[English] 日本語
 Yorodumi
Yorodumi- EMDB-28523: Structure of interleukin receptor common gamma chain (IL2Rgamma) ... -
+ Open data
Open data
- Basic information
Basic information
| Entry | 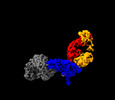 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Structure of interleukin receptor common gamma chain (IL2Rgamma) in complex with two antibodies | |||||||||
 Map data Map data | Structure of interleukin receptor common gamma chain (IL2Rgamma) in complex with two antibodies | |||||||||
 Sample Sample |
| |||||||||
| Function / homology |  Function and homology information Function and homology informationmature B cell differentiation / interleukin-15 receptor activity / interleukin-7-mediated signaling pathway / interleukin-2 binding / CD4-positive, CD25-positive, alpha-beta regulatory T cell differentiation / positive regulation of T cell differentiation in thymus / Interleukin-9 signaling / Interleukin-21 signaling / interleukin-9-mediated signaling pathway / interleukin-4-mediated signaling pathway ...mature B cell differentiation / interleukin-15 receptor activity / interleukin-7-mediated signaling pathway / interleukin-2 binding / CD4-positive, CD25-positive, alpha-beta regulatory T cell differentiation / positive regulation of T cell differentiation in thymus / Interleukin-9 signaling / Interleukin-21 signaling / interleukin-9-mediated signaling pathway / interleukin-4-mediated signaling pathway / positive regulation of CD4-positive, CD25-positive, alpha-beta regulatory T cell differentiation / interleukin-2-mediated signaling pathway / cellular homeostasis / interleukin-15-mediated signaling pathway / STAT3 nuclear events downstream of ALK signaling / Interleukin-15 signaling / Interleukin-2 signaling / cytokine receptor activity / positive regulation of B cell differentiation / cytokine binding / Interleukin receptor SHC signaling / coreceptor activity / positive regulation of phagocytosis / Interleukin-7 signaling / cytokine-mediated signaling pathway / RAF/MAP kinase cascade / gene expression / T cell differentiation in thymus / Interleukin-4 and Interleukin-13 signaling / receptor complex / endosome / immune response / external side of plasma membrane / positive regulation of gene expression / cell surface / signal transduction / endoplasmic reticulum / nucleoplasm / membrane / plasma membrane / cytosol Similarity search - Function | |||||||||
| Biological species |   Homo sapiens (human) Homo sapiens (human) | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 3.4 Å | |||||||||
 Authors Authors | Franklin MC / Romero Hernandez A | |||||||||
| Funding support | 1 items
| |||||||||
 Citation Citation |  Journal: Sci Transl Med / Year: 2023 Journal: Sci Transl Med / Year: 2023Title: Blocking common γ chain cytokine signaling ameliorates T cell-mediated pathogenesis in disease models. Authors: Audrey Le Floc'h / Kirsten Nagashima / Dylan Birchard / George Scott / Li-Hong Ben / Dharani Ajithdoss / Kaitlyn Gayvert / Annabel Romero Hernandez / Olivier Herbin / Amanda Tay / Pamela ...Authors: Audrey Le Floc'h / Kirsten Nagashima / Dylan Birchard / George Scott / Li-Hong Ben / Dharani Ajithdoss / Kaitlyn Gayvert / Annabel Romero Hernandez / Olivier Herbin / Amanda Tay / Pamela Farrales / Chandrashekhar K Korgaonkar / Hao Pan / Sweta Shah / Vishal Kamat / Ishita Chatterjee / Jon Popke / Adelekan Oyejide / Wei Keat Lim / Jee H Kim / Tammy Huang / Matthew Franklin / William Olson / Thomas Norton / Lorah Perlee / George D Yancopoulos / Andrew J Murphy / Matthew A Sleeman / Jamie M Orengo /  Abstract: The common γ chain (γc; IL-2RG) is a subunit of the interleukin (IL) receptors for the γc cytokines IL-2, IL-4, IL-7, IL-9, IL-15, and IL-21. The lack of appropriate neutralizing antibodies ...The common γ chain (γc; IL-2RG) is a subunit of the interleukin (IL) receptors for the γc cytokines IL-2, IL-4, IL-7, IL-9, IL-15, and IL-21. The lack of appropriate neutralizing antibodies recognizing IL-2RG has made it difficult to thoroughly interrogate the role of γc cytokines in inflammatory and autoimmune disease settings. Here, we generated a γc cytokine receptor antibody, REGN7257, to