[English] 日本語
 Yorodumi
Yorodumi- PDB-1n49: Viability of a Drug-Resistant HIV-1 Protease Variant: Structural ... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 1n49 | ||||||
|---|---|---|---|---|---|---|---|
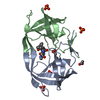
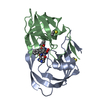
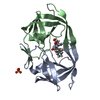
| Title | Viability of a Drug-Resistant HIV-1 Protease Variant: Structural Insights for Better Anti-Viral Therapy | ||||||
 Components Components | Protease | ||||||
 Keywords Keywords | HYDROLASE/HYDROLASE INHIBITOR /  HIV-1 protease / HIV-1 protease /  drug resistance / substrate recognition / drug resistance / substrate recognition /  inhibitor binding / inhibitor binding /  HYDROLASE / HYDROLASE-HYDROLASE INHIBITOR complex HYDROLASE / HYDROLASE-HYDROLASE INHIBITOR complex | ||||||
| Function / homology |  Function and homology information Function and homology information HIV-1 retropepsin / HIV-1 retropepsin /  : / : /  retroviral ribonuclease H / retroviral ribonuclease H /  exoribonuclease H / exoribonuclease H /  : / : /  exoribonuclease H activity / host multivesicular body / DNA integration / exoribonuclease H activity / host multivesicular body / DNA integration /  RNA-directed DNA polymerase / viral genome integration into host DNA ... RNA-directed DNA polymerase / viral genome integration into host DNA ... HIV-1 retropepsin / HIV-1 retropepsin /  : / : /  retroviral ribonuclease H / retroviral ribonuclease H /  exoribonuclease H / exoribonuclease H /  : / : /  exoribonuclease H activity / host multivesicular body / DNA integration / exoribonuclease H activity / host multivesicular body / DNA integration /  RNA-directed DNA polymerase / viral genome integration into host DNA / viral penetration into host nucleus / establishment of integrated proviral latency / RNA-directed DNA polymerase / viral genome integration into host DNA / viral penetration into host nucleus / establishment of integrated proviral latency /  RNA-directed DNA polymerase activity / RNA-directed DNA polymerase activity /  Transferases; Transferring phosphorus-containing groups; Nucleotidyltransferases / RNA-DNA hybrid ribonuclease activity / viral nucleocapsid / DNA recombination / Transferases; Transferring phosphorus-containing groups; Nucleotidyltransferases / RNA-DNA hybrid ribonuclease activity / viral nucleocapsid / DNA recombination /  Hydrolases; Acting on ester bonds / Hydrolases; Acting on ester bonds /  DNA-directed DNA polymerase / aspartic-type endopeptidase activity / DNA-directed DNA polymerase / aspartic-type endopeptidase activity /  DNA-directed DNA polymerase activity / symbiont entry into host cell / symbiont-mediated suppression of host gene expression / DNA-directed DNA polymerase activity / symbiont entry into host cell / symbiont-mediated suppression of host gene expression /  lipid binding / host cell nucleus / host cell plasma membrane / virion membrane / structural molecule activity / lipid binding / host cell nucleus / host cell plasma membrane / virion membrane / structural molecule activity /  proteolysis / proteolysis /  DNA binding / DNA binding /  RNA binding / zinc ion binding / RNA binding / zinc ion binding /  membrane membraneSimilarity search - Function | ||||||
| Biological species |    Human immunodeficiency virus 1 Human immunodeficiency virus 1 | ||||||
| Method |  X-RAY DIFFRACTION / X-RAY DIFFRACTION /  MOLECULAR REPLACEMENT / Resolution: 2.2 Å MOLECULAR REPLACEMENT / Resolution: 2.2 Å | ||||||
 Authors Authors | Prabu-Jeyabalan, M. / Nalivaika, E.A. / King, N.M. / Schiffer, C.A. | ||||||
 Citation Citation |  Journal: J.VIROL. / Year: 2003 Journal: J.VIROL. / Year: 2003Title: Viability of a Drug-Resistant Human Immunodeficiency Virus Type 1 Protease Variant: Structural Insights for Better Antiviral Therapy Authors: Prabu-Jeyabalan, M. / Nalivaika, E.A. / King, N.M. / Schiffer, C.A. #1:  Journal: J.Mol.Biol. / Year: 2000 Journal: J.Mol.Biol. / Year: 2000Title: How does a symmetric dimer recognize an asymmetric substrate? a substrate complex of HIV-1 protease Authors: Prabu-Jeyabalan, M. / Nalivaika, E. / Schiffer, C.A. #2:  Journal: Structure / Year: 2002 Journal: Structure / Year: 2002Title: Substrate shape determines specificity of recognition for HIV-1 protease: analysis of crystal structures of six substrate complexes Authors: Prabu-Jeyabalan, M. / Nalivaika, E. / Schiffer, C.A. #3:  Journal: Protein Sci. / Year: 2002 Journal: Protein Sci. / Year: 2002Title: Lack of synergy for inhibitors targeting a multi-drug resistant HIV-1 protease Authors: King, N.M. / Melnick, L. / Prabu-Jeyabalan, M. / Nalivaika, E.A. / Yang, S.S. / Gao, Y. / Nie, X. / Zepp, C. / Heefner, D.L. / Schiffer, C.A. #4:  Journal: Proc.Natl.Acad.Sci.USA / Year: 1995 Journal: Proc.Natl.Acad.Sci.USA / Year: 1995Title: ABT-538 Is a Potent Inhibitor of Human Immunodeficiency Virus Protease and has High Oral Bioavailability in Humans Authors: Kempf, D.J. / Marsh, K.C. / Denissen, J.F. / McDonald, E. / Vasavanonda, S. / Flentge, C.A. / Green, B.E. / Fino, L. / Park, C.H. / Kong, X. / Wideburg, N.E. / Saldivar, A. / Ruiz, L. / ...Authors: Kempf, D.J. / Marsh, K.C. / Denissen, J.F. / McDonald, E. / Vasavanonda, S. / Flentge, C.A. / Green, B.E. / Fino, L. / Park, C.H. / Kong, X. / Wideburg, N.E. / Saldivar, A. / Ruiz, L. / Kati, W.M. / Sham, H.L. / Robins, T. / Stewart, K.D. / Hsu, A. / Plattner, J.J. / Leonard, J.M. / Norbeck, D.W. