+ データを開く
データを開く
- 基本情報
基本情報
| 登録情報 |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| タイトル | Nav1.7 mutant class2 | |||||||||
 マップデータ マップデータ | ||||||||||
 試料 試料 |
| |||||||||
| 機能・相同性 |  機能・相同性情報 機能・相同性情報corticospinal neuron axon guidance / positive regulation of voltage-gated sodium channel activity / response to pyrethroid / detection of mechanical stimulus involved in sensory perception / voltage-gated sodium channel activity involved in Purkinje myocyte action potential / voltage-gated sodium channel activity involved in cardiac muscle cell action potential / regulation of sodium ion transmembrane transporter activity / voltage-gated potassium channel activity involved in ventricular cardiac muscle cell action potential repolarization / membrane depolarization during Purkinje myocyte cell action potential / regulation of atrial cardiac muscle cell membrane depolarization ...corticospinal neuron axon guidance / positive regulation of voltage-gated sodium channel activity / response to pyrethroid / detection of mechanical stimulus involved in sensory perception / voltage-gated sodium channel activity involved in Purkinje myocyte action potential / voltage-gated sodium channel activity involved in cardiac muscle cell action potential / regulation of sodium ion transmembrane transporter activity / voltage-gated potassium channel activity involved in ventricular cardiac muscle cell action potential repolarization / membrane depolarization during Purkinje myocyte cell action potential / regulation of atrial cardiac muscle cell membrane depolarization / cardiac conduction /  voltage-gated sodium channel complex / membrane depolarization during cardiac muscle cell action potential / positive regulation of sodium ion transport / voltage-gated sodium channel complex / membrane depolarization during cardiac muscle cell action potential / positive regulation of sodium ion transport /  ランヴィエの絞輪 / cardiac muscle cell action potential involved in contraction / high voltage-gated calcium channel activity / locomotion / ランヴィエの絞輪 / cardiac muscle cell action potential involved in contraction / high voltage-gated calcium channel activity / locomotion /  voltage-gated sodium channel activity / regulation of ventricular cardiac muscle cell membrane repolarization / voltage-gated sodium channel activity / regulation of ventricular cardiac muscle cell membrane repolarization /  sodium channel inhibitor activity / neuronal action potential propagation / Interaction between L1 and Ankyrins / sodium ion transport / sodium channel inhibitor activity / neuronal action potential propagation / Interaction between L1 and Ankyrins / sodium ion transport /  voltage-gated calcium channel complex / behavioral response to pain / Phase 0 - rapid depolarisation / regulation of heart rate by cardiac conduction / voltage-gated calcium channel complex / behavioral response to pain / Phase 0 - rapid depolarisation / regulation of heart rate by cardiac conduction /  脱分極 / detection of temperature stimulus involved in sensory perception of pain / calcium ion import across plasma membrane / sodium channel regulator activity / sodium ion transmembrane transport / 脱分極 / detection of temperature stimulus involved in sensory perception of pain / calcium ion import