-Search query
-Search result
Showing 1 - 50 of 367 items for (author: an & jy)

EMDB-36212: 
PhK holoenzyme in inactive state, muscle isoform
Method: single particle / : Yang XK, Xiao JY

EMDB-36213: 
PhK holoenzyme in active state, muscle isoform
Method: single particle / : Yang XK, Xiao JY

EMDB-36214: 
local map of hPhK alpha-beta-gamma-delta subcomplex in inactive state
Method: single particle / : Yang XK, Xiao JY

EMDB-36215: 
local map of hPhK gamma-delta subcomplex in inactive state
Method: single particle / : Yang XK, Xiao JY

EMDB-36216: 
local map of hPhK alpha-gamma subcomplex in active state
Method: single particle / : Yang XK, Xiao JY

EMDB-41907: 
Computationally Designed, Expandable O4 Octahedral Handshake Nanocage
Method: single particle / : Weidle C, Borst A
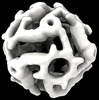
EMDB-42031: 
Computational Designed Nanocage O43_129_+8
Method: single particle / : Weidle C, Kibler RD

EMDB-43318: 
Twistless helix 12 repeat ring design R12B
Method: single particle / : Calise SJ, Kollman JM

EMDB-29974: 
Cryo-EM structure of synthetic tetrameric building block sC4
Method: single particle / : Redler RL, Huddy TF, Hsia Y, Baker D, Ekiert D, Bhabha G

EMDB-41364: 
CryoEM Structure of a Computationally Designed T3 Tetrahedral Nanocage
Method: single particle / : Weidle C, Borst AJ

EMDB-42906: 
Computational Designed Nanocage O43_129
Method: single particle / : Weidle C, Kibler RD

EMDB-42944: 
Computational Designed Nanocage O43_129_+4
Method: single particle / : Carr KD, Weidle C, Borst AJ

EMDB-41442: 
Acinetobacter GP16 Type IV pilus
Method: helical / : Meng R, Xing Z, Zhang J

EMDB-41443: 
Acinetobacter phage AP205
Method: single particle / : Meng R, Xing Z, Chang J, Zhang J

EMDB-41447: 
AP205 binding to one Acinetobacter GP16 type iv pilus
Method: single particle / : Xing Z, Meng R, Zhang J

EMDB-41634: 
Inner Mat-T4P complex
Method: single particle / : Meng R, Xing Z, Thongchol J, Zhang J

EMDB-41635: 
Outer Mat-T4P complex
Method: single particle / : Meng R, Xing Z, Thongchol J, Zhang J

EMDB-41646: 
AP205 phage Acinetobacter gp16 T4P complex
Method: single particle / : Meng R, Xing Z, Zhang J

EMDB-41657: 
Acinetobacter phage AP205 T=4 VLP
Method: single particle / : Meng R, Xing Z, Zhang J

EMDB-41666: 
Acinetobacter phage AP205 T=3 VLP
Method: single particle / : Meng R, Xing Z, Zhang J

EMDB-19431: 
Cryo-em structure of the rat Multidrug resistance-associated protein 2 (rMrp2) in an autoinhibited state (nucleotide-free)
Method: single particle / : Mazza T, Beis K

EMDB-19433: 
Cryo-em structure of the rat Multidrug resistance-associated protein 2 (rMrp2) in complex with probenecid
Method: single particle / : Mazza T, Beis K

PDB-8rq3: 
Cryo-em structure of the rat Multidrug resistance-associated protein 2 (rMrp2) in an autoinhibited state (nucleotide-free)
Method: single particle / : Mazza T, Beis K

PDB-8rq4: 
Cryo-em structure of the rat Multidrug resistance-associated protein 2 (rMrp2) in complex with probenecid
Method: single particle / : Mazza T, Beis K
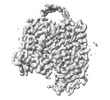
EMDB-35377: 
Cryo-EM structure of GPR156 of GPR156-miniGo-scFv16 complex (local refine)
Method: single particle / : Shin J, Park J, Cho Y

EMDB-35378: 
Cryo-EM structure of miniGo-scFv16 of GPR156-miniGo-scFv16 complex (local refine)
Method: single particle / : Shin J, Park J, Cho Y

EMDB-35380: 
Cryo-EM structure of GPR156-miniGo-scFv16 complex
Method: single particle / : Shin J, Park J, Cho Y

EMDB-35382: 
Cryo-EM structure of GPR156A/B of G-protein free GPR156 (local refine)
Method: single particle / : Shin J, Park J, Cho Y

