+ データを開く
データを開く
- 基本情報
基本情報
| 登録情報 |  | |||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| タイトル | RNA polymerase II transcribing a chromatosome (type I) | |||||||||||||||||||||
 マップデータ マップデータ | ||||||||||||||||||||||
 試料 試料 |
| |||||||||||||||||||||
| 機能・相同性 |  機能・相同性情報 機能・相同性情報: / : / regulation of septum digestion after cytokinesis / regulatory ncRNA-mediated heterochromatin formation / negative regulation of DNA recombination / nuclear DNA-directed RNA polymerase complex / RPB4-RPB7 complex /  RNA polymerase III activity / Apoptosis induced DNA fragmentation / nuclear-transcribed mRNA catabolic process, deadenylation-dependent decay ...: / : / regulation of septum digestion after cytokinesis / regulatory ncRNA-mediated heterochromatin formation / negative regulation of DNA recombination / nuclear DNA-directed RNA polymerase complex / RPB4-RPB7 complex / RNA polymerase III activity / Apoptosis induced DNA fragmentation / nuclear-transcribed mRNA catabolic process, deadenylation-dependent decay ...: / : / regulation of septum digestion after cytokinesis / regulatory ncRNA-mediated heterochromatin formation / negative regulation of DNA recombination / nuclear DNA-directed RNA polymerase complex / RPB4-RPB7 complex /  RNA polymerase III activity / Apoptosis induced DNA fragmentation / nuclear-transcribed mRNA catabolic process, deadenylation-dependent decay / chromosome condensation / nucleosomal DNA binding / : / Formation of Senescence-Associated Heterochromatin Foci (SAHF) / termination of RNA polymerase II transcription / termination of RNA polymerase III transcription / RNA polymerase III activity / Apoptosis induced DNA fragmentation / nuclear-transcribed mRNA catabolic process, deadenylation-dependent decay / chromosome condensation / nucleosomal DNA binding / : / Formation of Senescence-Associated Heterochromatin Foci (SAHF) / termination of RNA polymerase II transcription / termination of RNA polymerase III transcription /  RNA polymerase I activity / termination of RNA polymerase I transcription / RNA polymerase I activity / termination of RNA polymerase I transcription /  transcription initiation at RNA polymerase III promoter / tRNA transcription by RNA polymerase III / transcription initiation at RNA polymerase III promoter / tRNA transcription by RNA polymerase III /  transcription initiation at RNA polymerase I promoter / transcription elongation by RNA polymerase I / positive regulation of translational initiation / transcription initiation at RNA polymerase I promoter / transcription elongation by RNA polymerase I / positive regulation of translational initiation /  RNA polymerase II activity / negative regulation of tumor necrosis factor-mediated signaling pathway / RNA polymerase II activity / negative regulation of tumor necrosis factor-mediated signaling pathway /  RNA polymerase I complex / transcription by RNA polymerase I / RNA polymerase I complex / transcription by RNA polymerase I /  RNA polymerase III complex / negative regulation of megakaryocyte differentiation / transcription by RNA polymerase III / protein localization to CENP-A containing chromatin / pericentric heterochromatin / RNA polymerase III complex / negative regulation of megakaryocyte differentiation / transcription by RNA polymerase III / protein localization to CENP-A containing chromatin / pericentric heterochromatin /  RNA polymerase II, core complex / Chromatin modifying enzymes / Replacement of protamines by nucleosomes in the male pronucleus / CENP-A containing nucleosome / RNA polymerase II, core complex / Chromatin modifying enzymes / Replacement of protamines by nucleosomes in the male pronucleus / CENP-A containing nucleosome /  translation initiation factor binding / epigenetic regulation of gene expression / Packaging Of Telomere Ends / Recognition