determine whether γc cytokines might be targeted for T cell-mediated disease prevention and treatment. Biochemical, structural, and in vitro analysis showed that REGN7257 binds with high affinity to IL-2RG and potently blocks signaling of all γc cytokines. In nonhuman primates, REGN7257 efficiently suppressed T cells without affecting granulocytes, platelets, or red blood cells. Using REGN7257, we showed that γc cytokines drive T cell-mediated disease in mouse models of graft-versus-host disease (GVHD) and multiple sclerosis by affecting multiple aspects of the pathogenic response. We found that our xenogeneic GVHD mouse model recapitulates hallmarks of acute and chronic GVHD, with T cell expansion/infiltration into tissues and liver fibrosis, as well as hallmarks of immune aplastic anemia, with bone marrow aplasia and peripheral cytopenia. Our findings indicate that γc cytokines contribute to GVHD and aplastic anemia pathology by promoting these characteristic features. By demonstrating that broad inhibition of γc cytokine signaling with REGN7257 protects from immune-mediated disorders, our data provide evidence of γc cytokines as key drivers of pathogenic T cell responses, offering a potential strategy for the management of T cell-mediated diseases. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_28523.map.gz emd_28523.map.gz | 51.5 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-28523-v30.xml emd-28523-v30.xml emd-28523.xml emd-28523.xml | 20 KB 20 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_28523.png emd_28523.png | 35.3 KB | ||
| Others |  emd_28523_half_map_1.map.gz emd_28523_half_map_1.map.gz emd_28523_half_map_2.map.gz emd_28523_half_map_2.map.gz | 95.6 MB 95.6 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-28523 http://ftp.pdbj.org/pub/emdb/structures/EMD-28523 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-28523 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-28523 | HTTPS FTP |
-Validation report
| Summary document |  emd_28523_validation.pdf.gz emd_28523_validation.pdf.gz | 928.2 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_28523_full_validation.pdf.gz emd_28523_full_validation.pdf.gz | 927.7 KB | Display | |
| Data in XML |  emd_28523_validation.xml.gz emd_28523_validation.xml.gz | 13.3 KB | Display | |
| Data in CIF |  emd_28523_validation.cif.gz emd_28523_validation.cif.gz | 15.6 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-28523 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-28523 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-28523 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-28523 | HTTPS FTP |
-Related structure data
| Related structure data |  8epaMC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_28523.map.gz / Format: CCP4 / Size: 103 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_28523.map.gz / Format: CCP4 / Size: 103 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Structure of interleukin receptor common gamma chain (IL2Rgamma) in complex with two antibodies | ||||||||||||||||||||
| Voxel size | X=Y=Z: 0.86 Å | ||||||||||||||||||||
| Density |
| ||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Half map: Half Map 1
| File | emd_28523_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Half Map 1 | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: Half Map 2
| File | emd_28523_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Half Map 2 | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Complex of IL2Rgamma extracellular domain with Fab fragments of a...
| Entire | Name: Complex of IL2Rgamma extracellular domain with Fab fragments of antibodies REGN7257 and REGN9432 |
|---|---|
| Components |
|
-Supramolecule #1: Complex of IL2Rgamma extracellular domain with Fab fragments of a...
| Supramolecule | Name: Complex of IL2Rgamma extracellular domain with Fab fragments of antibodies REGN7257 and REGN9432 type: complex / ID: 1 / Chimera: Yes / Parent: 0 / Macromolecule list: all |
|---|---|
| Source (natural) | Organism:  |
-Macromolecule #1: Cytokine receptor common subunit gamma
