[English] 日本語
 Yorodumi
Yorodumi- PDB-6g8l: 14-3-3sigma in complex with a L132beta3L mutated YAP pS127 phosph... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 6g8l | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
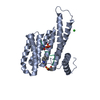
| Title | 14-3-3sigma in complex with a L132beta3L mutated YAP pS127 phosphopeptide | |||||||||
 Components Components |
| |||||||||
 Keywords Keywords | ONCOPROTEIN / beta amino acid / hippo pathway / YAP/TAZ | |||||||||
| Function / homology |  Function and homology information Function and homology informationenterocyte differentiation / regulation of keratinocyte proliferation / regulation of metanephric nephron tubule epithelial cell differentiation / cardiac muscle tissue regeneration / glandular epithelial cell differentiation / TEAD-YAP complex / lateral mesoderm development / RUNX3 regulates YAP1-mediated transcription / bud elongation involved in lung branching / polarized epithelial cell differentiation ...enterocyte differentiation / regulation of keratinocyte proliferation / regulation of metanephric nephron tubule epithelial cell differentiation / cardiac muscle tissue regeneration / glandular epithelial cell differentiation / TEAD-YAP complex / lateral mesoderm development / RUNX3 regulates YAP1-mediated transcription / bud elongation involved in lung branching / polarized epithelial cell differentiation / notochord development / negative regulation of cilium assembly / lung epithelial cell differentiation / YAP1- and WWTR1 (TAZ)-stimulated gene expression / heart process / trophectodermal cell differentiation / paraxial mesoderm development / regulation of stem cell proliferation / tissue homeostasis / hippo signaling / EGR2 and SOX10-mediated initiation of Schwann cell myelination / intestinal epithelial cell development / negative regulation of epithelial cell apoptotic process / negative regulation of stem cell differentiation / Formation of axial mesoderm / embryonic heart tube morphogenesis / female germ cell nucleus / Differentiation of keratinocytes in interfollicular epidermis in mammalian skin / proline-rich region binding / Signaling by Hippo / negative regulation of epithelial cell differentiation / organ growth / interleukin-6-mediated signaling pathway / positive regulation of Notch signaling pathway / negative regulation of fat cell differentiation / positive regulation of stem cell population maintenance / regulation of epidermal cell division / protein kinase C inhibitor activity / positive regulation of epidermal cell differentiation / keratinocyte development / keratinization / RUNX2 regulates osteoblast differentiation / Zygotic genome activation (ZGA) / somatic stem cell population maintenance / Regulation of localization of FOXO transcription factors / keratinocyte proliferation / phosphoserine residue binding / regulation of neurogenesis / Activation of BAD and translocation to mitochondria / canonical Wnt signaling pathway / negative regulation of keratinocyte proliferation / vasculogenesis / establishment of skin barrier / SARS-CoV-2 targets host intracellular signalling and regulatory pathways / positive regulation of osteoblast differentiation / Chk1/Chk2(Cds1) mediated inactivation of Cyclin B:Cdk1 complex / SARS-CoV-1 targets host intracellular signalling and regulatory pathways / negative regulation of stem cell proliferation / cellular response to retinoic acid / extrinsic apoptotic signaling pathway / Nuclear signaling by ERBB4 / positive regulation of epithelial cell proliferation / RHO GTPases activate PKNs / keratinocyte differentiation / positive regulation of cardiac muscle cell proliferation / transcription coregulator activity / epithelial cell proliferation / protein sequestering activity / protein kinase A signaling / negative regulation of innate immune response / protein export from nucleus / TP53 Regulates Transcription of Genes Involved in G2 Cell Cycle Arrest / release of cytochrome c from mitochondria / positive regulation of protein export from nucleus / stem cell proliferation / response to progesterone / Translocation of SLC2A4 (GLUT4) to the plasma membrane / negative regulation of extrinsic apoptotic signaling pathway / TP53 Regulates Metabolic Genes / negative regulation of protein kinase activity / wound healing / cell morphogenesis / cellular response to gamma radiation / positive regulation of protein localization to nucleus / positive regulation of canonical Wnt signaling pathway / transcription corepressor activity / intrinsic apoptotic signaling pathway in response to DNA damage / cell junction / cell-cell junction / protein localization / RUNX1 regulates transcription of genes involved in differentiation of HSCs / positive regulation of cell growth / DNA-binding transcription factor binding / protein-containing complex assembly / transcription coactivator activity / transcription cis-regulatory region binding / regulation of cell cycle / cadherin binding / RNA polymerase II cis-regulatory region sequence-specific DNA binding / negative regulation of gene expression Similarity search - Function | |||||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | |||||||||