and association of DNA glycosylase with site containing an affected purine / Cleavage of the damaged purine / Deposition of new CENPA-containing nucleosomes at the centromere / Recognition and association of DNA glycosylase with site containing an affected pyrimidine / Cleavage of the damaged pyrimidine / Inhibition of DNA recombination at telomere / Meiotic synapsis / telomere organization / RNA Polymerase I Promoter Opening / Interleukin-7 signaling / Assembly of the ORC complex at the origin of replication / SUMOylation of chromatin organization proteins / translation initiation factor binding / epigenetic regulation of gene expression / Packaging Of Telomere Ends / Recognition and association of DNA glycosylase with site containing an affected purine / Cleavage of the damaged purine / Deposition of new CENPA-containing nucleosomes at the centromere / Recognition and association of DNA glycosylase with site containing an affected pyrimidine / Cleavage of the damaged pyrimidine / Inhibition of DNA recombination at telomere / Meiotic synapsis / telomere organization / RNA Polymerase I Promoter Opening / Interleukin-7 signaling / Assembly of the ORC complex at the origin of replication / SUMOylation of chromatin organization proteins /  DNAメチル化 / DNAメチル化 /  DNA-directed RNA polymerase complex / Condensation of Prophase Chromosomes / ERCC6 (CSB) and EHMT2 (G9a) positively regulate rRNA expression / Chromatin modifications during the maternal to zygotic transition (MZT) / SIRT1 negatively regulates rRNA expression / HCMV Late Events / DNA-directed RNA polymerase complex / Condensation of Prophase Chromosomes / ERCC6 (CSB) and EHMT2 (G9a) positively regulate rRNA expression / Chromatin modifications during the maternal to zygotic transition (MZT) / SIRT1 negatively regulates rRNA expression / HCMV Late Events /  innate immune response in mucosa / PRC2 methylates histones and DNA / Defective pyroptosis / innate immune response in mucosa / PRC2 methylates histones and DNA / Defective pyroptosis /  P-body / HDACs deacetylate histones / transcription elongation by RNA polymerase II / P-body / HDACs deacetylate histones / transcription elongation by RNA polymerase II /  transcription initiation at RNA polymerase II promoter / RNA Polymerase I Promoter Escape / transcription initiation at RNA polymerase II promoter / RNA Polymerase I Promoter Escape /  lipopolysaccharide binding / Nonhomologous End-Joining (NHEJ) / Transcriptional regulation by small RNAs / Formation of the beta-catenin:TCF transactivating complex / lipopolysaccharide binding / Nonhomologous End-Joining (NHEJ) / Transcriptional regulation by small RNAs / Formation of the beta-catenin:TCF transactivating complex /  ユークロマチン / RUNX1 regulates genes involved in megakaryocyte differentiation and platelet function / Activated PKN1 stimulates transcription of AR (androgen receptor) regulated genes KLK2 and KLK3 / NoRC negatively regulates rRNA expression / B-WICH complex positively regulates rRNA expression / G2/M DNA damage checkpoint / HDMs demethylate histones / DNA Damage/Telomere Stress Induced Senescence / ユークロマチン / RUNX1 regulates genes involved in megakaryocyte differentiation and platelet function / Activated PKN1 stimulates transcription of AR (androgen receptor) regulated genes KLK2 and KLK3 / NoRC negatively regulates rRNA expression / B-WICH complex positively regulates rRNA expression / G2/M DNA damage checkpoint / HDMs demethylate histones / DNA Damage/Telomere Stress Induced Senescence /  ribonucleoside binding / Metalloprotease DUBs / chromatin DNA binding / PKMTs methylate histone lysines / RMTs methylate histone arginines / ribonucleoside binding / Metalloprotease DUBs / chromatin DNA binding / PKMTs methylate histone lysines / RMTs methylate histone arginines /  遺伝的組換え / Pre-NOTCH Transcription