| Macromolecule | Name: Cytokine receptor common subunit gamma / type: protein_or_peptide / ID: 1 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 31.617098 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: LNTTILTPNG NEDTTADFFL TTMPTDSLSV STLPLPEVQC FVFNVEYMNC TWNSSSEPQP TNLTLHYWYK NSDNDKVQKC SHYLFSEEI TSGCQLQKKE IHLYQTFVVQ LQDPREPRRQ ATQMLKLQNL VIPWAPENLT LHKLSESQLE LNWNNRFLNH C LEHLVQYR ...String: LNTTILTPNG NEDTTADFFL TTMPTDSLSV STLPLPEVQC FVFNVEYMNC TWNSSSEPQP TNLTLHYWYK NSDNDKVQKC SHYLFSEEI TSGCQLQKKE IHLYQTFVVQ LQDPREPRRQ ATQMLKLQNL VIPWAPENLT LHKLSESQLE LNWNNRFLNH C LEHLVQYR TDWDHSWTEQ SVDYRHKFSL PSVDGQKRYT FRVRSRFNPL CGSAQHWSEW SHPIHWGSNT SKENPFLFAL EA EQKLISE EDLGGEQKLI SEEDLHHHHH H |
-Macromolecule #2: REGN7257 Fab heavy chain
| Macromolecule | Name: REGN7257 Fab heavy chain / type: protein_or_peptide / ID: 2 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 23.607463 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: EVQLVESGGG LVQPGGSLRL SCAASGFIFS SYEMHWVRQA PGKGLEWISY ISSSGTTIYY ADSVKGRFTI SRDNAKNSLY LHMNSLRAE DTAVYYCTRA RITGTFDVFD IWGQGTMVTV SSASTKGPSV FPLAPCSRST SESTAALGCL VKDYFPEPVT V SWNSGALT ...String: EVQLVESGGG LVQPGGSLRL SCAASGFIFS SYEMHWVRQA PGKGLEWISY ISSSGTTIYY ADSVKGRFTI SRDNAKNSLY LHMNSLRAE DTAVYYCTRA RITGTFDVFD IWGQGTMVTV SSASTKGPSV FPLAPCSRST SESTAALGCL VKDYFPEPVT V SWNSGALT SGVHTFPAVL QSSGLYSLSS VVTVPSSSLG TKTYTCNVDH KPSNTKVDKR VE |
-Macromolecule #3: REGN7257 Fab light chain
| Macromolecule | Name: REGN7257 Fab light chain / type: protein_or_peptide / ID: 3 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 23.541125 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: DIQMTQSPSS LSASVGDRVT ITCRASQSIS SYLNWYQQKP GKAPKLLIFA ASNLQSGVPS RFSGSRSGTD FTLTISSLQP EDFATYYCQ QNYNIPYTFG QGTKLEIKRT VAAPSVFIFP PSDEQLKSGT ASVVCLLNNF YPREAKVQWK VDNALQSGNS Q ESVTEQDS ...String: DIQMTQSPSS LSASVGDRVT ITCRASQSIS SYLNWYQQKP GKAPKLLIFA ASNLQSGVPS RFSGSRSGTD FTLTISSLQP EDFATYYCQ QNYNIPYTFG QGTKLEIKRT VAAPSVFIFP PSDEQLKSGT ASVVCLLNNF YPREAKVQWK VDNALQSGNS Q ESVTEQDS KDSTYSLSST LTLSKADYEK HKVYACEVTH QGLSSPVTKS FNRGEC |
-Macromolecule #4: REGN9432 Fab heavy chain
| Macromolecule | Name: REGN9432 Fab heavy chain / type: protein_or_peptide / ID: 4 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 23.323006 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: EVQLVESGGG VVRPGGSLRL SCAASGFTFD DFDMSWVRQG PGKGLEWVSG INWHGSSTGY ADSVKGRFTI SRDNAKNSLY LQMSSLRAE DTALYHCVRG GTIVGATTPL DYWGQGTLVT VSSASTKGPS VFPLAPCSRS TSESTAALGC LVKDYFPEPV T VSWNSGAL ...String: EVQLVESGGG VVRPGGSLRL SCAASGFTFD DFDMSWVRQG PGKGLEWVSG INWHGSSTGY ADSVKGRFTI SRDNAKNSLY LQMSSLRAE DTALYHCVRG GTIVGATTPL DYWGQGTLVT VSSASTKGPS VFPLAPCSRS TSESTAALGC LVKDYFPEPV T VSWNSGAL TSGVHTFPAV LQSSGLYSLS SVVTVPSSSL GTKTYTCNVD HKPSNTKVDK RVE |
-Macromolecule #5: REGN9432 Fab light chain
| Macromolecule | Name: REGN9432 Fab light chain / type: protein_or_peptide / ID: 5 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 23.247803 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: DIQMTQSPSS LSASVGDRVT MTCRASRTIS SYLSWYQQKS GKVPNLLIFG ASSLQSGVPS RFSASGSGTD FTLIISSLQP EDFATYYCQ QSYSSPLTFG GGTKVEIKRT VAAPSVFIFP PSDEQLKSGT ASVVCLLNNF YPREAKVQWK VDNALQSGNS Q ESVTEQDS ...String: DIQMTQSPSS LSASVGDRVT MTCRASRTIS SYLSWYQQKS GKVPNLLIFG ASSLQSGVPS RFSASGSGTD FTLIISSLQP EDFATYYCQ QSYSSPLTFG GGTKVEIKRT VAAPSVFIFP PSDEQLKSGT ASVVCLLNNF YPREAKVQWK VDNALQSGNS Q ESVTEQDS KDSTYSLSST LTLSKADYEK HKVYACEVTH QGLSSPVTKS FNRGEC |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.5 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 BIOQUANTUM (6k x 4k) / Average electron dose: 40.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 2.4 µm / Nominal defocus min: 1.4000000000000001 µm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
- Image processing
Image processing
| Startup model | Type of model: NONE / Details: Ab initio model generation in CryoSparc |
|---|---|
| Final reconstruction | Applied symmetry - Point group: C1 (asymmetric) / Resolution.type: BY AUTHOR / Resolution: 3.4 Å / Resolution method: FSC 0.143 CUT-OFF / Number images used: 435563 |
| Initial angle assignment | Type: MAXIMUM LIKELIHOOD |
| Final angle assignment | Type: MAXIMUM LIKELIHOOD |
 Movie
Movie Controller
Controller






 Z
Z Y
Y X
X

















