-Search query
-Search result
Showing 1 - 50 of 139 items for (author: lee & jh)
EMDB-35377: 
Cryo-EM structure of GPR156 of GPR156-miniGo-scFv16 complex (local refine)
Method: single particle / : Shin J, Park J, Cho Y

EMDB-35378: 
Cryo-EM structure of miniGo-scFv16 of GPR156-miniGo-scFv16 complex (local refine)
Method: single particle / : Shin J, Park J, Cho Y

EMDB-35380: 
Cryo-EM structure of GPR156-miniGo-scFv16 complex
Method: single particle / : Shin J, Park J, Cho Y

EMDB-35382: 
Cryo-EM structure of GPR156A/B of G-protein free GPR156 (local refine)
Method: single particle / : Shin J, Park J, Cho Y

EMDB-35389: 
Cryo-EM structure of GPR156C/D of G-protein free GPR156 (local refine)
Method: single particle / : Shin J, Park J, Cho Y

EMDB-35390: 
Cryo-EM structure of G-protein free GPR156
Method: single particle / : Shin J, Park J, Cho Y

EMDB-35234: 
Cryo-EM structure of Acipimox bound human hydroxy-carboxylic acid receptor 2 in complex with Gi heterotrimer
Method: single particle / : Park JH, Ishimoto N, Park SY

EMDB-35235: 
Cryo-EM structure of GSK256073 bound human hydroxy-carboxylic acid receptor 2 in complex with Gi heterotrimer
Method: single particle / : Park JH, Ishimoto N, Park SY

EMDB-36900: 
Cryo-EM structure of niacin bound human hydroxy-carboxylic acid receptor 2 (Local refinement)
Method: single particle / : Park JH, Ishimoto N, Park SY

EMDB-36901: 
Cryo-EM structure of Acipimox bound human hydroxy-carboxylic acid receptor 2 (Local refinement)
Method: single particle / : Park JH, Ishimoto N, Park SY

EMDB-36902: 
Cryo-EM structure of GSK256073 bound human hydroxy-carboxylic acid receptor 2 (Local refinement)
Method: single particle / : Park JH, Ishimoto N, Park SY

EMDB-40088: 
HIV-1 Env subtype C CZA97.12 SOSIP.664 in complex with 3BNC117 Fab
Method: single particle / : Ozorowski G, Lee JH, Ward AB

EMDB-34437: 
Cryo-EM structure of niacin bound human hydroxy-carboxylic acid receptor 2 in complex with Gi heterotrimer
Method: single particle / : Park JH, Ishimoto N, Park SY

EMDB-33734: 
Cryo-EM structure of SARS-CoV-2 spike in complex with K202.B bispecific antibody
Method: single particle / : Yoo Y, Cho HS

EMDB-33569: 
Higher-ordered assembly of mouse TRIM72 WT on the Phosphatidylserine/Cholesterol liposome bilayer
Method: subtomogram averaging / : Park SH, Hyun J, Jeong H, Song HK

EMDB-33582: 
Higher-ordered assembly of mouse TRIM72 M138R on the Phosphatidylserine/Cholesterol liposome bilayer
Method: subtomogram averaging / : Park SH, Hyun J, Jeong H, Song HK

EMDB-27867: 
E. coli 50S ribosome bound to D-linker solithromycin conjugate
Method: single particle / : Pellegrino J, Lee DJ, Fraser JS, Seiple IB

EMDB-27868: 
E. coli 50S ribosome bound to L-linker solithromycin conjugate
Method: single particle / : Pellegrino J, Lee DJ, Fraser JS, Seiple IB

EMDB-27869: 
E. coli 50S ribosome bound to solithromycin and VM1
Method: single particle / : Pellegrino J, Lee DJ, Fraser JS, Seiple IB

EMDB-28650: 
Human S1P transporter Spns2 in an inward-facing open conformation (state 1)
Method: single particle / : Ahmed S, Zhao H, Dai Y, Lee CH

EMDB-28651: 
Human S1P transporter Spns2 in an outward-facing open conformation (state 4)
Method: single particle / : Ahmed S, Zhao H, Dai Y, Lee CH

EMDB-28652: 
Human S1P transporter Spns2 in an inward-facing open conformation (state 1*)
Method: single particle / : Ahmed S, Zhao H, Dai Y, Lee CH

EMDB-28653: 
Human S1P transporter Spns2 in an outward-facing partially occluded conformation (state 3)
Method: single particle / : Ahmed S, Zhao H, Dai Y, Lee CH

EMDB-28654: 
Human S1P transporter Spns2 in an outward-facing partially occluded conformation (state 2)
Method: single particle / : Ahmed S, Zhao H, Dai Y, Lee CH

EMDB-29860: 
Structure of inhibitor 16d-bound SPNS2
Method: single particle / : Chen H, Li X

EMDB-27973: 
Erwinia amylovora 70S Ribosome
Method: subtomogram averaging / : Laughlin TG, Villa E