and Translation / DNA-directed 5'-3' RNA polymerase activity / 遺伝的組換え / Pre-NOTCH Transcription and Translation / DNA-directed 5'-3' RNA polymerase activity /  ポリメラーゼ / ポリメラーゼ /  nucleosome assembly / Activation of anterior HOX genes in hindbrain development during early embryogenesis / HCMV Early Events / Transcriptional regulation of granulopoiesis / structural constituent of chromatin / UCH proteinases / nucleosome assembly / Activation of anterior HOX genes in hindbrain development during early embryogenesis / HCMV Early Events / Transcriptional regulation of granulopoiesis / structural constituent of chromatin / UCH proteinases /  ヌクレオソーム / antimicrobial humoral immune response mediated by antimicrobial peptide / ヌクレオソーム / antimicrobial humoral immune response mediated by antimicrobial peptide /  single-stranded DNA binding / E3 ubiquitin ligases ubiquitinate target proteins / Recruitment and ATM-mediated phosphorylation of repair and signaling proteins at DNA double strand breaks / RUNX1 regulates transcription of genes involved in differentiation of HSCs / single-stranded DNA binding / E3 ubiquitin ligases ubiquitinate target proteins / Recruitment and ATM-mediated phosphorylation of repair and signaling proteins at DNA double strand breaks / RUNX1 regulates transcription of genes involved in differentiation of HSCs /  遺伝子発現 遺伝子発現類似検索 - 分子機能 | |||||||||||||||||||||
| 生物種 |  Komagataella phaffii (菌類) / Komagataella phaffii (菌類) /   Homo sapiens (ヒト) / synthetic construct (人工物) Homo sapiens (ヒト) / synthetic construct (人工物) | |||||||||||||||||||||
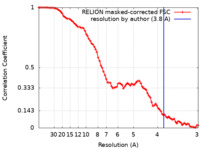
| 手法 |  単粒子再構成法 / 単粒子再構成法 /  クライオ電子顕微鏡法 / 解像度: 3.8 Å クライオ電子顕微鏡法 / 解像度: 3.8 Å | |||||||||||||||||||||
 データ登録者 データ登録者 | Hirano R / Ehara H / Tomoya K / Takizawa Y / Sekine S / Kurumizaka H | |||||||||||||||||||||
| 資金援助 |  日本, 6件 日本, 6件
| |||||||||||||||||||||
 引用 引用 |  ジャーナル: Nat Commun / 年: 2022 ジャーナル: Nat Commun / 年: 2022タイトル: Structural basis of RNA polymerase II transcription on the chromatosome containing linker histone H1. 著者: Rina Hirano / Haruhiko Ehara / Tomoya Kujirai / Tamami Uejima / Yoshimasa Takizawa / Shun-Ichi Sekine / Hitoshi Kurumizaka /  要旨: In chromatin, linker histone H1 binds to nucleosomes, forming chromatosomes, and changes the transcription status. However, the mechanism by which RNA polymerase II (RNAPII) transcribes the DNA in ...In chromatin, linker histone H1 binds to nucleosomes, forming chromatosomes, and changes the transcription status. However, the mechanism by which RNA polymerase II (RNAPII) transcribes the DNA in the chromatosome has remained enigmatic. Here we report the cryo-electron microscopy (cryo-EM) structures of transcribing RNAPII-chromatosome complexes (forms I and II), in which RNAPII is paused at the entry linker DNA region of the chromatosome due to H1 binding. In the form I complex, the H1 bound to the nucleosome restricts the linker DNA orientation, and the exit linker DNA is captured by the RNAPII DNA binding cleft. In the form II complex, the RNAPII progresses a few bases ahead by releasing the exit linker DNA from the RNAPII cleft, and directly clashes with the H1 bound to the nucleosome. The transcription elongation factor Spt4/5 masks the RNAPII DNA binding region, and drastically reduces the H1-mediated RNAPII pausing. | |||||||||||||||||||||
| 履歴 |
|
- 構造の表示
構造の表示
| 添付画像 |
|---|
- ダウンロードとリンク
ダウンロードとリンク
-EMDBアーカイブ
| マップデータ |  emd_34415.map.gz emd_34415.map.gz | 40.7 MB |  EMDBマップデータ形式 EMDBマップデータ形式 | |
|---|---|---|---|---|
| ヘッダ (付随情報) |  emd-34415-v30.xml emd-34415-v30.xml emd-34415.xml emd-34415.xml | 45.2 KB 45.2 KB | 表示 表示 |  EMDBヘッダ EMDBヘッダ |
| FSC (解像度算出) |  emd_34415_fsc.xml emd_34415_fsc.xml | 8.5 KB | 表示 |  FSCデータファイル FSCデータファイル |
| 画像 |  emd_34415.png emd_34415.png | 63.7 KB | ||