across plasma membrane / sodium channel regulator activity / sodium ion transmembrane transport /  介在板 / cardiac muscle contraction / sensory perception of pain / 介在板 / cardiac muscle contraction / sensory perception of pain /  横行小管 / post-embryonic development / 横行小管 / post-embryonic development /  軸索誘導 / response to toxic substance / Sensory perception of sweet, bitter, and umami (glutamate) taste / positive regulation of neuron projection development / 軸索誘導 / response to toxic substance / Sensory perception of sweet, bitter, and umami (glutamate) taste / positive regulation of neuron projection development /  概日リズム / 概日リズム /  nervous system development / nervous system development /  遺伝子発現 / response to heat / 遺伝子発現 / response to heat /  perikaryon / chemical synaptic transmission / transmembrane transporter binding / perikaryon / chemical synaptic transmission / transmembrane transporter binding /  細胞接着 / 細胞接着 /  炎症 / 炎症 /  神経繊維 / 神経繊維 /  シナプス / extracellular region / シナプス / extracellular region /  細胞膜 細胞膜類似検索 - 分子機能 | |||||||||
| 生物種 |   Homo sapiens (ヒト) Homo sapiens (ヒト) | |||||||||
| 手法 |  単粒子再構成法 / 単粒子再構成法 /  クライオ電子顕微鏡法 / 解像度: 2.8 Å クライオ電子顕微鏡法 / 解像度: 2.8 Å | |||||||||
 データ登録者 データ登録者 | Huang G / Wu Q / Li Z / Pan X / Yan N | |||||||||
| 資金援助 |  米国, 米国,  中国, 2件 中国, 2件
| |||||||||
 引用 引用 |  ジャーナル: Proc Natl Acad Sci U S A / 年: 2022 ジャーナル: Proc Natl Acad Sci U S A / 年: 2022タイトル: Unwinding and spiral sliding of S4 and domain rotation of VSD during the electromechanical coupling in Na1.7. 著者: Gaoxingyu Huang / Qiurong Wu / Zhangqiang Li / Xueqin Jin / Xiaoshuang Huang / Tong Wu / Xiaojing Pan / Nieng Yan /  要旨: Voltage-gated sodium (Na) channel Na1.7 has been targeted for the development of nonaddictive pain killers. Structures of Na1.7 in distinct functional states will offer an advanced mechanistic ...Voltage-gated sodium (Na) channel Na1.7 has been targeted for the development of nonaddictive pain killers. Structures of Na1.7 in distinct functional states will offer an advanced mechanistic understanding and aid drug discovery. Here we report the cryoelectron microscopy analysis of a human Na1.7 variant that, with 11 rationally introduced point mutations, has a markedly right-shifted activation voltage curve with V reaching 69 mV. The voltage-sensing domain in the first repeat (VSD) in a 2.7-Å resolution structure displays a completely down (deactivated) conformation. Compared to the structure of WT Na1.7, three gating charge (GC) residues in VSD are transferred to the cytosolic side through a combination of helix unwinding and spiral sliding of S4 and ∼20° domain rotation. A conserved WNФФD motif on the cytoplasmic end of S3 stabilizes the down conformation of VSD. One GC residue is transferred in VSD mainly through helix sliding. Accompanying GC transfer in VSD and VSD, rearrangement and contraction of the intracellular gate is achieved through concerted movements of adjacent segments, including S4-5, S4-5, S5, and all S6 segments. Our studies provide important insight into the electromechanical coupling mechanism of the single-chain voltage-gated ion channels and afford molecular interpretations for a number of pain-associated mutations whose pathogenic mechanism cannot be revealed from previously reported Na structures. | |||||||||
