-Search query
-Search result
Showing 1 - 50 of 61 items for (author: courbet & a)

EMDB-40602: 
cryoEM map for design HE0537, a D4 symmetric homo-oligomer designed with RFdiffusion.
Method: single particle / : Courbet A, Baker D, Watson J, Juergens D, Eisenach E

EMDB-29939: 
EM map of de novo designed protein nanopore RNR_C6_3 using a reinforcement learning approach
Method: single particle / : Courbet A, Baker D

EMDB-41907: 
Computationally Designed, Expandable O4 Octahedral Handshake Nanocage
Method: single particle / : Weidle C, Borst A
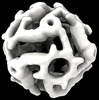
EMDB-42031: 
Computational Designed Nanocage O43_129_+8
Method: single particle / : Weidle C, Kibler RD

EMDB-43318: 
Twistless helix 12 repeat ring design R12B
Method: single particle / : Calise SJ, Kollman JM

EMDB-29974: 
Cryo-EM structure of synthetic tetrameric building block sC4
Method: single particle / : Redler RL, Huddy TF, Hsia Y, Baker D, Ekiert D, Bhabha G

EMDB-41364: 
CryoEM Structure of a Computationally Designed T3 Tetrahedral Nanocage
Method: single particle / : Weidle C, Borst AJ

EMDB-42906: 
Computational Designed Nanocage O43_129
Method: single particle / : Weidle C, Kibler RD

EMDB-42944: 
Computational Designed Nanocage O43_129_+4
Method: single particle / : Carr KD, Weidle C, Borst AJ

PDB-8gel: 
Cryo-EM structure of synthetic tetrameric building block sC4
Method: single particle / : Redler RL, Huddy TF, Hsia Y, Baker D, Ekiert D, Bhabha G

PDB-8tl7: 
CryoEM Structure of a Computationally Designed T3 Tetrahedral Nanocage
Method: single particle / : Weidle C, Borst AJ

PDB-8v3b: 
Computational Designed Nanocage O43_129_+4
Method: single particle / : Carr KD, Weidle C, Borst AJ

EMDB-29893: 
Computationally designed strut_C6_21 de novo protein assembly
Method: single particle / : Courbet A, Huddy TF, Hsia Y

EMDB-27031: 
Accurate computational design of genetically encoded 3D protein crystals
Method: single particle / : Li Z, Borst AJ, Baker D

EMDB-40926: 
CryoEM Structure of Computationally Designed Nanocage O32-ZL4
Method: single particle / : Weidle C, Borst A

PDB-8cwy: 
Accurate computational design of genetically encoded 3D protein crystals
Method: single particle / : Li Z, Borst AJ, Baker D

PDB-8szz: 
CryoEM Structure of Computationally Designed Nanocage O32-ZL4
Method: single particle / : Weidle C, Borst A

EMDB-29915: 
CryoEM map of a de novo designed octahedral nanocage with programmable volume; design cage_O4_34
Method: single particle / : Kibler RD, Borst AJ

EMDB-40070: 
Cryo-EM map of synthetic cage_O3_10 reconstructed without symmetry (C1)
Method: single particle / : Coudray N, Redler R, Hsia Y, Huddy TF, Baker D, Ekiert D, Bhabha G

EMDB-40071: 
Cryo-EM map of synthetic cage_O3_10 reconstructed with O symmetry
Method: single particle / : Coudray N, Redler R, Hsia Y, Huddy TF, Baker D, Ekiert D, Bhabha G

EMDB-40073: 
Cryo-EM map of synthetic cage_T3_5 reconstructed without symmetry (C1), with 1 monomer missing (class 3.0)
Method: single particle / : Coudray N, Redler R, Huddy TF, Hsia Y, Baker D, Ekiert D, Bhabha G

EMDB-40074: 
Cryo-EM map of synthetic cage_T3_5 reconstructed with T symmetry
Method: single particle / : Coudray N, Redler R, Huddy TF, Hsia Y, Baker D, Ekiert D, Bhabha G

EMDB-40075: 
Cryo-EM map of synthetic cage_T3_5 reconstructed without symmetry (C1)
Method: single particle / : Coudray N, Redler R, Huddy TF, Hsia Y, Baker D, Ekiert D, Bhabha G