| その他 |  emd_34415_additional_1.map.gz emd_34415_additional_1.map.gz emd_34415_additional_2.map.gz emd_34415_additional_2.map.gz emd_34415_half_map_1.map.gz emd_34415_half_map_1.map.gz emd_34415_half_map_2.map.gz emd_34415_half_map_2.map.gz | 40.8 MB 40.7 MB 40.8 MB 40.8 MB | ||
| アーカイブディレクトリ |  http://ftp.pdbj.org/pub/emdb/structures/EMD-34415 http://ftp.pdbj.org/pub/emdb/structures/EMD-34415 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-34415 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-34415 | HTTPS FTP |
-関連構造データ
| 関連構造データ |  8h0vMC  8h0wC M: このマップから作成された原子モデル C: 同じ文献を引用 ( |
|---|---|
| 類似構造データ | 類似検索 - 機能・相同性  F&H 検索 F&H 検索 |
- リンク
リンク
| EMDBのページ |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| 「今月の分子」の関連する項目 |
- マップ
マップ
| ファイル |  ダウンロード / ファイル: emd_34415.map.gz / 形式: CCP4 / 大きさ: 52.7 MB / タイプ: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) ダウンロード / ファイル: emd_34415.map.gz / 形式: CCP4 / 大きさ: 52.7 MB / タイプ: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| ボクセルのサイズ | X=Y=Z: 1.49 Å | ||||||||||||||||||||
| 密度 |
| ||||||||||||||||||||
| 対称性 | 空間群: 1 | ||||||||||||||||||||
| 詳細 | EMDB XML:
|
-添付データ
-追加マップ: RNAP reconstruction
| ファイル | emd_34415_additional_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 注釈 | RNAP reconstruction | ||||||||||||
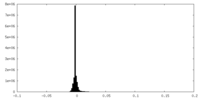
| 投影像・断面図 |
| ||||||||||||
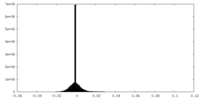
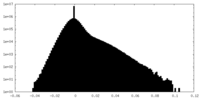
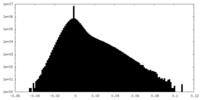
| 密度ヒストグラム |
-追加マップ: Nucleosome reconstruction
| ファイル | emd_34415_additional_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 注釈 | Nucleosome reconstruction | ||||||||||||
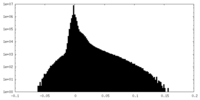
| 投影像・断面図 |
| ||||||||||||
| 密度ヒストグラム |
-ハーフマップ: #1
| ファイル | emd_34415_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
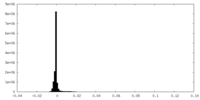
| 投影像・断面図 |
| ||||||||||||
| 密度ヒストグラム |
-ハーフマップ: #2
| ファイル | emd_34415_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
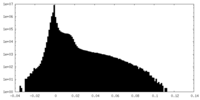
| 投影像・断面図 |
| ||||||||||||
| 密度ヒストグラム |
- 試料の構成要素
試料の構成要素
+全体 : RNA polymerase II transcribing a chromatosome (type I)
+超分子 #1: RNA polymerase II transcribing a chromatosome (type I)
+超分子 #2: RNA polymerase II
+超分子 #3: DNA,RNA
+超分子 #4: Histone
+分子 #1: DNA-directed RNA polymerase subunit
+分子 #2: DNA-directed RNA polymerase subunit beta
+分子 #3: RNA polymerase II third largest subunit B44, part of central core
+分子 #4: RNA polymerase II subunit B32
+分子 #5: DNA-directed RNA polymerases I, II, and III subunit RPABC1
+分子 #6: RNA polymerase subunit ABC23, common to RNA polymerases I, II, and III
+分子 #7: RNA polymerase II subunit
+分子 #8: DNA-directed RNA polymerases I, II, and III subunit RPABC3
+分子 #9: DNA-directed RNA polymerase subunit
+分子 #10: RNA polymerase subunit ABC10-beta, common to RNA polymerases I, I...
+分子 #11: RNA polymerase II subunit B12.5
+分子 #12: RNA polymerase subunit ABC10-alpha
+分子 #16: Histone H3.1
+分子 #17: Histone H4
+分子 #18: Histone H2A type 1-B/E
+分子 #19: Histone H2B type 1-J
+分子 #20: Histone H1.2
+分子 #13: RNA (5'-R(P*AP*GP*CP*AP*AP*UP*AP*GP*GP*AP*GP*CP*UP*U)-3')
+分子 #14: DNA (261-MER)
+分子 #15: DNA (261-MER)
+分子 #21: ZINC ION
+分子 #22: MAGNESIUM ION
-実験情報
-構造解析
| 手法 |  クライオ電子顕微鏡法 クライオ電子顕微鏡法 |
|---|---|
 解析 解析 |  単粒子再構成法 単粒子再構成法 |
| 試料の集合状態 | particle |
- 試料調製
試料調製
| 緩衝液 | pH: 7.5 |
|---|---|
| 凍結 | 凍結剤: ETHANE |
- 電子顕微鏡法
電子顕微鏡法
| 顕微鏡 | FEI TITAN KRIOS |
|---|---|
| 電子線 | 加速電圧: 300 kV / 電子線源:  FIELD EMISSION GUN FIELD EMISSION GUN |
| 電子光学系 | 照射モード: FLOOD BEAM / 撮影モード: BRIGHT FIELD Bright-field microscopy Bright-field microscopy最大 デフォーカス(公称値): 2.3000000000000003 µm 最小 デフォーカス(公称値): 1.2 µm |
| 撮影 | フィルム・検出器のモデル: GATAN K3 BIOQUANTUM (6k x 4k) 平均電子線量: 56.2 e/Å2 |
| 実験機器 |  モデル: Titan Krios / 画像提供: FEI Company |
 ムービー
ムービー コントローラー
コントローラー
























 Z
Z Y
Y X
X