman PG, Egelman EH

PDB-5w5f: 
Cryo-EM structure of the T4 tail tube
Method: helical / : Zheng W, Wang F, Taylor NM, Guerrero-Ferreira RC, Leiman PG, Egelman EH

EMDB-8396: 
Asymmetric reconstruction of bacteriophage MS2 mutant NEO1
Method: single particle / : Guerrero-Ferreira RC, Nazarov SY, Zhong Q, Kohn T, Leiman PG
Pages:
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search




 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