| Method |  X-RAY DIFFRACTION / X-RAY DIFFRACTION /  SYNCHROTRON / SYNCHROTRON /  MOLECULAR REPLACEMENT / MOLECULAR REPLACEMENT /  molecular replacement / Resolution: 1.37 Å molecular replacement / Resolution: 1.37 Å | |||||||||
 Authors Authors | Andrei, S.A. / Thijssen, V. / Brunsveld, L. / Ottmann, C. / Milroy, L.G. | |||||||||
| Funding support |  Netherlands, 2items Netherlands, 2items
| |||||||||
 Citation Citation |  Journal: Chem.Commun.(Camb.) / Year: 2019 Journal: Chem.Commun.(Camb.) / Year: 2019Title: A study on the effect of synthetic alpha-to-beta3-amino acid mutations on the binding of phosphopeptides to 14-3-3 proteins. Authors: Andrei, S.A. / Thijssen, V. / Brunsveld, L. / Ottmann, C. / Milroy, L.G. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  6g8l.cif.gz 6g8l.cif.gz | 180 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb6g8l.ent.gz pdb6g8l.ent.gz | 143.9 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  6g8l.json.gz 6g8l.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Summary document |  6g8l_validation.pdf.gz 6g8l_validation.pdf.gz | 428.3 KB | Display |  wwPDB validaton report wwPDB validaton report |
|---|---|---|---|---|
| Full document |  6g8l_full_validation.pdf.gz 6g8l_full_validation.pdf.gz | 429.2 KB | Display | |
| Data in XML |  6g8l_validation.xml.gz 6g8l_validation.xml.gz | 15.9 KB | Display | |
| Data in CIF |  6g8l_validation.cif.gz 6g8l_validation.cif.gz | 25.5 KB | Display | |
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/g8/6g8l https://data.pdbj.org/pub/pdb/validation_reports/g8/6g8l ftp://data.pdbj.org/pub/pdb/validation_reports/g8/6g8l ftp://data.pdbj.org/pub/pdb/validation_reports/g8/6g8l | HTTPS FTP |
-Related structure data
| Related structure data |  6g6xC  6g8iC  6g8jC  6g8kC  6g8pC  6g8qC  3mhrS C: citing same article ( S: Starting model for refinement |
|---|---|
| Similar structure data |
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| 1 | 
| |||||||||
| Unit cell |
| |||||||||
| Components on special symmetry positions |
|
- Components
Components
-Protein / Protein/peptide , 2 types, 2 molecules AP
| #1: Protein | Mass: 26542.914 Da / Num. of mol.: 1 Source method: isolated from a genetically manipulated source Source: (gene. exp.)  Homo sapiens (human) / Gene: SFN, HME1 / Production host: Homo sapiens (human) / Gene: SFN, HME1 / Production host:  |
|---|---|
| #2: Protein/peptide | Mass: 1175.190 Da / Num. of mol.: 1 / Source method: obtained synthetically / Details: BLE is a beta3-leucine unnatural amino acid / Source: (synth.)  Homo sapiens (human) / References: UniProt: P46937*PLUS Homo sapiens (human) / References: UniProt: P46937*PLUS |
-Non-polymers , 4 types, 429 molecules 






| #3: Chemical | ChemComp-CL / |
|---|---|
| #4: Chemical | ChemComp-MG / |
| #5: Chemical | ChemComp-NA / |
| #6: Water | ChemComp-HOH / |
-Experimental details
-Experiment
| Experiment | Method:  X-RAY DIFFRACTION / Number of used crystals: 1 X-RAY DIFFRACTION / Number of used crystals: 1 |
|---|
- Sample preparation
Sample preparation
| Crystal | Density Matthews: 2.6 Å3/Da / Density % sol: 52.76 % |
|---|---|
| Crystal grow | Temperature: 277 K / Method: vapor diffusion, sitting drop Details: 0.095 M HEPES, pH 7.1, 29% PEG 400, 0.19 M CaCl2, 5% glycerol |
-Data collection
| Diffraction | Mean temperature: 100 K | ||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Diffraction source | Source:  SYNCHROTRON / Site: SYNCHROTRON / Site:  PETRA III, DESY PETRA III, DESY  / Beamline: P11 / Wavelength: 1.0332 Å / Beamline: P11 / Wavelength: 1.0332 Å | ||||||||||||||||||||||||
| Detector | Type: DECTRIS PILATUS 6M-F / Detector: PIXEL / Date: Apr 8, 2016 | ||||||||||||||||||||||||
| Radiation | Protocol: SINGLE WAVELENGTH / Monochromatic (M) / Laue (L): M / Scattering type: x-ray | ||||||||||||||||||||||||
| Radiation wavelength | Wavelength: 1.0332 Å / Relative weight: 1 | ||||||||||||||||||||||||
| Reflection | Resolution: 1.37→66.29 Å / Num. obs: 60861 / % possible obs: 99.8 % / Redundancy: 9.9 % / Biso Wilson estimate: 9.6 Å2 / CC1/2: 0.992 / Rmerge(I) obs: 0.133 / Rpim(I) all: 0.043 / Rrim(I) all: 0.14 / Net I/σ(I): 10.8 / Num. measured all: 602191 / Scaling rejects: 1475 | ||||||||||||||||||||||||
| Reflection shell | Diffraction-ID: 1
|
-Phasing
| Phasing | Method:  molecular replacement molecular replacement | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Phasing MR |
|
- Processing
Processing
| Software |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method to determine structure:  MOLECULAR REPLACEMENT MOLECULAR REPLACEMENTStarting model: 3mhr Resolution: 1.37→45.533 Å / SU ML: 0.13 / Cross valid method: THROUGHOUT / σ(F): 1.34 / Phase error: 14.19
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Solvent computation | Shrinkage radii: 0.9 Å / VDW probe radii: 1.11 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Displacement parameters | Biso max: 64.48 Å2 / Biso mean: 14.4806 Å2 / Biso min: 0.85 Å2 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement step | Cycle: final / Resolution: 1.37→45.533 Å
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| LS refinement shell | Refine-ID: X-RAY DIFFRACTION / Rfactor Rfree error: 0 / Total num. of bins used: 11
|
 Movie
Movie Controller
Controller





















 PDBj
PDBj





















