+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 1mbj | ||||||
|---|---|---|---|---|---|---|---|
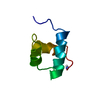
| Title | MOUSE C-MYB DNA-BINDING DOMAIN REPEAT 3 | ||||||
 Components Components | MYB PROTO-ONCOGENE PROTEIN | ||||||
 Keywords Keywords | DNA BINDING PROTEIN / PROTOONCOGENE PRODUCT | ||||||
| Function / homology |  Function and homology information Function and homology informationpositive regulation of testosterone secretion / myeloid cell development / positive regulation of hepatic stellate cell proliferation / positive regulation of hepatic stellate cell activation / positive regulation of transforming growth factor beta production / embryonic digestive tract development / stem cell division / myeloid cell differentiation / cellular response to interleukin-6 / WD40-repeat domain binding ...positive regulation of testosterone secretion / myeloid cell development / positive regulation of hepatic stellate cell proliferation / positive regulation of hepatic stellate cell activation / positive regulation of transforming growth factor beta production / embryonic digestive tract development / stem cell division / myeloid cell differentiation / cellular response to interleukin-6 / WD40-repeat domain binding / positive regulation of glial cell proliferation / homeostasis of number of cells / positive regulation of collagen biosynthetic process / spleen development / B cell differentiation / positive regulation of smooth muscle cell proliferation / thymus development / cellular response to leukemia inhibitory factor / G1/S transition of mitotic cell cycle / RNA polymerase II transcription regulator complex / calcium ion transport / positive regulation of neuron apoptotic process / regulation of gene expression / mitotic cell cycle / DNA-binding transcription activator activity, RNA polymerase II-specific / in utero embryonic development / DNA-binding transcription factor activity, RNA polymerase II-specific / RNA polymerase II cis-regulatory region sequence-specific DNA binding / DNA-binding transcription factor activity / regulation of DNA-templated transcription / positive regulation of transcription by RNA polymerase II / DNA binding / nucleoplasm / nucleus / cytosol Similarity search - Function | ||||||
| Biological species |  | ||||||
| Method | SOLUTION NMR | ||||||
 Authors Authors | Ogata, K. / Morikawa, S. / Nakamura, H. / Hojo, H. / Yoshimura, S. / Zhang, R. / Aimoto, S. / Ametani, Y. / Hirata, Z. / Sarai, A. ...Ogata, K. / Morikawa, S. / Nakamura, H. / Hojo, H. / Yoshimura, S. / Zhang, R. / Aimoto, S. / Ametani, Y. / Hirata, Z. / Sarai, A. / Ishii, S. / Nishimura, Y. | ||||||
 Citation Citation |  Journal: Nat.Struct.Biol. / Year: 1995 Journal: Nat.Struct.Biol. / Year: 1995Title: Comparison of the free and DNA-complexed forms of the DNA-binding domain from c-Myb. Authors: Ogata, K. / Morikawa, S. / Nakamura, H. / Hojo, H. / Yoshimura, S. / Zhang, R. / Aimoto, S. / Ametani, Y. / Hirata, Z. / Sarai, A. / Ishii, S. / Nishimura, Y. #1:  Journal: Cell(Cambridge,Mass.) / Year: 1994 Journal: Cell(Cambridge,Mass.) / Year: 1994Title: Solution Structure of a Specific DNA Complex of the Myb DNA-Binding Domain with Cooperative Recognition Helices Authors: Ogata, K. / Morikawa, S. / Nakamura, H. / Sekikawa, A. / Inoue, T. / Kanai, H. / Sarai, A. / Ishii, S. / Nishimura, Y. #2:  Journal: Proc.Natl.Acad.Sci.USA / Year: 1992 Journal: Proc.Natl.Acad.Sci.USA / Year: 1992Title: Solution Structure of a DNA-Binding Unit of Myb: A Helix-Turn-Helix-Related Motif with Conserved Tryptophans Forming a Hydrophobic Core Authors: Ogata, K. / Hojo, H. / Aimoto, S. / Nakai, T. / Nakamura, H. / Sarai, A. / Ishii, S. / Nishimura, Y. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  1mbj.cif.gz 1mbj.cif.gz | 28.2 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb1mbj.ent.gz pdb1mbj.ent.gz | 17.7 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  1mbj.json.gz 1mbj.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Summary document |  1mbj_validation.pdf.gz 1mbj_validation.pdf.gz | 326.4 KB | Display |  wwPDB validaton report wwPDB validaton report |
|---|---|---|---|---|
| Full document |  1mbj_full_validation.pdf.gz 1mbj_full_validation.pdf.gz | 326.1 KB | Display | |
| Data in XML |  1mbj_validation.xml.gz 1mbj_validation.xml.gz | 2.4 KB | Display | |
| Data in CIF |  1mbj_validation.cif.gz 1mbj_validation.cif.gz | 2.8 KB | Display | |
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/mb/1mbj https://data.pdbj.org/pub/pdb/validation_reports/mb/1mbj ftp://data.pdbj.org/pub/pdb/validation_reports/mb/1mbj ftp://data.pdbj.org/pub/pdb/validation_reports/mb/1mbj | HTTPS FTP |
-Related structure data
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| 1 |
| |||||||||
| NMR ensembles |
|
- Components
Components
| #1: Protein | Mass: 6259.194 Da / Num. of mol.: 1 / Source method: isolated from a natural source Details: GENE C-MYB, NMR, MINIMIZED AVERAGE STRUCTURE; ENGINEERED Source: (natural)  |
|---|
-Experimental details
-Experiment
| Experiment | Method: SOLUTION NMR |
|---|
- Sample preparation
Sample preparation
| Sample conditions |
| |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Crystal grow | *PLUS Method: other / Details: NMR |
- Processing
Processing
| Software | Name:  AMBER / Classification: refinement AMBER / Classification: refinement | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| NMR software |
| |||||||||
| Refinement | Software ordinal: 1 Details: A TOTAL OF 50 STRUCTURES WERE CALCULATED. THE COORDINATES OF THE RESTRAINED MINIMIZED AVERAGED STRUCTURE WERE OBTAINED BY AVERAGING THE COORDINATES OF THE INDIVIDUAL STRUCTURES, AND THEN BY ...Details: A TOTAL OF 50 STRUCTURES WERE CALCULATED. THE COORDINATES OF THE RESTRAINED MINIMIZED AVERAGED STRUCTURE WERE OBTAINED BY AVERAGING THE COORDINATES OF THE INDIVIDUAL STRUCTURES, AND THEN BY SUBJECTING THE RESULTING COORDINATES TO THE RESTRAINED ENERGY MINIMIZATION BY PRESTO. | |||||||||
| NMR ensemble | Conformers submitted total number: 1 |
 Movie
Movie Controller
Controller

















 PDBj
PDBj