| 履歴 |
|
- 構造の表示
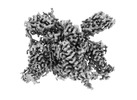
構造の表示
| 添付画像 |
|---|
- ダウンロードとリンク
ダウンロードとリンク
-EMDBアーカイブ
| マップデータ |  emd_33485.map.gz emd_33485.map.gz | 118 MB |  EMDBマップデータ形式 EMDBマップデータ形式 | |
|---|---|---|---|---|
| ヘッダ (付随情報) |  emd-33485-v30.xml emd-33485-v30.xml emd-33485.xml emd-33485.xml | 20.6 KB 20.6 KB | 表示 表示 |  EMDBヘッダ EMDBヘッダ |
| 画像 |  emd_33485.png emd_33485.png | 55.5 KB | ||
| その他 |  emd_33485_half_map_1.map.gz emd_33485_half_map_1.map.gz emd_33485_half_map_2.map.gz emd_33485_half_map_2.map.gz | 116 MB 116 MB | ||
| アーカイブディレクトリ |  http://ftp.pdbj.org/pub/emdb/structures/EMD-33485 http://ftp.pdbj.org/pub/emdb/structures/EMD-33485 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-33485 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-33485 | HTTPS FTP |
-関連構造データ
| 関連構造データ |  7xvfMC  7xveC C: 同じ文献を引用 ( M: このマップから作成された原子モデル |
|---|---|
| 類似構造データ | 類似検索 - 機能・相同性  F&H 検索 F&H 検索 |
- リンク
リンク
| EMDBのページ |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| 「今月の分子」の関連する項目 |
- マップ
マップ
| ファイル |  ダウンロード / ファイル: emd_33485.map.gz / 形式: CCP4 / 大きさ: 125 MB / タイプ: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) ダウンロード / ファイル: emd_33485.map.gz / 形式: CCP4 / 大きさ: 125 MB / タイプ: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| ボクセルのサイズ | X=Y=Z: 1.0825 Å | ||||||||||||||||||||
| 密度 |
| ||||||||||||||||||||
| 対称性 | 空間群: 1 | ||||||||||||||||||||
| 詳細 | EMDB XML:
|
-添付データ
-ハーフマップ: #2
| ファイル | emd_33485_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
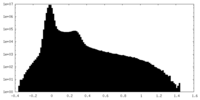
| 投影像・断面図 |
| ||||||||||||
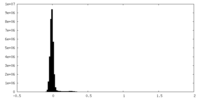
| 密度ヒストグラム |
-ハーフマップ: #1
| ファイル | emd_33485_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
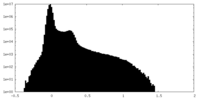
| 投影像・断面図 |
| ||||||||||||
| 密度ヒストグラム |
- 試料の構成要素
試料の構成要素
+全体 : alpha1-beta1-beta2 ternary complex
+超分子 #1: alpha1-beta1-beta2 ternary complex
+分子 #1: Sodium channel protein type 9 subunit alpha
+分子 #2: Sodium channel subunit beta-1
+分子 #3: Sodium channel subunit beta-2
+分子 #5: 2-acetamido-2-deoxy-beta-D-glucopyranose
+分子 #6: CHOLESTEROL HEMISUCCINATE
+分子 #7: CHOLESTEROL
+分子 #8: 1-O-OCTADECYL-SN-GLYCERO-3-PHOSPHOCHOLINE
+分子 #9: (2S,3R,4E)-2-(acetylamino)-3-hydroxyoctadec-4-en-1-yl dihydrogen ...
+分子 #10: 1,2-DIOLEOYL-SN-GLYCERO-3-PHOSPHOCHOLINE
+分子 #11: O-[(R)-{[(2R)-2,3-bis(octadecanoyloxy)propyl]oxy}(hydroxy)phospho...
-実験情報
-構造解析
| 手法 |  クライオ電子顕微鏡法 クライオ電子顕微鏡法 |
|---|---|
 解析 解析 |  単粒子再構成法 単粒子再構成法 |
| 試料の集合状態 | particle |
- 試料調製
試料調製
| 緩衝液 | pH: 7.5 |
|---|---|
| 凍結 | 凍結剤: ETHANE |
- 電子顕微鏡法
電子顕微鏡法
| 顕微鏡 | FEI TITAN KRIOS |
|---|---|
| 電子線 | 加速電圧: 300 kV / 電子線源:  FIELD EMISSION GUN FIELD EMISSION GUN |
| 電子光学系 | 照射モード: FLOOD BEAM / 撮影モード: BRIGHT FIELD Bright-field microscopy / 最大 デフォーカス(公称値): 1.8 µm / 最小 デフォーカス(公称値): 1.5 µm Bright-field microscopy / 最大 デフォーカス(公称値): 1.8 µm / 最小 デフォーカス(公称値): 1.5 µm |
| 撮影 | フィルム・検出器のモデル: GATAN K3 (6k x 4k) / 平均電子線量: 50.0 e/Å2 |
| 実験機器 |  モデル: Titan Krios / 画像提供: FEI Company |
- 画像解析
画像解析
| 初期モデル | モデルのタイプ: EMDB MAP EMDB ID: |
|---|---|
| 初期 角度割当 | タイプ: MAXIMUM LIKELIHOOD |
| 最終 角度割当 | タイプ: MAXIMUM LIKELIHOOD |
| 最終 再構成 | 想定した対称性 - 点群: C1 (非対称) / 解像度のタイプ: BY AUTHOR / 解像度: 2.8 Å / 解像度の算出法: FSC 0.143 CUT-OFF / 使用した粒子像数: 394163 |
 ムービー
ムービー コントローラー
コントローラー









 Z
Z Y
Y X
X