EMDB-27993: 
Erwinia amlyavora 50S ribosome
Method: subtomogram averaging / : Prichard A, Lee J, Laughlin TG, Lee A, Thomas K, Sy A, Spencer T, Asavavimol A, Cafferata A, Cameron M, Chiu N, Davydov D, Desai I, Diaz G, Guereca M, Hearst K, Huang L, Jacobs E, Johnson A, Kahn S, Koch R, Martinez A, McDonald-Uyeno A, Norquist M, Pau T, Prasad G, Saam K, Sandu M, Sarabia A, Schumaker S, Sonin A, Zhao A, Corbett A, Pogliano K, Meyer J, Grose J, Villa E, Dutton R, Pogliano J

EMDB-28003: 
Erwinia phage vB_EamM_RAY (RAY) Capsid Vertex
Method: subtomogram averaging / : Laughlin TG, Villa E

EMDB-28004: 
Erwinia phage vB_EamM_RAY (RAY) Capsid Collar
Method: subtomogram averaging / : Laughlin TG, Villa E

EMDB-28005: 
Erwinia phage vB_EamM_RAY (RAY) Tail Sheath
Method: subtomogram averaging / : Laughlin TG, Villa E

EMDB-28006: 
Erwinia phage vB_EamM_RAY (RAY) Baseplate
Method: subtomogram averaging / : Laughlin TG, Villa E

EMDB-28007: 
Erwinia phage vB_EamM_RAY (RAY) Chimallin
Method: subtomogram averaging / : Laughlin TG, Villa E

EMDB-28008: 
Erwinia phage vB_EamM_RAY (RAY) Putative PhuZ Filament
Method: subtomogram averaging / : Laughlin TG, Villa E

EMDB-28009: 
Pseudomonas chlororaphis phage 201phi2-1 PhuZ Filament
Method: subtomogram averaging / : Laughlin TG, Villa E

EMDB-28010: 
Pseudomonas phage phiKZ PhuZ Filament
Method: subtomogram averaging / : Laughlin TG, Villa E
EMDB-28523: 
Structure of interleukin receptor common gamma chain (IL2Rgamma) in complex with two antibodies
Method: single particle / : Franklin MC, Romero Hernandez A

EMDB-26639: 
SARS-CoV-2 replication-transcription complex bound to Remdesivir triphosphate, in a pre-catalytic state
Method: single particle / : Malone BF, Perry JK, Appleby TC, Feng JY, Campbell EA, Darst SA

EMDB-26641: 
SARS-CoV-2 replication-transcription complex bound to ATP, in a pre-catalytic state
Method: single particle / : Malone BF, Perry JK, Appleby TC, Feng JY, Campbell EA, Darst SA

EMDB-26642: 
SARS-CoV-2 replication-transcription complex bound to UTP, in a pre-catalytic state
Method: single particle / : Malone BF, Perry JK, Appleby TC, Feng JY, Campbell EA, Darst SA

EMDB-26645: 
SARS-CoV-2 replication-transcription complex bound to GTP, in a pre-catalytic state
Method: single particle / : Malone BF, Perry JK, Appleby TC, Feng JY, Campbell EA, Darst SA

EMDB-26646: 
SARS-CoV-2 replication-transcription complex bound to CTP, in a pre-catalytic state
Method: single particle / : Malone BF, Perry JK, Appleby TC, Feng JY, Campbell EA, Darst SA
EMDB-31152: 
Reconstituted proteoliposomes of TRIM72 in positive curvature #2
Method: electron tomography / : Park SH, Song HK

EMDB-27610: 
BG505 MD39 SOSIP in complex with Rh.NJ82 wk13 N611 and base epitope pAbs
Method: single particle / : Torres JL, Lee WH, Ozorowski G, Ward AB
EMDB-27611: 
BG505 MD39 SOSIP in complex with Rh.NJ95 wk13 base epitope pAb
Method: single particle / : Torres JL, Lee WH, Ozorowski G, Ward AB
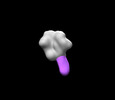
EMDB-27612: 
BG505 MD39 SOSIP in complex with Rh.NK05 wk13 base epitope pAb
Method: single particle / : Torres JL, Lee WH, Ozorowski G, Ward AB
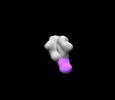
EMDB-27614: 
BG505 MD39 SOSIP in complex with Rh.NJ79 wk13 base epitope pAb
Method: single particle / : Torres JL, Lee WH, Ozorowski G, Ward AB

EMDB-27615: 
BG505 MD39 SOSIP in complex with Rh.NJ93 wk13 base and V5/C3 epitope pAbs
Method: single particle / : Torres JL, Lee WH, Ozorowski G, Ward AB

EMDB-31139: 
Reconstituted proteoliposomes of TRIM72 in negative curvature #1
Method: electron tomography / : Park SH, Song HK
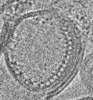
EMDB-31150: 
Reconstituted proteoliposomes of TRIM72 in negative curvature #2
Method: electron tomography / : Park SH, Song HK
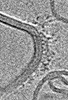
EMDB-31151: 
Reconstituted proteoliposomes of TRIM72 in positive curvature #1
Method: electron tomography / : Park SH, Song HK
Pages:
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search



 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