EMDB-35389: 
Cryo-EM structure of GPR156C/D of G-protein free GPR156 (local refine)
Method: single particle / : Shin J, Park J, Cho Y

EMDB-35390: 
Cryo-EM structure of G-protein free GPR156
Method: single particle / : Shin J, Park J, Cho Y

EMDB-36951: 
Cannabinoid Receptor 1 bound to Fenofibrate coupling MiniGsq and Nb35 Complex
Method: single particle / : Tang WQ, Wang TX, Li FH, Wang JY

EMDB-36952: 
Cannabinoid Receptor 1 bound to Fenofibrate coupling MiniGsq and Nb35 Complex-Global Refine
Method: single particle / : Tang WQ, Wang TX, Li FH, Wang JY

EMDB-36953: 
Cannabinoid Receptor 1 bound to Fenofibrate coupling MiniGsq and Nb35 Complex-Local Refine
Method: single particle / : Tang WQ, Wang TX, Li FH, Wang JY

EMDB-35758: 
Cryo-EM structure of mouse BIRC6, N-terminal section optimized
Method: single particle / : Liu S, Jiang T, Bu F, Zhao J, Wang G, Li N, Gao N, Qiu X

EMDB-35759: 
Cryo-EM structure of mouse BIRC6, Global map
Method: single particle / : Liu S, Jiang T, Bu F, Zhao J, Wang G, Li N, Gao N, Qiu X
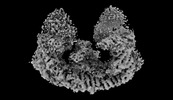
EMDB-35760: 
Cryo-EM structure of mouse BIRC6, with endogenous Smac binding
Method: single particle / : Liu S, Jiang T, Bu F, Zhao J, Wang G, Li N, Gao N, Qiu X

EMDB-38461: 
Cryo-EM structure of mouse BIRC6, Core region
Method: single particle / : Liu S, Jiang T, Bu F, Zhao J, Wang G, Li N, Gao N, Qiu X

EMDB-38462: 
Cryo-EM structure of mouse BIRC6, Half map of the core region
Method: single particle / : Liu S, Jiang T, Bu F, Zhao J, Wang G, Li N, Gao N, Qiu X

EMDB-38464: 
Cryo-EM structure of mouse BIRC6, Composite map
Method: single particle / : Liu S, Jiang T, Bu F, Zhao J, Wang G, Li N, Gao N, Qiu X
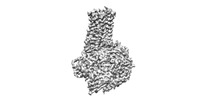
EMDB-37356: 
Cryo-EM structure of the GPR101-Gs complex
Method: single particle / : Sun JP, Gao N, Yu X, Wang GP, Yang F, Wang JY, Yang Z, Guan Y

EMDB-37357: 
Cryo-EM structure of the AA-14-bound GPR101-Gs complex
Method: single particle / : Sun JP, Yu X, Gao N, Yang F, Wang JY, Yang Z, Guan Y, Wang GP

EMDB-37358: 
Cryo-EM structure of the AA14-bound GPR101 complex
Method: single particle / : Sun JP, Yu X, Gao N, Yang F, Wang JY, Yang Z, Guan Y, Wang GP

EMDB-34908: 
Cryo-EM structure of ligand histamine-bound Histamine H4 receptor Gi complex
Method: single particle / : Tang WQ, Sun XY, Li FH, Wang JY

EMDB-34925: 
Cryo-EM structure of ligand histamine-bound Histamine H4 receptor Gi complex
Method: single particle / : Tang WQ, Sun XY, Li FH, Wang JY

EMDB-36636: 
Cryo-EM structure of SARS-CoV-1 2P spike protein in complex with W328-6A1 IgG
Method: single particle / : Zhu JY

EMDB-36647: 
Cryo-EM structure of SARS-CoV-1 2P spike protein in complex with W328-6E10 Fab
Method: single particle / : Zhu JY

EMDB-36648: 
Cryo-EM structure of SARS-CoV-1 2P spike protein protomer in complex with W328-6E10 (local refine)
Method: single particle / : Zhu JY

EMDB-29013: 
96nm doublet microtubule repeat from wild type mouse sperm
Method: subtomogram averaging / : Hwang JY, Chai P, Nawaz S

EMDB-34550: 
Immune complex of W322-3D10 IgG binding the RBD of SARS-CoV-1 2P spike protein
Method: single particle / : Zhu JY, Zhou BN

EMDB-34553: 
Immune complex of W328-5B1 IgG binding the RBD of SARS-CoV-1 2P spike protein
Method: single particle / : Zhu JY, Zhou BN
Pages:
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search



 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
