[English] 日本語
 Yorodumi
Yorodumi- PDB-263d: ISOHELICITY AND PHASING IN DRUG-DNA SEQUENCE RECOGNITION: CRYSTAL... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 263d | ||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Title | ISOHELICITY AND PHASING IN DRUG-DNA SEQUENCE RECOGNITION: CRYSTAL STRUCTURE OF A TRIS(BENZIMIDAZOLE)-OLIGONUCLEOTIDE COMPLEX | ||||||||||||||||||
 Components Components | DNA (5'-D(* Keywords KeywordsDNA / B-DNA / DOUBLE HELIX / COMPLEXED WITH DRUG | Function / homology | Chem-TBZ / DNA / DNA (> 10) |  Function and homology information Function and homology informationMethod |  X-RAY DIFFRACTION / X-RAY DIFFRACTION /  MOLECULAR REPLACEMENT / Resolution: 2.2 Å MOLECULAR REPLACEMENT / Resolution: 2.2 Å  Authors AuthorsClark, G.R. / Gray, E.J. / Neidle, S. / Li, Y.-H. / Leupin, W. |  Citation Citation Journal: Biochemistry / Year: 1996 Journal: Biochemistry / Year: 1996Title: Isohelicity and phasing in drug--DNA sequence recognition: crystal structure of a tris(benzimidazole)--oligonucleotide complex. Authors: Clark, G.R. / Gray, E.J. / Neidle, S. / Li, Y.H. / Leupin, W. #1:  Journal: Nucleic Acids Res. / Year: 1994 Journal: Nucleic Acids Res. / Year: 1994Title: Sequence-Dependent Effects in Drug-DNA Interaction: The Crystal Structure of Hoechst 33258 Bound to the d(CGCAAATTTGCG)2 Duplex Authors: Spink, N. / Skelly, J.V. / Neidle, S. #2:  Journal: Eur.J.Biochem. / Year: 1994 Journal: Eur.J.Biochem. / Year: 1994Title: Three-Dimensional Crystal Structure of the A-Tract DNA Dodecamer d(CGCAAATTTGCG) Complexed with the Minor-Groove-Binding Drug Hoechst 33258 Authors: Vega, M.C. / Saez, I.G. / Aymami, J. / Eritja, R. / van der Marel, G.A. / van Boom, J.H. / Rich, A. / Coll, M. History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  263d.cif.gz 263d.cif.gz | 25 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb263d.ent.gz pdb263d.ent.gz | 16.2 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  263d.json.gz 263d.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Summary document |  263d_validation.pdf.gz 263d_validation.pdf.gz | 566.6 KB | Display |  wwPDB validaton report wwPDB validaton report |
|---|---|---|---|---|
| Full document |  263d_full_validation.pdf.gz 263d_full_validation.pdf.gz | 568.9 KB | Display | |
| Data in XML |  263d_validation.xml.gz 263d_validation.xml.gz | 4.4 KB | Display | |
| Data in CIF |  263d_validation.cif.gz 263d_validation.cif.gz | 5.6 KB | Display | |
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/63/263d https://data.pdbj.org/pub/pdb/validation_reports/63/263d ftp://data.pdbj.org/pub/pdb/validation_reports/63/263d ftp://data.pdbj.org/pub/pdb/validation_reports/63/263d | HTTPS FTP |
-Related structure data
| Similar structure data |
|---|
- Links
Links
- Assembly
Assembly
| Deposited unit | 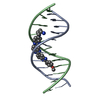
| ||||||||
|---|---|---|---|---|---|---|---|---|---|
| 1 |
| ||||||||
| Unit cell |
|
- Components
Components
| #1: DNA chain | Mass: 3662.404 Da / Num. of mol.: 2 / Source method: obtained synthetically #2: Chemical | ChemComp-TBZ / | #3: Water | ChemComp-HOH / | |
|---|
-Experimental details
-Experiment
| Experiment | Method:  X-RAY DIFFRACTION / Number of used crystals: 1 X-RAY DIFFRACTION / Number of used crystals: 1 |
|---|
- Sample preparation
Sample preparation
| Crystal | Density Matthews: 2.23 Å3/Da / Density % sol: 44.93 % | ||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Crystal grow | Temperature: 286 K / Method: vapor diffusion, hanging drop / pH: 7 Details: pH 7.00, VAPOR DIFFUSION, HANGING DROP, temperature 286.00K | ||||||||||||||||||||||||||||||||||||||||||||||||
| Components of the solutions |
| ||||||||||||||||||||||||||||||||||||||||||||||||
| Crystal grow | *PLUS pH: 7 | ||||||||||||||||||||||||||||||||||||||||||||||||
| Components of the solutions | *PLUS
|
-Data collection
| Diffraction | Mean temperature: 287 K |
|---|---|
| Diffraction source | Source:  ROTATING ANODE / Wavelength: 1.5418 ROTATING ANODE / Wavelength: 1.5418 |
| Detector | Type: XENTRONICS / Detector: AREA DETECTOR / Date: May 15, 1996 |
| Radiation | Monochromator: GRAPHITE / Monochromatic (M) / Laue (L): M / Scattering type: x-ray |
| Radiation wavelength | Wavelength: 1.5418 Å / Relative weight: 1 |
| Reflection | Resolution: 2.2→8 Å / Num. obs: 3594 / % possible obs: 92 % / Observed criterion σ(I): 2 / Rmerge(I) obs: 0.083 |
| Reflection shell | Resolution: 2.2→2.35 Å / % possible all: 57 |
| Reflection | *PLUS Highest resolution: 2.2 Å / Lowest resolution: 8 Å / % possible obs: 92 % / Observed criterion σ(I): 2 |
| Reflection shell | *PLUS Highest resolution: 2.2 Å / Lowest resolution: 2.35 Å / % possible obs: 57 % |
- Processing
Processing
| Software |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method to determine structure:  MOLECULAR REPLACEMENT MOLECULAR REPLACEMENTStarting model: GDL023 Resolution: 2.2→8 Å / σ(F): 2
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Displacement parameters | Biso mean: 24 Å2 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine Biso | Class: ALL ATOMS / Details: TR / Treatment: isotropic | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement step | Cycle: LAST / Resolution: 2.2→8 Å
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine LS restraints |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Software | *PLUS Name:  X-PLOR / Classification: refinement X-PLOR / Classification: refinement | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement | *PLUS Highest resolution: 2.2 Å / Lowest resolution: 8 Å / σ(F): 2 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Solvent computation | *PLUS | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Displacement parameters | *PLUS |
 Movie
Movie Controller
Controller












 PDBj
PDBj