EMDB-40076: 
Cryo-EM map of synthetic cage_T3_5+2 reconstructed without symmetry (C1)
Method: single particle / : Coudray N, Redler R, Huddy TF, Hsia Y, Baker D, Ekiert D, Bhabha G

EMDB-40072: 
Cryo-EM map of synthetic cage_T3_5 reconstructed without symmetry (C1), with 1 trimer missing (class 3.1)
Method: single particle / : Coudray N, Redler R, Huddy TF, Hsia Y, Baker D, Ekiert D, Bhabha G
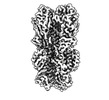
EMDB-40557: 
Cryo-EM structure of designed Influenza HA binder, HA_20, bound to Influenza HA (Strain: Iowa43)
Method: single particle / : Borst AJ, Bennett NR

PDB-8sk7: 
Cryo-EM structure of designed Influenza HA binder, HA_20, bound to Influenza HA (Strain: Iowa43)
Method: single particle / : Borst AJ, Bennett NR

EMDB-28858: 
Top-down design of protein architectures with reinforcement learning
Method: single particle / : Borst AJ, Baker D

EMDB-28859: 
Top-down design of protein architectures with reinforcement learning
Method: single particle / : Borst AJ, Baker D

EMDB-28860: 
Top-down design of protein architectures with reinforcement learning
Method: single particle / : Borst AJ, Baker D

PDB-8f4x: 
Top-down design of protein architectures with reinforcement learning
Method: single particle / : Borst AJ, Baker D

PDB-8f53: 
Top-down design of protein architectures with reinforcement learning
Method: single particle / : Borst AJ, Baker D

PDB-8f54: 
Top-down design of protein architectures with reinforcement learning
Method: single particle / : Borst AJ, Baker D

EMDB-27659: 
cryoEM map of de novo hallucinated homooligomer of C18-C6 symmetry, design HALC18-6_265.
Method: single particle / : Wicky BIM, Milles LF, Courbet AC, Baker D

EMDB-27658: 
cryoEM map of de novo hallucinated homooligomer of C15-C5 symmetry, design HALC15-5_262.
Method: single particle / : Wicky BIM, Milles LF, Courbet AC, Baker D

EMDB-27660: 
cryoEM map of de novo hallucinated homooligomer of C33-C3 symmetry, design HALC33-3_343
Method: single particle / : Wicky BIM, Milles LF, Courbet A, Baker D

EMDB-25575: 
D8-C4 computationally-designed Rotor
Method: single particle / : Hansen JM, Courbet A, Quispe J, Kollman JM, Baker D

EMDB-25576: 
D3-C5 computationally-designed rotor
Method: single particle / : Hansen JM, Courbet A, Quispe J, Kollman JM, Baker D

EMDB-25578: 
D3-C5 computationally-designed rotor (axel only)
Method: single particle / : Hansen JM, Courbet A, Quispe J, Kollman JM, Baker D

EMDB-25579: 
D3-C3 computationally-designed rotor
Method: single particle / : Hansen JM, Courbet A, Quispe J, Kollman JM, Baker D

EMDB-25580: 
C3-C3 computationally-designed rotor
Method: single particle / : Hansen JM, Courbet A, Quispe J, Kollman JM, Baker D

EMDB-23199: 
Generation of ordered protein assemblies using rigid three-body fusion
Method: single particle / : Yao Q, Vulovic I, Baker D, Jensen G

EMDB-23531: 
D3-19.19
Method: single particle / : Park YJ, Ivan V, Baker D, Veesler D

EMDB-23532: 
D3-19.14
Method: single particle / : Park YJ, Ivan V, Baker D, Veesler D

EMDB-23533: 
D3-1.5C
Method: single particle / : Park YJ, Ivan V, Baker D, Veesler D

EMDB-23534: 
Designed oligomer D2-1.1B
Method: single particle / : Park YJ, Ivan V, Baker D, Veesler D

EMDB-23535: 
Designed oligomer D2-1.4H
Method: single particle / : Park YJ, Ivan V, Baker D, Veesler D

EMDB-23536: 
Designed oligomer D2-1.1D
Method: single particle / : Park YJ, Ivan V, Baker D, Veesler D

EMDB-23537: 
DARPin 21.8.HSA-C9.v2 with HSA complex
Method: single particle / : Park YJ, Ivan V, Baker D, Veesler D
Pages:
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search




 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
