+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 1qwf | ||||||
|---|---|---|---|---|---|---|---|
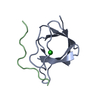
| Title | C-SRC SH3 DOMAIN COMPLEXED WITH LIGAND VSL12 | ||||||
 Components Components |
| ||||||
 Keywords Keywords | COMPLEX (SIGNAL TRANSDUCTION/PEPTIDE) / SRC SH3 DOMAIN / CLASS I LIGAND COMPLEX / COMPLEX (SIGNAL TRANSDUCTION-PEPTIDE) COMPLEX | ||||||
| Function / homology |  Function and homology information Function and homology information non-specific protein-tyrosine kinase / non-membrane spanning protein tyrosine kinase activity / non-specific protein-tyrosine kinase / non-membrane spanning protein tyrosine kinase activity /  phosphorylation / phosphorylation /  ATP binding ATP bindingSimilarity search - Function | ||||||
| Biological species |  Avian sarcoma virus Avian sarcoma virus | ||||||
| Method |  SOLUTION NMR SOLUTION NMR | ||||||
 Authors Authors | Feng, S. / Chiyoshi, K. / Rickles, R.J. / Schreiber, S.L. | ||||||
 Citation Citation |  Journal: Proc.Natl.Acad.Sci.USA / Year: 1995 Journal: Proc.Natl.Acad.Sci.USA / Year: 1995Title: Specific interactions outside the proline-rich core of two classes of Src homology 3 ligands. Authors: Feng, S. / Kasahara, C. / Rickles, R.J. / Schreiber, S.L. #1:  Journal: Cell(Cambridge,Mass.) / Year: 1994 Journal: Cell(Cambridge,Mass.) / Year: 1994Title: Structural Basis for the Binding of Proline-Rich Peptides to SH3 Domains Authors: Yu, H. / Chen, J.K. / Feng, S. / Dalgarno, D.C. / Brauer, A.W. / Schreiber, S.L. #2:  Journal: Science / Year: 1994 Journal: Science / Year: 1994Title: Two Binding Orientations for Peptides to the Src SH3 Domain: Development of a General Model for SH3-Ligand Interactions Authors: Feng, S. / Chen, J.K. / Yu, H. / Simon, J.A. / Schreiber, S.L. #3:  Journal: Science / Year: 1992 Journal: Science / Year: 1992Title: Solution Structure of the SH3 Domain of Src and Identification of its Ligand-Binding Site Authors: Yu, H. / Rosen, M.K. / Shin, T.B. / Seidel-Dugan, C. / Brugge, J.S. / Schreiber, S.L. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  1qwf.cif.gz 1qwf.cif.gz | 30.1 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb1qwf.ent.gz pdb1qwf.ent.gz | 23.2 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  1qwf.json.gz 1qwf.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/qw/1qwf https://data.pdbj.org/pub/pdb/validation_reports/qw/1qwf ftp://data.pdbj.org/pub/pdb/validation_reports/qw/1qwf ftp://data.pdbj.org/pub/pdb/validation_reports/qw/1qwf | HTTPS FTP |
|---|
-Related structure data
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| 1 |
| |||||||||
| NMR ensembles |
|
- Components
Components
| #1: Protein | Mass: 7104.749 Da / Num. of mol.: 1 / Fragment: SH3 DOMAIN Source method: isolated from a genetically manipulated source Source: (gene. exp.)  Avian sarcoma virus / Genus: Alpharetrovirus Avian sarcoma virus / Genus: Alpharetrovirus / Description: GST-FUSION / Gene: CHICKEN / Plasmid: PGEX-2T / Gene (production host): CHICKEN / Production host: / Description: GST-FUSION / Gene: CHICKEN / Plasmid: PGEX-2T / Gene (production host): CHICKEN / Production host:   Escherichia coli (E. coli) / References: UniProt: P00525, EC: 2.7.1.112 Escherichia coli (E. coli) / References: UniProt: P00525, EC: 2.7.1.112 |
|---|---|
| #2: Protein/peptide | Mass: 1317.622 Da / Num. of mol.: 1 Source method: isolated from a genetically manipulated source |
-Experimental details
-Experiment
| Experiment | Method:  SOLUTION NMR SOLUTION NMR |
|---|
- Sample preparation
Sample preparation
Crystal grow | *PLUS Method: other |
|---|
- Processing
Processing
| Software |
| ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| NMR software | Name:  X-PLOR / Version: 3.1 / Developer: BRUNGER / Classification: refinement X-PLOR / Version: 3.1 / Developer: BRUNGER / Classification: refinement | ||||||||||||
| NMR ensemble | Conformers submitted total number: 1 |
 Movie
Movie Controller
Controller








 PDBj
PDBj