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  1n49.cif.gz 1n49.cif.gz | 86.2 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb1n49.ent.gz pdb1n49.ent.gz | 65.4 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  1n49.json.gz 1n49.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/n4/1n49 https://data.pdbj.org/pub/pdb/validation_reports/n4/1n49 ftp://data.pdbj.org/pub/pdb/validation_reports/n4/1n49 ftp://data.pdbj.org/pub/pdb/validation_reports/n4/1n49 | HTTPS FTP |
|---|
-Related structure data
| Related structure data |  1mt7C  1mt8C  1mt9C  1mtbC  1f7aS S: Starting model for refinement C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| ||||||||
|---|---|---|---|---|---|---|---|---|---|
| 1 | 
| ||||||||
| 2 | 
| ||||||||
| Unit cell |
|
- Components
Components
| #1: Protein |  / Retropepsin / RetropepsinMass: 10772.724 Da / Num. of mol.: 4 / Mutation: Q7K, D25N, V82A Source method: isolated from a genetically manipulated source Source: (gene. exp.)    Human immunodeficiency virus 1 / Genus: Lentivirus Human immunodeficiency virus 1 / Genus: Lentivirus / Gene: POL / Plasmid: pEN18 / Production host: / Gene: POL / Plasmid: pEN18 / Production host:   Escherichia coli (E. coli) / Strain (production host): TAP106 / References: UniProt: P03369, Escherichia coli (E. coli) / Strain (production host): TAP106 / References: UniProt: P03369,  HIV-1 retropepsin HIV-1 retropepsin#2: Chemical |  / A-84538 / / A-84538 /  Ritonavir Ritonavir#3: Water | ChemComp-HOH / |  Water Water |
|---|
-Experimental details
-Experiment
| Experiment | Method:  X-RAY DIFFRACTION / Number of used crystals: 1 X-RAY DIFFRACTION / Number of used crystals: 1 |
|---|
- Sample preparation
Sample preparation
| Crystal | Density Matthews: 2.14 Å3/Da / Density % sol: 42.59 % | ||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
Crystal grow | Method: vapor diffusion, hanging drop / pH: 6.2 Details: sodium phosphate, sodium citrate, ammonium sulphate, pH 6.2, VAPOR DIFFUSION, HANGING DROP | ||||||||||||||||||||||||||||||
| Crystal grow | *PLUS | ||||||||||||||||||||||||||||||
| Components of the solutions | *PLUS
|
-Data collection
| Diffraction | Mean temperature: 200 K |
|---|---|
| Diffraction source | Source:  ROTATING ANODE / Type: RIGAKU / Wavelength: 1.5418 Å ROTATING ANODE / Type: RIGAKU / Wavelength: 1.5418 Å |
| Detector | Type: RIGAKU RAXIS IV / Detector: IMAGE PLATE / Date: Aug 7, 2001 / Details: Yale Mirrors |
| Radiation | Monochromator: Yale Mirrors / Protocol: SINGLE WAVELENGTH / Monochromatic (M) / Laue (L): M / Scattering type: x-ray |
| Radiation wavelength | Wavelength : 1.5418 Å / Relative weight: 1 : 1.5418 Å / Relative weight: 1 |
| Reflection | Resolution: 2.2→50 Å / Num. all: 13097 / Num. obs: 13097 / % possible obs: 71.2 % / Observed criterion σ(F): 0 / Observed criterion σ(I): 0 / Rmerge(I) obs: 0.064 / Net I/σ(I): 7.6 |
| Reflection shell | Resolution: 2.2→2.28 Å / Rmerge(I) obs: 0.27 |
| Reflection | *PLUS Lowest resolution: 50 Å / Num. measured all: 19193 |
| Reflection shell | *PLUS % possible obs: 66 % / Num. unique obs: 1184 |
- Processing
Processing
| Software |
| |||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method to determine structure : :  MOLECULAR REPLACEMENT MOLECULAR REPLACEMENTStarting model: 1F7A Resolution: 2.2→50 Å / Isotropic thermal model: restrained / Cross valid method: THROUGHOUT / σ(F): 0 / σ(I): 0 / Stereochemistry target values: Engh & Huber Details: NCS RESTRAINTS IMPOSED BETWEEN THE DIMERS (BUT NOT WITHIN THE DIMERS) TO IMPROVE OBSERVABLES TO PARAMETER RATIO.
| |||||||||||||||||||||||||
| Refinement step | Cycle: LAST / Resolution: 2.2→50 Å
| |||||||||||||||||||||||||
| Refine LS restraints |
| |||||||||||||||||||||||||
| LS refinement shell | Resolution: 2.2→2.28 Å
| |||||||||||||||||||||||||
| Refinement | *PLUS Lowest resolution: 50 Å | |||||||||||||||||||||||||
| Solvent computation | *PLUS | |||||||||||||||||||||||||
| Displacement parameters | *PLUS | |||||||||||||||||||||||||
| Refine LS restraints | *PLUS
|
 Movie
Movie Controller
Controller
















 PDBj
PDBj