[English] 日本語
 Yorodumi
Yorodumi- PDB-1eq8: THREE-DIMENSIONAL STRUCTURE OF THE PENTAMERIC HELICAL BUNDLE OF T... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 1eq8 | ||||||
|---|---|---|---|---|---|---|---|
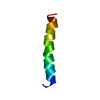
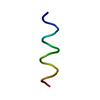
| Title | THREE-DIMENSIONAL STRUCTURE OF THE PENTAMERIC HELICAL BUNDLE OF THE ACETYLCHOLINE RECEPTOR M2 TRANSMEMBRANE SEGMENT | ||||||
 Components Components | ACETYLCHOLINE RECEPTOR PROTEIN | ||||||
 Keywords Keywords | SIGNALING PROTEIN / NEUROTRANSMITTER RECEPTOR / M2 / LIPID BILAYERS / ION-CHANNEL / HELICAL BUNDLE / PENTAMERIC BUNDLE | ||||||
| Function / homology |  Function and homology information Function and homology informationacetylcholine-gated monoatomic cation-selective channel activity / transmembrane signaling receptor activity / postsynaptic membrane / neuron projection Similarity search - Function | ||||||
| Biological species |  | ||||||
| Method | SOLID-STATE NMR / X-PLOR 3.1, distance geometry, simulated annealing, molecular dynamics, FISI, FINGERPRINT, HOLE | ||||||
 Authors Authors | Marassi, F.M. / Gesell, J.J. / Kim, Y. / Valente, A.P. / Oblatt-Montal, M. / Montal, M. / Opella, S.J. | ||||||
 Citation Citation |  Journal: Nat.Struct.Biol. / Year: 1999 Journal: Nat.Struct.Biol. / Year: 1999Title: Structures of the M2 channel-lining segments from nicotinic acetylcholine and NMDA receptors by NMR spectroscopy. Authors: Opella, S.J. / Marassi, F.M. / Gesell, J.J. / Valente, A.P. / Kim, Y. / Oblatt-Montal, M. / Montal, M. #1:  Journal: J.BIOMOL.NMR / Year: 1999 Journal: J.BIOMOL.NMR / Year: 1999Title: Dilute Spin-Exchange Assignment of Solid-State NMR Spectra of Oriented Proteins: Acetylcholine M2 in Bilayers Authors: Marassi, F.M. / Gesell, J.J. / Valente, A.P. / Kim, Y. / Oblatt-Montal, M. / Montal, M. / Opella, S.J. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  1eq8.cif.gz 1eq8.cif.gz | 43.2 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb1eq8.ent.gz pdb1eq8.ent.gz | 34.7 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  1eq8.json.gz 1eq8.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Summary document |  1eq8_validation.pdf.gz 1eq8_validation.pdf.gz | 307.4 KB | Display |  wwPDB validaton report wwPDB validaton report |
|---|---|---|---|---|
| Full document |  1eq8_full_validation.pdf.gz 1eq8_full_validation.pdf.gz | 307.2 KB | Display | |
| Data in XML |  1eq8_validation.xml.gz 1eq8_validation.xml.gz | 3 KB | Display | |
| Data in CIF |  1eq8_validation.cif.gz 1eq8_validation.cif.gz | 4.3 KB | Display | |
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/eq/1eq8 https://data.pdbj.org/pub/pdb/validation_reports/eq/1eq8 ftp://data.pdbj.org/pub/pdb/validation_reports/eq/1eq8 ftp://data.pdbj.org/pub/pdb/validation_reports/eq/1eq8 | HTTPS FTP |
-Related structure data
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| 1 |
| |||||||||
| NMR ensembles |
|
- Components
Components
| #1: Protein/peptide | Mass: 2522.013 Da / Num. of mol.: 5 / Fragment: M2 SEGMENT Source method: isolated from a genetically manipulated source Source: (gene. exp.)  Cell: NEURON / Cellular location: POST-SYNAPTIC MEMBRANE / Organ: BRAIN / Plasmid: PMAL / Species (production host): Escherichia coli / Production host:  #2: Chemical | ChemComp-OH / | |
|---|
-Experimental details
-Experiment
| Experiment | Method: SOLID-STATE NMR | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| NMR experiment |
| ||||||||||||
| NMR details | Text: The orientation of each monomer in the pentameric bundle was obtained from the combination of the solution NMR (PDB file 1a11) and solid-state NMR (PDB file 1cek) structures |
- Sample preparation
Sample preparation
| Details |
| ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Sample conditions |
| ||||||||||||
| Crystal grow | *PLUS Method: other / Details: NMR |
-NMR measurement
| NMR spectrometer |
|
|---|
- Processing
Processing
| NMR software |
| ||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method: X-PLOR 3.1, distance geometry, simulated annealing, molecular dynamics, FISI, FINGERPRINT, HOLE Software ordinal: 1 Details: The backbone coordinates obtained from solution NMR were superimposed on the coordinates obtained from solid-state NMR to fix the helix orientation and rotation in the membrane. The ...Details: The backbone coordinates obtained from solution NMR were superimposed on the coordinates obtained from solid-state NMR to fix the helix orientation and rotation in the membrane. The pentameric array was then optimized using molecular dynamics. Pore contours were calculated with the program HOLE. | ||||||||||||||||||||
| NMR representative | Selection criteria: minimized average structure | ||||||||||||||||||||
| NMR ensemble | Conformer selection criteria: structures with favorable non-bond energy Conformers calculated total number: 30 / Conformers submitted total number: 1 |
 Movie
Movie Controller
Controller









 PDBj
PDBj






 HSQC
HSQC